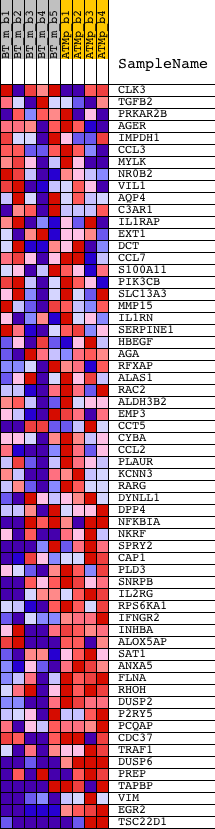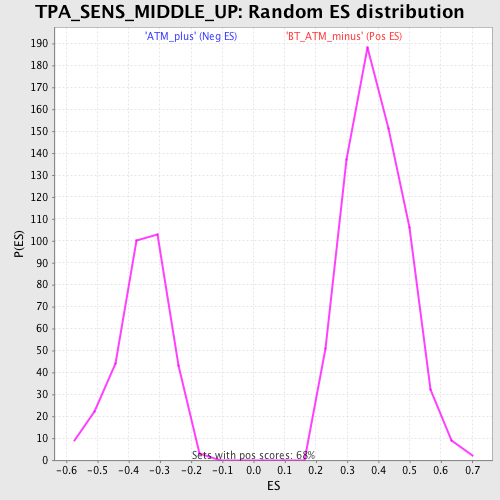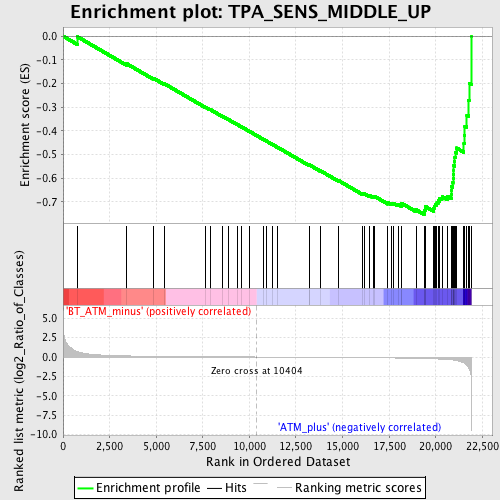
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | TPA_SENS_MIDDLE_UP |
| Enrichment Score (ES) | -0.7538494 |
| Normalized Enrichment Score (NES) | -2.1127124 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0070471945 |
| FWER p-Value | 0.0060 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | CLK3 | 1423849_a_at 1441472_at | 763 | 0.690 | -0.0020 | No | ||
| 2 | TGFB2 | 1423250_a_at 1438303_at 1446141_at 1450922_a_at 1450923_at | 3404 | 0.140 | -0.1161 | No | ||
| 3 | PRKAR2B | 1430640_a_at 1438664_at 1456475_s_at | 4847 | 0.090 | -0.1777 | No | ||
| 4 | AGER | 1420428_at 1441958_s_at | 5423 | 0.078 | -0.2003 | No | ||
| 5 | IMPDH1 | 1423239_at | 7635 | 0.038 | -0.2996 | No | ||
| 6 | CCL3 | 1419561_at | 7905 | 0.034 | -0.3103 | No | ||
| 7 | MYLK | 1425504_at 1425505_at 1425506_at | 8585 | 0.024 | -0.3401 | No | ||
| 8 | NR0B2 | 1449854_at | 8867 | 0.021 | -0.3520 | No | ||
| 9 | VIL1 | 1448837_at | 9365 | 0.014 | -0.3741 | No | ||
| 10 | AQP4 | 1425382_a_at 1434449_at 1447745_at | 9570 | 0.011 | -0.3829 | No | ||
| 11 | C3AR1 | 1419482_at 1419483_at 1442082_at | 10021 | 0.005 | -0.4032 | No | ||
| 12 | IL1RAP | 1421843_at 1439697_at 1442614_at 1449585_at | 10761 | -0.004 | -0.4368 | No | ||
| 13 | EXT1 | 1417730_at 1439877_at 1440011_at 1440092_at 1456655_at 1458054_at 1458296_at | 10901 | -0.007 | -0.4428 | No | ||
| 14 | DCT | 1418028_at | 11232 | -0.011 | -0.4574 | No | ||
| 15 | CCL7 | 1421228_at | 11527 | -0.015 | -0.4701 | No | ||
| 16 | S100A11 | 1460351_at | 13207 | -0.037 | -0.5451 | No | ||
| 17 | PIK3CB | 1445731_at 1453069_at | 13212 | -0.037 | -0.5435 | No | ||
| 18 | SLC13A3 | 1416560_at 1438377_x_at 1439384_at 1439385_x_at | 13802 | -0.046 | -0.5683 | No | ||
| 19 | MMP15 | 1422597_at 1437462_x_at 1439734_at | 14782 | -0.061 | -0.6101 | No | ||
| 20 | IL1RN | 1423017_a_at 1425663_at 1451798_at | 16076 | -0.084 | -0.6653 | No | ||
| 21 | SERPINE1 | 1419149_at | 16158 | -0.085 | -0.6649 | No | ||
| 22 | HBEGF | 1418349_at 1418350_at | 16439 | -0.092 | -0.6733 | No | ||
| 23 | AGA | 1434665_at | 16652 | -0.096 | -0.6784 | No | ||
| 24 | RFXAP | 1418859_at | 16707 | -0.098 | -0.6763 | No | ||
| 25 | ALAS1 | 1424126_at 1442331_at 1455282_x_at | 17439 | -0.117 | -0.7041 | No | ||
| 26 | RAC2 | 1417620_at 1440208_at | 17615 | -0.121 | -0.7064 | No | ||
| 27 | ALDH3B2 | 1456317_at | 17739 | -0.125 | -0.7060 | No | ||
| 28 | EMP3 | 1417104_at | 17994 | -0.133 | -0.7113 | No | ||
| 29 | CCT5 | 1417258_at | 18164 | -0.138 | -0.7125 | No | ||
| 30 | CYBA | 1454268_a_at | 18170 | -0.138 | -0.7062 | No | ||
| 31 | CCL2 | 1420380_at | 18955 | -0.168 | -0.7341 | No | ||
| 32 | PLAUR | 1452521_a_at | 19389 | -0.190 | -0.7448 | Yes | ||
| 33 | KCNN3 | 1421632_at 1459308_at | 19418 | -0.191 | -0.7370 | Yes | ||
| 34 | RARG | 1419415_a_at 1419416_a_at | 19450 | -0.194 | -0.7292 | Yes | ||
| 35 | DYNLL1 | 1448682_at | 19471 | -0.195 | -0.7208 | Yes | ||
| 36 | DPP4 | 1416697_at 1438279_at 1441242_at 1441342_at 1459973_x_at | 19898 | -0.228 | -0.7295 | Yes | ||
| 37 | NFKBIA | 1420088_at 1420089_at 1438157_s_at 1448306_at 1449731_s_at | 19924 | -0.230 | -0.7197 | Yes | ||
| 38 | NKRF | 1434398_at 1441325_at 1445983_at | 19970 | -0.233 | -0.7107 | Yes | ||
| 39 | SPRY2 | 1421656_at 1436584_at | 20076 | -0.244 | -0.7039 | Yes | ||
| 40 | CAP1 | 1417461_at 1417462_at | 20136 | -0.252 | -0.6946 | Yes | ||
| 41 | PLD3 | 1416013_at | 20208 | -0.260 | -0.6855 | Yes | ||
| 42 | SNRPB | 1419260_a_at 1437193_s_at | 20347 | -0.275 | -0.6787 | Yes | ||
| 43 | IL2RG | 1416295_a_at 1416296_at | 20630 | -0.317 | -0.6765 | Yes | ||
| 44 | RPS6KA1 | 1416896_at | 20840 | -0.358 | -0.6690 | Yes | ||
| 45 | IFNGR2 | 1423557_at 1423558_at | 20850 | -0.361 | -0.6522 | Yes | ||
| 46 | INHBA | 1422053_at 1458291_at | 20861 | -0.364 | -0.6354 | Yes | ||
| 47 | ALOX5AP | 1452016_at 1459234_at | 20929 | -0.383 | -0.6202 | Yes | ||
| 48 | SAT1 | 1420502_at | 20941 | -0.385 | -0.6024 | Yes | ||
| 49 | ANXA5 | 1425567_a_at | 20950 | -0.387 | -0.5844 | Yes | ||
| 50 | FLNA | 1426677_at | 20973 | -0.396 | -0.5666 | Yes | ||
| 51 | RHOH | 1429319_at 1441385_at 1443015_at | 20979 | -0.397 | -0.5479 | Yes | ||
| 52 | DUSP2 | 1450698_at | 21013 | -0.410 | -0.5299 | Yes | ||
| 53 | P2RY5 | 1428615_at 1444627_at | 21037 | -0.420 | -0.5110 | Yes | ||
| 54 | PCQAP | 1441617_at 1444680_at 1444688_at 1448435_at | 21085 | -0.435 | -0.4924 | Yes | ||
| 55 | CDC37 | 1416819_at 1416820_at | 21126 | -0.454 | -0.4727 | Yes | ||
| 56 | TRAF1 | 1423602_at 1445452_at | 21524 | -0.759 | -0.4547 | Yes | ||
| 57 | DUSP6 | 1415834_at | 21545 | -0.791 | -0.4180 | Yes | ||
| 58 | PREP | 1422796_at 1436198_at 1439942_at | 21574 | -0.826 | -0.3800 | Yes | ||
| 59 | TAPBP | 1421812_at 1450378_at | 21684 | -1.070 | -0.3341 | Yes | ||
| 60 | VIM | 1438118_x_at 1450641_at 1456292_a_at | 21782 | -1.421 | -0.2709 | Yes | ||
| 61 | EGR2 | 1427682_a_at 1427683_at | 21804 | -1.550 | -0.1981 | Yes | ||
| 62 | TSC22D1 | 1425742_a_at 1433899_x_at 1447360_at 1454758_a_at 1454971_x_at 1456132_x_at | 21928 | -4.288 | 0.0003 | Yes |