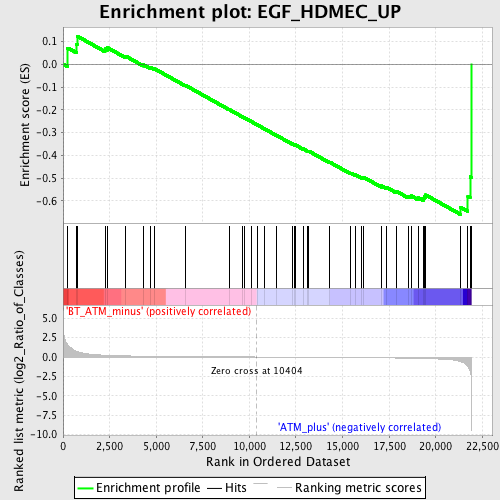
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | EGF_HDMEC_UP |
| Enrichment Score (ES) | -0.6593377 |
| Normalized Enrichment Score (NES) | -1.7979705 |
| Nominal p-value | 0.0053619305 |
| FDR q-value | 0.15207137 |
| FWER p-Value | 0.87 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ITGB1 | 1426918_at 1426919_at 1426920_x_at 1427771_x_at 1438119_at 1452545_a_at | 251 | 1.572 | 0.0716 | No | ||
| 2 | CCND2 | 1416122_at 1416123_at 1416124_at 1430127_a_at 1434745_at 1448229_s_at 1455956_x_at | 722 | 0.724 | 0.0884 | No | ||
| 3 | MDM2 | 1423605_a_at 1427718_a_at 1457929_at | 754 | 0.697 | 0.1238 | No | ||
| 4 | ITGA6 | 1422444_at 1422445_at | 2250 | 0.230 | 0.0676 | No | ||
| 5 | GNAS | 1421740_at 1427789_s_at 1443007_at 1443375_at 1444767_at 1450186_s_at 1453413_at | 2363 | 0.216 | 0.0740 | No | ||
| 6 | ITGA5 | 1423267_s_at 1423268_at 1457561_at 1458996_at | 3362 | 0.142 | 0.0359 | No | ||
| 7 | GRB2 | 1418508_a_at 1449111_a_at | 4312 | 0.105 | -0.0019 | No | ||
| 8 | GSTP1 | 1449575_a_at | 4710 | 0.094 | -0.0151 | No | ||
| 9 | CTGF | 1416953_at | 4924 | 0.089 | -0.0201 | No | ||
| 10 | ITGAV | 1421198_at 1432296_a_at 1432678_at 1432679_at 1452784_at | 6565 | 0.056 | -0.0921 | No | ||
| 11 | CCNG2 | 1416488_at 1448364_at | 8912 | 0.020 | -0.1982 | No | ||
| 12 | GADD45A | 1449519_at | 9649 | 0.010 | -0.2313 | No | ||
| 13 | CASP4 | 1449591_at | 9760 | 0.009 | -0.2358 | No | ||
| 14 | MARCKSL1 | 1415922_s_at 1435415_x_at 1437226_x_at | 10094 | 0.004 | -0.2508 | No | ||
| 15 | NDUFB7 | 1416417_a_at 1448331_at | 10464 | -0.001 | -0.2677 | No | ||
| 16 | CASP3 | 1426165_a_at 1430192_at 1449839_at | 10808 | -0.005 | -0.2830 | No | ||
| 17 | TEK | 1418788_at 1444931_at | 11477 | -0.014 | -0.3128 | No | ||
| 18 | IGFBP2 | 1454159_a_at | 12305 | -0.025 | -0.3493 | No | ||
| 19 | GPX1 | 1460671_at | 12439 | -0.026 | -0.3540 | No | ||
| 20 | PRDX2 | 1418506_a_at 1430979_a_at | 12466 | -0.027 | -0.3537 | No | ||
| 21 | DAD1 | 1418528_a_at 1454860_x_at | 12884 | -0.032 | -0.3711 | No | ||
| 22 | CXCL2 | 1449984_at | 13143 | -0.036 | -0.3810 | No | ||
| 23 | MAP3K11 | 1450669_at | 13197 | -0.037 | -0.3814 | No | ||
| 24 | SKI | 1426373_at 1429192_at 1444785_at 1456547_at | 14315 | -0.054 | -0.4296 | No | ||
| 25 | RHOA | 1437628_s_at | 15444 | -0.073 | -0.4773 | No | ||
| 26 | HSPB1 | 1422943_a_at 1425964_x_at 1427853_a_at 1427854_x_at 1453914_at | 15708 | -0.077 | -0.4852 | No | ||
| 27 | RPL6 | 1416546_a_at | 16023 | -0.083 | -0.4952 | No | ||
| 28 | DDIT3 | 1417516_at 1443897_at | 16124 | -0.085 | -0.4953 | No | ||
| 29 | TIE1 | 1416238_at | 17087 | -0.107 | -0.5335 | No | ||
| 30 | HSP90AA1 | 1426645_at 1437497_a_at 1438902_a_at 1455078_at | 17352 | -0.114 | -0.5396 | No | ||
| 31 | RELA | 1419536_a_at | 17908 | -0.130 | -0.5581 | No | ||
| 32 | RAD23B | 1424222_s_at 1450903_at 1455420_at 1456822_at | 18541 | -0.151 | -0.5790 | No | ||
| 33 | CDH2 | 1418815_at 1439307_at 1449244_at | 18688 | -0.156 | -0.5774 | No | ||
| 34 | GSTM1 | 1416416_x_at 1448330_at | 19075 | -0.173 | -0.5859 | No | ||
| 35 | CCND1 | 1417419_at 1417420_at 1448698_at | 19337 | -0.187 | -0.5879 | No | ||
| 36 | PLAUR | 1452521_a_at | 19389 | -0.190 | -0.5802 | No | ||
| 37 | DYNLL1 | 1448682_at | 19471 | -0.195 | -0.5736 | No | ||
| 38 | AXL | 1423586_at 1438621_x_at | 21349 | -0.587 | -0.6283 | Yes | ||
| 39 | ITGB3 | 1421511_at 1455257_at | 21738 | -1.266 | -0.5792 | Yes | ||
| 40 | TCEB1 | 1427914_a_at 1427915_s_at 1436226_at 1452593_a_at | 21856 | -1.724 | -0.4934 | Yes | ||
| 41 | TMSB10 | 1417219_s_at 1436682_at 1436902_x_at 1437185_s_at 1455946_x_at | 21934 | -9.405 | -0.0000 | Yes |

