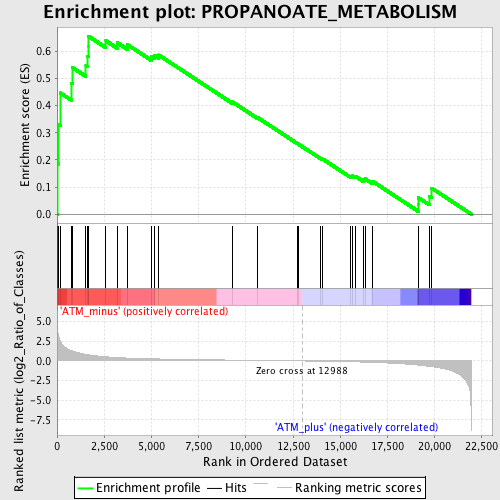
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_02_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_minus |
| GeneSet | PROPANOATE_METABOLISM |
| Enrichment Score (ES) | 0.6558419 |
| Normalized Enrichment Score (NES) | 1.7569009 |
| Nominal p-value | 0.0045454544 |
| FDR q-value | 0.076593824 |
| FWER p-Value | 0.731 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ACADL | 1448987_at 1448988_at | 25 | 3.939 | 0.1875 | Yes | ||
| 2 | ALDH1B1 | 1451260_at | 89 | 3.024 | 0.3294 | Yes | ||
| 3 | SUCLG2 | 1427441_a_at 1435841_s_at 1444309_at | 169 | 2.521 | 0.4465 | Yes | ||
| 4 | MUT | 1416838_at 1416839_at 1448486_at | 761 | 1.301 | 0.4818 | Yes | ||
| 5 | ACAT1 | 1421558_at 1424182_at 1424183_at 1451271_a_at | 805 | 1.260 | 0.5401 | Yes | ||
| 6 | LDHA | 1419737_a_at | 1527 | 0.839 | 0.5473 | Yes | ||
| 7 | ALDH9A1 | 1436689_a_at 1437398_a_at | 1628 | 0.798 | 0.5810 | Yes | ||
| 8 | ECHS1 | 1452341_at | 1648 | 0.793 | 0.6181 | Yes | ||
| 9 | PCCB | 1450969_at | 1651 | 0.791 | 0.6558 | Yes | ||
| 10 | SUCLG1 | 1415891_at | 2584 | 0.528 | 0.6386 | No | ||
| 11 | ACACA | 1444810_at | 3182 | 0.424 | 0.6316 | No | ||
| 12 | LDHC | 1415846_a_at 1415847_at | 3716 | 0.363 | 0.6247 | No | ||
| 13 | PCCA | 1447577_x_at 1451405_at | 4981 | 0.271 | 0.5799 | No | ||
| 14 | ALDH1A2 | 1422789_at | 5144 | 0.262 | 0.5851 | No | ||
| 15 | ALDH3A1 | 1418752_at | 5380 | 0.250 | 0.5863 | No | ||
| 16 | SUCLA2 | 1430402_at 1452206_at | 9272 | 0.109 | 0.4139 | No | ||
| 17 | ACAT2 | 1435630_s_at | 10596 | 0.071 | 0.3569 | No | ||
| 18 | MLYCD | 1449964_a_at | 12719 | 0.009 | 0.2605 | No | ||
| 19 | SDS | 1424744_at | 12776 | 0.007 | 0.2583 | No | ||
| 20 | ALDH1A1 | 1416468_at 1445980_at | 13959 | -0.034 | 0.2059 | No | ||
| 21 | ALDH2 | 1434987_at 1434988_x_at 1437410_at 1448143_at | 14079 | -0.039 | 0.2024 | No | ||
| 22 | ACADSB | 1455446_x_at | 15551 | -0.102 | 0.1401 | No | ||
| 23 | ALDH1A3 | 1417642_at 1427395_a_at 1448789_at | 15647 | -0.107 | 0.1409 | No | ||
| 24 | ALDH3A2 | 1415776_at | 15805 | -0.116 | 0.1393 | No | ||
| 25 | LDHB | 1416183_a_at 1434499_a_at 1442618_at 1448237_x_at 1455235_x_at | 16211 | -0.142 | 0.1276 | No | ||
| 26 | ACADM | 1415984_at | 16311 | -0.149 | 0.1303 | No | ||
| 27 | ABAT | 1433855_at | 16726 | -0.181 | 0.1200 | No | ||
| 28 | HADHA | 1452173_at | 19127 | -0.527 | 0.0357 | No | ||
| 29 | ALDH6A1 | 1448104_at | 19132 | -0.529 | 0.0609 | No | ||
| 30 | EHHADH | 1448382_at | 19720 | -0.681 | 0.0666 | No | ||
| 31 | MCEE | 1438477_a_at | 19833 | -0.718 | 0.0959 | No |

