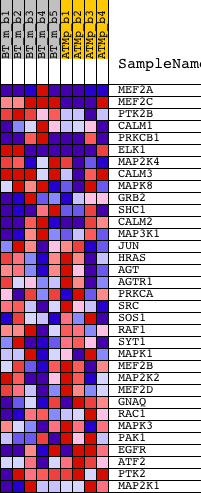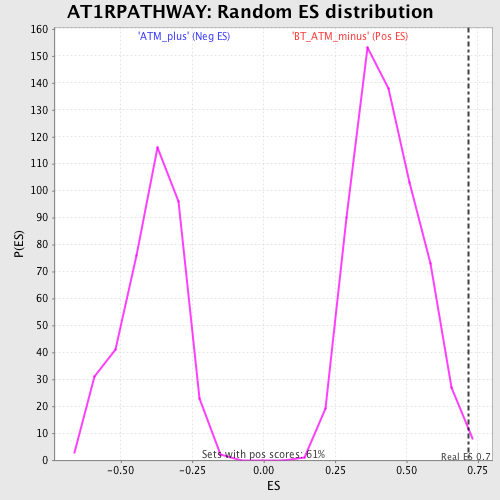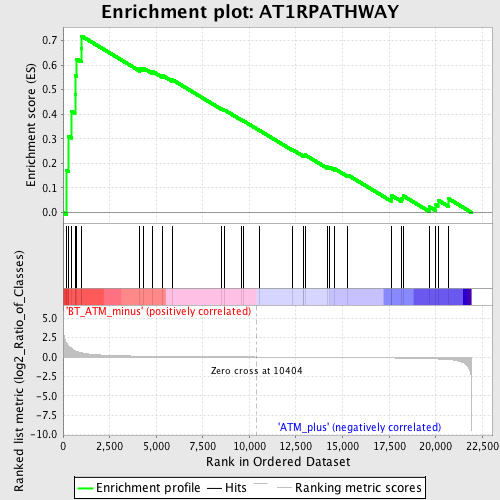
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus.phenotype_BT_ATM_minus_versus_ATM_plus.cls #BT_ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_BT_ATM_minus_versus_ATM_plus.cls#BT_ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | BT_ATM_minus |
| GeneSet | AT1RPATHWAY |
| Enrichment Score (ES) | 0.71820354 |
| Normalized Enrichment Score (NES) | 1.6653539 |
| Nominal p-value | 0.001633987 |
| FDR q-value | 0.39344546 |
| FWER p-Value | 0.993 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | MEF2A | 1421252_a_at 1425426_a_at 1427185_at 1427186_a_at 1452347_at | 173 | 1.840 | 0.1712 | Yes | ||
| 2 | MEF2C | 1421027_a_at 1421028_a_at 1424852_at 1439946_at 1445420_at 1446484_at 1451506_at 1451507_at 1458551_at | 284 | 1.486 | 0.3109 | Yes | ||
| 3 | PTK2B | 1434653_at 1442437_at 1442927_at | 453 | 1.115 | 0.4118 | Yes | ||
| 4 | CALM1 | 1417365_a_at 1417366_s_at 1433592_at 1454611_a_at 1455571_x_at | 639 | 0.812 | 0.4824 | Yes | ||
| 5 | PRKCB1 | 1423478_at 1438981_at 1443144_at 1459674_at 1460419_a_at | 662 | 0.786 | 0.5579 | Yes | ||
| 6 | ELK1 | 1421896_at 1421897_at 1446390_at | 729 | 0.717 | 0.6247 | Yes | ||
| 7 | MAP2K4 | 1426233_at 1442679_at 1451982_at | 982 | 0.548 | 0.6665 | Yes | ||
| 8 | CALM3 | 1426710_at 1438825_at 1438826_x_at 1450864_at | 998 | 0.538 | 0.7182 | Yes | ||
| 9 | MAPK8 | 1420931_at 1420932_at 1437045_at 1440856_at 1457936_at | 4125 | 0.111 | 0.5863 | No | ||
| 10 | GRB2 | 1418508_a_at 1449111_a_at | 4312 | 0.105 | 0.5880 | No | ||
| 11 | SHC1 | 1422853_at 1422854_at | 4792 | 0.092 | 0.5751 | No | ||
| 12 | CALM2 | 1422414_a_at 1423807_a_at | 5336 | 0.080 | 0.5581 | No | ||
| 13 | MAP3K1 | 1424850_at | 5859 | 0.069 | 0.5409 | No | ||
| 14 | JUN | 1417409_at 1448694_at | 8513 | 0.025 | 0.4223 | No | ||
| 15 | HRAS | 1422407_s_at 1424132_at | 8692 | 0.023 | 0.4164 | No | ||
| 16 | AGT | 1423396_at | 9556 | 0.011 | 0.3781 | No | ||
| 17 | AGTR1 | 1436739_at 1446527_at | 9689 | 0.010 | 0.3730 | No | ||
| 18 | PRKCA | 1427562_a_at 1443069_at 1445028_at 1445395_at 1446598_at 1450945_at | 10534 | -0.002 | 0.3346 | No | ||
| 19 | SRC | 1423240_at 1450918_s_at | 12329 | -0.025 | 0.2551 | No | ||
| 20 | SOS1 | 1421884_at 1421885_at 1421886_at | 12908 | -0.033 | 0.2319 | No | ||
| 21 | RAF1 | 1416078_s_at 1425419_a_at | 12932 | -0.033 | 0.2341 | No | ||
| 22 | SYT1 | 1421990_at 1431191_a_at 1433884_at 1438282_at 1442557_at 1457497_at | 12993 | -0.034 | 0.2347 | No | ||
| 23 | MAPK1 | 1419568_at 1426585_s_at 1442876_at 1453104_at | 14178 | -0.052 | 0.1856 | No | ||
| 24 | MEF2B | 1421541_a_at | 14313 | -0.054 | 0.1848 | No | ||
| 25 | MAP2K2 | 1415974_at 1443436_at 1460636_at | 14594 | -0.058 | 0.1777 | No | ||
| 26 | MEF2D | 1421388_at 1434487_at 1436642_x_at 1437300_at 1458901_at | 15295 | -0.070 | 0.1525 | No | ||
| 27 | GNAQ | 1428938_at 1428939_s_at 1428940_at 1429559_at 1446688_at 1447593_x_at 1450115_at 1455729_at 1458159_at | 17611 | -0.121 | 0.0586 | No | ||
| 28 | RAC1 | 1423734_at 1437674_at 1451086_s_at | 17638 | -0.122 | 0.0693 | No | ||
| 29 | MAPK3 | 1427060_at | 18157 | -0.138 | 0.0590 | No | ||
| 30 | PAK1 | 1420979_at 1420980_at 1450070_s_at 1459029_at | 18252 | -0.140 | 0.0684 | No | ||
| 31 | EGFR | 1424932_at 1432647_at 1435888_at 1451530_at 1454313_at 1457563_at 1460420_a_at | 19653 | -0.207 | 0.0246 | No | ||
| 32 | ATF2 | 1426582_at 1426583_at 1427559_a_at 1437777_at 1452116_s_at 1459237_at | 20021 | -0.238 | 0.0310 | No | ||
| 33 | PTK2 | 1423059_at 1430827_a_at 1439198_at 1440082_at 1441475_at 1443384_at 1445137_at | 20132 | -0.251 | 0.0505 | No | ||
| 34 | MAP2K1 | 1416351_at | 20684 | -0.326 | 0.0571 | No |