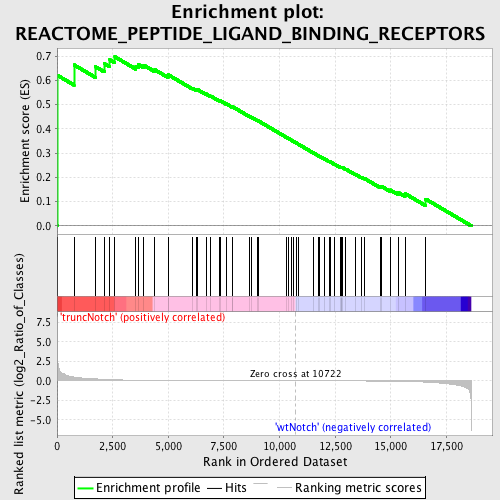
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_03_truncNotch_versus_wtNotch.phenotype_truncNotch_versus_wtNotch.cls #truncNotch_versus_wtNotch |
| Phenotype | phenotype_truncNotch_versus_wtNotch.cls#truncNotch_versus_wtNotch |
| Upregulated in class | truncNotch |
| GeneSet | REACTOME_PEPTIDE_LIGAND_BINDING_RECEPTORS |
| Enrichment Score (ES) | 0.6971042 |
| Normalized Enrichment Score (NES) | 1.5424408 |
| Nominal p-value | 0.012522361 |
| FDR q-value | 0.2876011 |
| FWER p-Value | 0.982 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GAL | 3930279 | 9 | 3.618 | 0.6212 | Yes | ||
| 2 | CCR7 | 2060687 | 763 | 0.484 | 0.6639 | Yes | ||
| 3 | CCL5 | 3710397 | 1721 | 0.258 | 0.6566 | Yes | ||
| 4 | F2 | 5720280 | 2123 | 0.199 | 0.6692 | Yes | ||
| 5 | PDYN | 5720408 | 2370 | 0.167 | 0.6847 | Yes | ||
| 6 | EDNRA | 6900133 | 2598 | 0.143 | 0.6971 | Yes | ||
| 7 | BDKRB1 | 730592 | 3541 | 0.075 | 0.6592 | No | ||
| 8 | NTSR1 | 6510136 | 3658 | 0.070 | 0.6649 | No | ||
| 9 | AVPR2 | 360064 | 3895 | 0.060 | 0.6626 | No | ||
| 10 | CX3CL1 | 3990707 | 4394 | 0.046 | 0.6437 | No | ||
| 11 | AGT | 7000575 | 4988 | 0.034 | 0.6176 | No | ||
| 12 | AVP | 2100113 | 4998 | 0.034 | 0.6229 | No | ||
| 13 | KNG1 | 6400576 6770347 | 6084 | 0.020 | 0.5679 | No | ||
| 14 | MC5R | 2640176 | 6284 | 0.018 | 0.5603 | No | ||
| 15 | AGTR1 | 4780524 2680592 | 6306 | 0.018 | 0.5623 | No | ||
| 16 | XCR1 | 5080072 | 6692 | 0.015 | 0.5442 | No | ||
| 17 | CCKBR | 2760128 | 6896 | 0.014 | 0.5356 | No | ||
| 18 | HCRTR2 | 2350463 | 7286 | 0.012 | 0.5167 | No | ||
| 19 | PPYR1 | 1410072 | 7328 | 0.011 | 0.5164 | No | ||
| 20 | IL8RA | 1580100 | 7607 | 0.010 | 0.5032 | No | ||
| 21 | MC2R | 2970504 | 7874 | 0.009 | 0.4904 | No | ||
| 22 | HCRTR1 | 1580273 | 7895 | 0.009 | 0.4909 | No | ||
| 23 | DARC | 3940050 | 8640 | 0.006 | 0.4519 | No | ||
| 24 | OPRM1 | 5360279 | 8737 | 0.006 | 0.4477 | No | ||
| 25 | CCR3 | 50427 | 8997 | 0.005 | 0.4346 | No | ||
| 26 | OXT | 3850692 | 9042 | 0.005 | 0.4330 | No | ||
| 27 | HCRT | 2970463 | 10328 | 0.001 | 0.3640 | No | ||
| 28 | PPY | 2340373 | 10388 | 0.001 | 0.3610 | No | ||
| 29 | NPFF | 2030280 | 10541 | 0.001 | 0.3529 | No | ||
| 30 | CCR10 | 4050097 | 10634 | 0.000 | 0.3480 | No | ||
| 31 | TACR1 | 70358 3840411 | 10741 | -0.000 | 0.3423 | No | ||
| 32 | C5AR1 | 4540402 | 10869 | -0.000 | 0.3355 | No | ||
| 33 | TACR3 | 4780017 | 11510 | -0.002 | 0.3014 | No | ||
| 34 | TACR2 | 1740358 | 11762 | -0.003 | 0.2884 | No | ||
| 35 | IL8RB | 450592 1170537 | 11796 | -0.003 | 0.2872 | No | ||
| 36 | EDN1 | 1770047 | 12016 | -0.004 | 0.2760 | No | ||
| 37 | CCR6 | 5720368 6020176 | 12251 | -0.005 | 0.2643 | No | ||
| 38 | GRPR | 6020170 | 12272 | -0.005 | 0.2641 | No | ||
| 39 | GALR1 | 6020452 | 12482 | -0.006 | 0.2538 | No | ||
| 40 | XCL1 | 3800504 | 12744 | -0.007 | 0.2409 | No | ||
| 41 | NTS | 380300 2120592 | 12776 | -0.007 | 0.2405 | No | ||
| 42 | SST | 6590142 | 12786 | -0.007 | 0.2412 | No | ||
| 43 | TAC1 | 7000195 380706 | 12833 | -0.007 | 0.2400 | No | ||
| 44 | CCK | 6370368 | 12965 | -0.008 | 0.2344 | No | ||
| 45 | CXCL11 | 1090551 | 13390 | -0.011 | 0.2134 | No | ||
| 46 | AVPR1A | 2120300 | 13659 | -0.013 | 0.2012 | No | ||
| 47 | CCL25 | 450541 540435 | 13809 | -0.014 | 0.1956 | No | ||
| 48 | KISS1 | 1580398 | 14542 | -0.025 | 0.1605 | No | ||
| 49 | CXCR6 | 3190440 | 14571 | -0.026 | 0.1634 | No | ||
| 50 | SSTR1 | 2630471 | 14971 | -0.036 | 0.1481 | No | ||
| 51 | PNOC | 1990114 | 15353 | -0.052 | 0.1364 | No | ||
| 52 | NMB | 3360735 | 15639 | -0.069 | 0.1330 | No | ||
| 53 | F2R | 4810180 | 16562 | -0.159 | 0.1106 | No |

