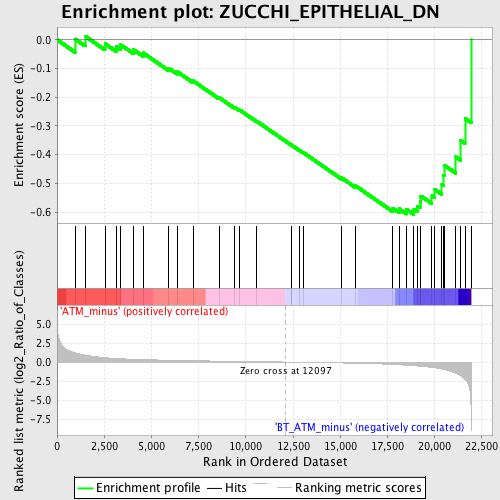
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_BT_ATM_minus.phenotype_ATM_minus_versus_BT_ATM_minus.cls #ATM_minus_versus_BT_ATM_minus.phenotype_ATM_minus_versus_BT_ATM_minus.cls #ATM_minus_versus_BT_ATM_minus_repos |
| Phenotype | phenotype_ATM_minus_versus_BT_ATM_minus.cls#ATM_minus_versus_BT_ATM_minus_repos |
| Upregulated in class | BT_ATM_minus |
| GeneSet | ZUCCHI_EPITHELIAL_DN |
| Enrichment Score (ES) | -0.60825425 |
| Normalized Enrichment Score (NES) | -1.63905 |
| Nominal p-value | 0.009041592 |
| FDR q-value | 0.35805035 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PDCD5 | 1450638_at | 950 | 1.216 | 0.0028 | No | ||
| 2 | HMGN1 | 1422495_a_at 1422496_at 1438940_x_at 1455897_x_at | 1483 | 0.903 | 0.0129 | No | ||
| 3 | ANXA1 | 1448213_at | 2538 | 0.571 | -0.0136 | No | ||
| 4 | EIF4A1 | 1427058_at 1430980_a_at 1434985_a_at | 3158 | 0.462 | -0.0243 | No | ||
| 5 | LGALS3 | 1426808_at 1445626_at | 3374 | 0.432 | -0.0177 | No | ||
| 6 | PDLIM1 | 1416554_at | 4053 | 0.361 | -0.0349 | No | ||
| 7 | SEMA3B | 1416644_a_at 1431795_a_at 1448415_a_at | 4558 | 0.320 | -0.0458 | No | ||
| 8 | EIF3S4 | 1417718_at | 5914 | 0.234 | -0.0988 | No | ||
| 9 | CXCL2 | 1449984_at | 6363 | 0.212 | -0.1112 | No | ||
| 10 | SEC61B | 1417083_at | 7200 | 0.173 | -0.1428 | No | ||
| 11 | PPM1G | 1416792_at | 8580 | 0.119 | -0.2012 | No | ||
| 12 | RASD1 | 1423619_at | 9400 | 0.090 | -0.2352 | No | ||
| 13 | IL6 | 1450297_at | 9669 | 0.082 | -0.2443 | No | ||
| 14 | LIF | 1421206_at 1421207_at 1450160_at | 10555 | 0.052 | -0.2828 | No | ||
| 15 | LMNA | 1421654_a_at 1425472_a_at 1426868_x_at 1452212_at 1457670_s_at | 12390 | -0.011 | -0.3661 | No | ||
| 16 | TM4SF1 | 1439925_at 1448069_at 1450958_at | 12391 | -0.011 | -0.3657 | No | ||
| 17 | YWHAE | 1426384_a_at 1426385_x_at 1435702_s_at 1438839_a_at 1440841_at 1457173_at | 12851 | -0.028 | -0.3855 | No | ||
| 18 | TACSTD2 | 1423323_at | 13038 | -0.035 | -0.3927 | No | ||
| 19 | CCL20 | 1422029_at | 15058 | -0.119 | -0.4804 | No | ||
| 20 | AREG | 1421134_at | 15803 | -0.158 | -0.5083 | No | ||
| 21 | DDB1 | 1415735_at | 17772 | -0.312 | -0.5863 | No | ||
| 22 | CXCL1 | 1419209_at 1441855_x_at 1457644_s_at | 18118 | -0.354 | -0.5886 | No | ||
| 23 | PFN1 | 1449018_at | 18512 | -0.415 | -0.5908 | Yes | ||
| 24 | PICALM | 1441328_at 1443218_at 1446968_at 1451316_a_at 1455773_at | 18895 | -0.486 | -0.5898 | Yes | ||
| 25 | BAP1 | 1426354_at 1452039_a_at | 19100 | -0.527 | -0.5791 | Yes | ||
| 26 | CST3 | 1426195_a_at | 19222 | -0.555 | -0.5635 | Yes | ||
| 27 | ACTG1 | 1415779_s_at | 19262 | -0.562 | -0.5439 | Yes | ||
| 28 | STRN4 | 1426209_at | 19853 | -0.713 | -0.5437 | Yes | ||
| 29 | DDX5 | 1419653_a_at 1423645_a_at 1433809_at 1433810_x_at 1438371_x_at 1454793_x_at | 19988 | -0.757 | -0.5211 | Yes | ||
| 30 | ZYX | 1417240_at 1438552_x_at | 20379 | -0.916 | -0.5041 | Yes | ||
| 31 | SOD2 | 1417193_at 1417194_at 1442994_at 1444531_at 1448610_a_at 1454976_at | 20453 | -0.952 | -0.4712 | Yes | ||
| 32 | GADD45B | 1420197_at 1449773_s_at 1450971_at | 20505 | -0.977 | -0.4364 | Yes | ||
| 33 | CLDN4 | 1418283_at | 21122 | -1.491 | -0.4079 | Yes | ||
| 34 | JUNB | 1415899_at | 21378 | -1.811 | -0.3507 | Yes | ||
| 35 | ST13 | 1416851_at 1442775_at 1460193_at | 21603 | -2.284 | -0.2742 | Yes | ||
| 36 | PNRC1 | 1433668_at 1438524_x_at 1439513_at | 21933 | -7.611 | 0.0000 | Yes |

