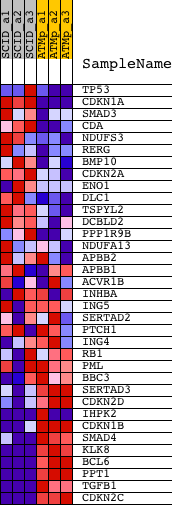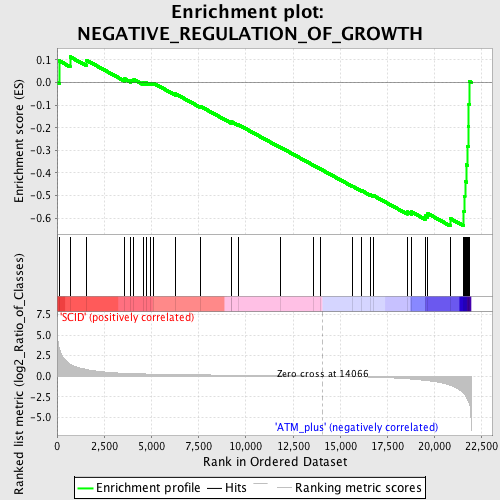
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus_repos |
| Phenotype | phenotype_SCID_versus_ATM_plus.cls#SCID_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | NEGATIVE_REGULATION_OF_GROWTH |
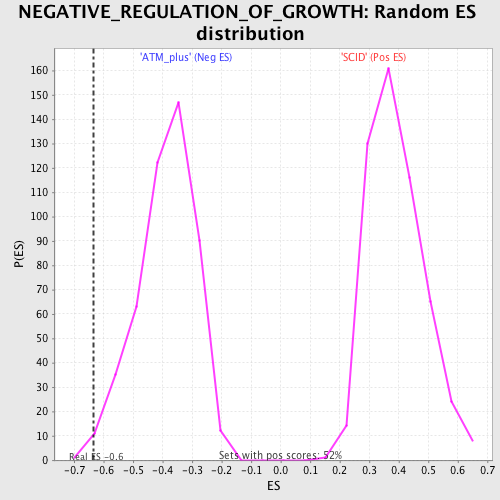
| Enrichment Score (ES) | -0.63474226 |
| Normalized Enrichment Score (NES) | -1.6404217 |
| Nominal p-value | 0.008316008 |
| FDR q-value | 0.28837267 |
| FWER p-Value | 0.985 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | TP53 | 1426538_a_at 1427739_a_at 1438808_at 1457623_x_at 1459780_at 1459781_x_at | 111 | 3.368 | 0.0961 | No | ||
| 2 | CDKN1A | 1421679_a_at 1424638_at | 689 | 1.463 | 0.1137 | No | ||
| 3 | SMAD3 | 1450471_at 1450472_s_at 1454960_at | 1555 | 0.815 | 0.0987 | No | ||
| 4 | CDA | 1427357_at | 3588 | 0.343 | 0.0162 | No | ||
| 5 | NDUFS3 | 1423737_at | 3907 | 0.316 | 0.0112 | No | ||
| 6 | RERG | 1447446_at 1451236_at | 4067 | 0.303 | 0.0130 | No | ||
| 7 | BMP10 | 1421763_at | 4565 | 0.273 | -0.0015 | No | ||
| 8 | CDKN2A | 1450140_a_at | 4728 | 0.265 | -0.0010 | No | ||
| 9 | ENO1 | 1419023_x_at | 4940 | 0.255 | -0.0029 | No | ||
| 10 | DLC1 | 1436173_at 1450206_at 1458220_at 1460602_at | 5090 | 0.249 | -0.0022 | No | ||
| 11 | TSPYL2 | 1434849_at | 6274 | 0.202 | -0.0502 | No | ||
| 12 | DCBLD2 | 1420526_at 1437635_at 1449891_a_at | 7588 | 0.161 | -0.1053 | No | ||
| 13 | PPP1R9B | 1447247_at 1447486_at | 9214 | 0.119 | -0.1759 | No | ||
| 14 | NDUFA13 | 1430713_s_at | 9251 | 0.118 | -0.1740 | No | ||
| 15 | APBB2 | 1426719_at 1426720_at 1440135_at 1446481_at 1452342_at | 9600 | 0.109 | -0.1866 | No | ||
| 16 | APBB1 | 1423892_at 1423893_x_at | 11855 | 0.056 | -0.2879 | No | ||
| 17 | ACVR1B | 1422098_at 1433725_at | 13556 | 0.014 | -0.3651 | No | ||
| 18 | INHBA | 1422053_at 1458291_at | 13937 | 0.004 | -0.3823 | No | ||
| 19 | ING5 | 1431134_at 1436069_at 1442094_at 1454235_a_at | 15634 | -0.053 | -0.4581 | No | ||
| 20 | SERTAD2 | 1417209_at 1444645_at 1454815_at | 16129 | -0.077 | -0.4784 | No | ||
| 21 | PTCH1 | 1428853_at 1439663_at 1447039_at 1450824_at | 16592 | -0.105 | -0.4964 | No | ||
| 22 | ING4 | 1448054_at 1460728_s_at | 16734 | -0.114 | -0.4994 | No | ||
| 23 | RB1 | 1417850_at 1441494_at 1444400_at 1444454_at 1457447_at | 18543 | -0.312 | -0.5725 | No | ||
| 24 | PML | 1448757_at 1456103_at 1459137_at | 18760 | -0.347 | -0.5720 | No | ||
| 25 | BBC3 | 1423315_at | 19494 | -0.509 | -0.5902 | No | ||
| 26 | SERTAD3 | 1421076_at 1421077_at | 19635 | -0.547 | -0.5801 | No | ||
| 27 | CDKN2D | 1416253_at | 20821 | -1.081 | -0.6018 | Yes | ||
| 28 | IHPK2 | 1428373_at 1435319_at | 21544 | -2.129 | -0.5708 | Yes | ||
| 29 | CDKN1B | 1419497_at 1434045_at | 21588 | -2.239 | -0.5055 | Yes | ||
| 30 | SMAD4 | 1422485_at 1422486_a_at 1422487_at 1444205_at | 21604 | -2.282 | -0.4377 | Yes | ||
| 31 | KLK8 | 1419722_at | 21678 | -2.565 | -0.3640 | Yes | ||
| 32 | BCL6 | 1421818_at 1450381_a_at | 21747 | -2.870 | -0.2809 | Yes | ||
| 33 | PPT1 | 1420015_s_at 1420016_at 1422467_at 1422468_at 1444884_at | 21776 | -2.991 | -0.1923 | Yes | ||
| 34 | TGFB1 | 1420653_at 1445360_at | 21809 | -3.227 | -0.0969 | Yes | ||
| 35 | CDKN2C | 1416868_at 1439164_at | 21847 | -3.418 | 0.0041 | Yes |