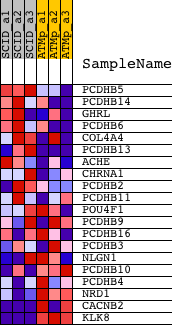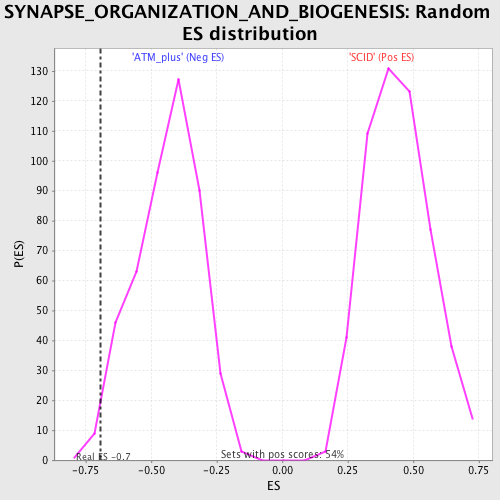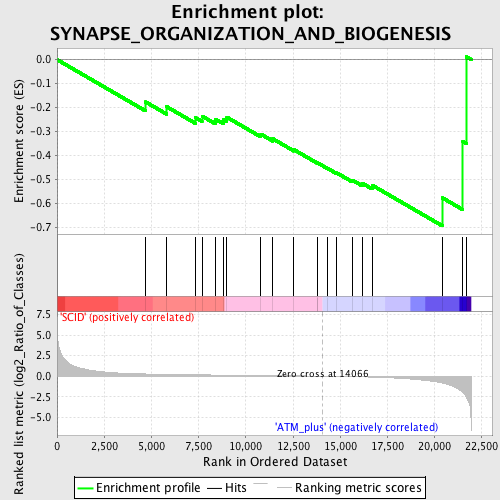
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus_repos |
| Phenotype | phenotype_SCID_versus_ATM_plus.cls#SCID_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | SYNAPSE_ORGANIZATION_AND_BIOGENESIS |
| Enrichment Score (ES) | -0.69325626 |
| Normalized Enrichment Score (NES) | -1.5905764 |
| Nominal p-value | 0.012931035 |
| FDR q-value | 0.3841201 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PCDHB5 | 1450263_at | 4687 | 0.267 | -0.1762 | No | ||
| 2 | PCDHB14 | 1421437_x_at | 5806 | 0.220 | -0.1962 | No | ||
| 3 | GHRL | 1448980_at | 7332 | 0.169 | -0.2420 | No | ||
| 4 | PCDHB6 | 1423014_at | 7677 | 0.158 | -0.2354 | No | ||
| 5 | COL4A4 | 1425772_at 1440250_at 1445328_at | 8395 | 0.139 | -0.2484 | No | ||
| 6 | PCDHB13 | 1450316_at | 8820 | 0.129 | -0.2496 | No | ||
| 7 | ACHE | 1422635_at | 8990 | 0.124 | -0.2398 | No | ||
| 8 | CHRNA1 | 1418852_at | 10792 | 0.081 | -0.3105 | No | ||
| 9 | PCDHB2 | 1421548_at | 11426 | 0.067 | -0.3299 | No | ||
| 10 | PCDHB11 | 1450270_at 1456710_at | 12544 | 0.040 | -0.3752 | No | ||
| 11 | POU4F1 | 1429667_at 1429668_at | 13781 | 0.008 | -0.4305 | No | ||
| 12 | PCDHB9 | 1422640_at | 13786 | 0.008 | -0.4297 | No | ||
| 13 | PCDHB16 | 1441358_at 1441994_at 1450216_at | 14305 | -0.007 | -0.4523 | No | ||
| 14 | PCDHB3 | 1420429_at | 14810 | -0.023 | -0.4720 | No | ||
| 15 | NLGN1 | 1421648_at 1437160_at | 15640 | -0.054 | -0.5023 | No | ||
| 16 | PCDHB10 | 1422729_at | 16162 | -0.078 | -0.5150 | No | ||
| 17 | PCDHB4 | 1422951_at 1440632_at | 16689 | -0.111 | -0.5234 | No | ||
| 18 | NRD1 | 1424391_at 1449716_s_at | 20412 | -0.829 | -0.5763 | Yes | ||
| 19 | CACNB2 | 1441521_at 1444693_at 1446632_at 1452476_at 1456401_at | 21490 | -2.012 | -0.3415 | Yes | ||
| 20 | KLK8 | 1419722_at | 21678 | -2.565 | 0.0118 | Yes |