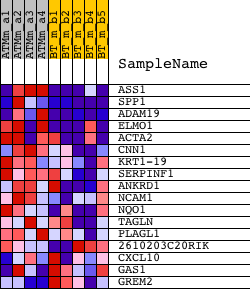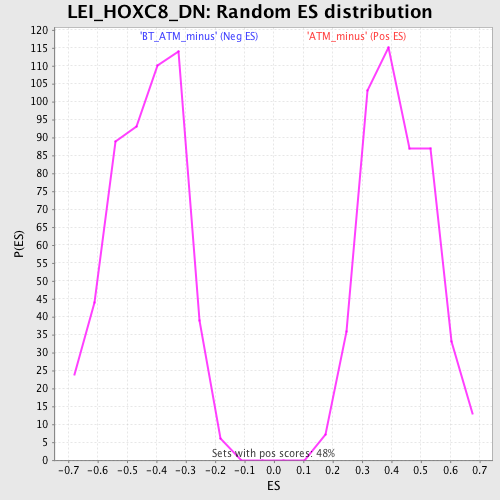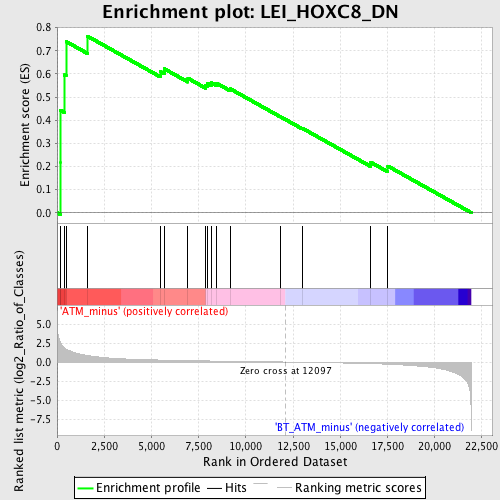
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_BT_ATM_minus.phenotype_ATM_minus_versus_BT_ATM_minus.cls #ATM_minus_versus_BT_ATM_minus.phenotype_ATM_minus_versus_BT_ATM_minus.cls #ATM_minus_versus_BT_ATM_minus_repos |
| Phenotype | phenotype_ATM_minus_versus_BT_ATM_minus.cls#ATM_minus_versus_BT_ATM_minus_repos |
| Upregulated in class | ATM_minus |
| GeneSet | LEI_HOXC8_DN |
| Enrichment Score (ES) | 0.7634083 |
| Normalized Enrichment Score (NES) | 1.8011585 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.07694372 |
| FWER p-Value | 0.53 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ASS1 | 1416239_at 1459937_at | 167 | 2.594 | 0.2192 | Yes | ||
| 2 | SPP1 | 1449254_at | 170 | 2.570 | 0.4438 | Yes | ||
| 3 | ADAM19 | 1418402_at 1418403_at | 379 | 1.845 | 0.5956 | Yes | ||
| 4 | ELMO1 | 1424523_at 1446610_at 1450208_a_at | 478 | 1.702 | 0.7399 | Yes | ||
| 5 | ACTA2 | 1416454_s_at 1444105_at 1456658_at | 1602 | 0.855 | 0.7634 | Yes | ||
| 6 | CNN1 | 1417917_at | 5471 | 0.261 | 0.6098 | No | ||
| 7 | KRT1-19 | 1417156_at | 5680 | 0.247 | 0.6219 | No | ||
| 8 | SERPINF1 | 1416168_at 1453724_a_at | 6927 | 0.185 | 0.5812 | No | ||
| 9 | ANKRD1 | 1420991_at 1420992_at | 7863 | 0.145 | 0.5513 | No | ||
| 10 | NCAM1 | 1421966_at 1425126_at 1426864_a_at 1426865_a_at 1439556_at 1440067_at 1441995_at 1442680_at 1443018_at 1450437_a_at 1450438_at 1454140_at | 7949 | 0.142 | 0.5598 | No | ||
| 11 | NQO1 | 1423627_at | 8158 | 0.134 | 0.5621 | No | ||
| 12 | TAGLN | 1423505_at | 8436 | 0.124 | 0.5603 | No | ||
| 13 | PLAGL1 | 1426208_x_at 1438588_at 1450533_a_at | 9175 | 0.098 | 0.5352 | No | ||
| 14 | 2610203C20RIK | 1427437_at 1442373_at 1447047_at 1455160_at | 11827 | 0.009 | 0.4151 | No | ||
| 15 | CXCL10 | 1418930_at | 12970 | -0.033 | 0.3659 | No | ||
| 16 | GAS1 | 1416855_at 1448494_at 1456322_at | 16591 | -0.208 | 0.2189 | No | ||
| 17 | GREM2 | 1418492_at | 17516 | -0.284 | 0.2016 | No |