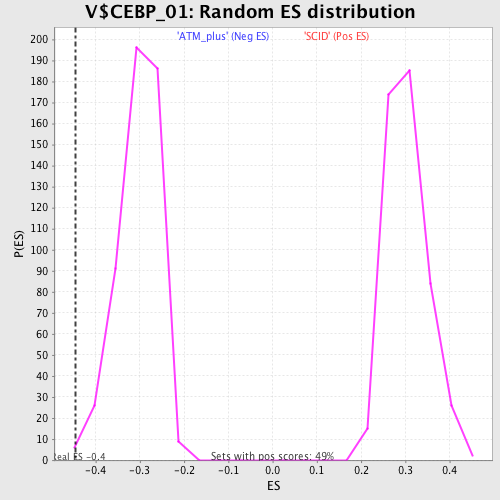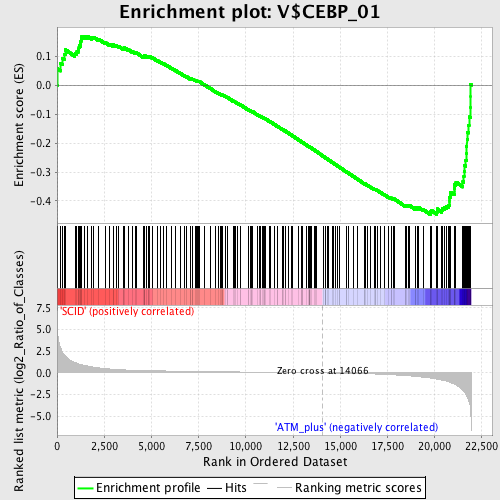
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus_repos |
| Phenotype | phenotype_SCID_versus_ATM_plus.cls#SCID_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | V$CEBP_01 |
| Enrichment Score (ES) | -0.4465447 |
| Normalized Enrichment Score (NES) | -1.4767832 |
| Nominal p-value | 0.0019455253 |
| FDR q-value | 0.2108593 |
| FWER p-Value | 0.932 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PSMA1 | 1415695_at 1436769_at 1436770_x_at 1458292_at | 8 | 6.245 | 0.0567 | No | ||
| 2 | BCL11B | 1440137_at 1441762_at 1450339_a_at | 186 | 2.915 | 0.0752 | No | ||
| 3 | CSK | 1423518_at 1439744_at | 289 | 2.344 | 0.0919 | No | ||
| 4 | DMD | 1417307_at 1430320_at 1446156_at 1448665_at 1457022_at | 376 | 2.102 | 0.1072 | No | ||
| 5 | SUPT16H | 1419741_at 1449578_at 1456449_at | 449 | 1.937 | 0.1216 | No | ||
| 6 | UTX | 1427235_at 1427672_a_at 1440524_at 1441201_at 1445198_at 1446234_at 1446278_at | 947 | 1.192 | 0.1096 | No | ||
| 7 | PPP1CB | 1426205_at 1431328_at 1433540_x_at 1456462_x_at | 1050 | 1.104 | 0.1150 | No | ||
| 8 | EVI2A | 1443894_at 1450241_a_at 1458788_at 1460055_at | 1110 | 1.055 | 0.1220 | No | ||
| 9 | STOML2 | 1448774_at | 1157 | 1.019 | 0.1292 | No | ||
| 10 | ZNF462 | 1439106_at 1456789_at | 1172 | 1.009 | 0.1377 | No | ||
| 11 | SYNE2 | 1427982_s_at 1438585_at 1442285_at 1447159_at 1459322_at | 1249 | 0.965 | 0.1431 | No | ||
| 12 | CS | 1422577_at 1422578_at 1450667_a_at | 1256 | 0.961 | 0.1516 | No | ||
| 13 | PCDH8 | 1417051_at 1432491_at 1447825_x_at | 1269 | 0.954 | 0.1597 | No | ||
| 14 | EYA1 | 1421727_at 1443117_at 1457424_at | 1310 | 0.929 | 0.1664 | No | ||
| 15 | TNRC15 | 1439837_at 1446275_at 1451856_at 1457472_at | 1428 | 0.873 | 0.1690 | No | ||
| 16 | FABP1 | 1417556_at 1448764_a_at | 1591 | 0.796 | 0.1688 | No | ||
| 17 | KPNA3 | 1421828_at 1445527_at 1446538_at 1450385_at 1450386_at | 1831 | 0.695 | 0.1642 | No | ||
| 18 | NPR3 | 1435184_at 1448024_at 1450286_at | 1929 | 0.659 | 0.1657 | No | ||
| 19 | RBMS1 | 1418703_at 1434005_at 1441437_at 1458578_at 1459968_at | 2213 | 0.577 | 0.1580 | No | ||
| 20 | TMEM10 | 1435854_at 1447487_at | 2548 | 0.499 | 0.1472 | No | ||
| 21 | NUP155 | 1418727_at 1449200_at | 2793 | 0.453 | 0.1401 | No | ||
| 22 | PVRL1 | 1438111_at 1438421_at 1440131_at 1446488_at 1450819_at | 2967 | 0.425 | 0.1360 | No | ||
| 23 | SEPHS2 | 1418325_at 1449040_a_at | 2985 | 0.423 | 0.1391 | No | ||
| 24 | DNM3 | 1425403_at 1436624_at 1436875_at 1438801_at 1445540_at 1446265_at 1446431_at 1446694_at 1457405_at | 3145 | 0.396 | 0.1354 | No | ||
| 25 | IGSF4 | 1417376_a_at 1417377_at 1417378_at 1431611_a_at | 3240 | 0.383 | 0.1346 | No | ||
| 26 | RALY | 1418573_a_at | 3532 | 0.348 | 0.1244 | No | ||
| 27 | FOXD3 | 1422210_at | 3541 | 0.347 | 0.1272 | No | ||
| 28 | SPA17 | 1417635_at | 3569 | 0.344 | 0.1291 | No | ||
| 29 | PLXNC1 | 1423213_at 1423214_at 1443433_at 1443434_s_at 1450905_at 1450906_at | 3804 | 0.324 | 0.1213 | No | ||
| 30 | PRDM10 | 1441764_at 1444812_at | 3974 | 0.310 | 0.1163 | No | ||
| 31 | GPX1 | 1460671_at | 4159 | 0.297 | 0.1106 | No | ||
| 32 | COL2A1 | 1450567_a_at | 4182 | 0.296 | 0.1123 | No | ||
| 33 | PFKFB1 | 1427213_at | 4583 | 0.272 | 0.0964 | No | ||
| 34 | KCNJ1 | 1418613_at 1418614_at | 4611 | 0.271 | 0.0976 | No | ||
| 35 | SLITRK3 | 1439250_at | 4623 | 0.270 | 0.0996 | No | ||
| 36 | ITGB1BP2 | 1423238_at | 4624 | 0.270 | 0.1020 | No | ||
| 37 | CALN1 | 1425846_a_at | 4738 | 0.265 | 0.0993 | No | ||
| 38 | WWP2 | 1438482_at 1442162_at 1448145_at 1448146_at 1456714_at 1457499_at | 4832 | 0.260 | 0.0974 | No | ||
| 39 | MAP4K4 | 1422615_at 1440609_at 1448050_s_at 1459912_at | 4839 | 0.259 | 0.0995 | No | ||
| 40 | PRDM1 | 1420425_at 1444997_at | 4910 | 0.257 | 0.0986 | No | ||
| 41 | CLDN2 | 1417231_at 1417232_at | 5049 | 0.251 | 0.0945 | No | ||
| 42 | POU3F4 | 1422164_at 1422165_at | 5317 | 0.239 | 0.0844 | No | ||
| 43 | CD86 | 1420404_at 1449858_at | 5473 | 0.232 | 0.0794 | No | ||
| 44 | POU3F3 | 1422331_at 1435197_at | 5623 | 0.226 | 0.0746 | No | ||
| 45 | IL1RAPL1 | 1445821_at 1458964_at | 5785 | 0.221 | 0.0692 | No | ||
| 46 | CSAD | 1427981_a_at | 6082 | 0.209 | 0.0575 | No | ||
| 47 | OSR2 | 1426155_a_at 1430950_at | 6279 | 0.202 | 0.0504 | No | ||
| 48 | HOXA11 | 1420414_at | 6511 | 0.195 | 0.0415 | No | ||
| 49 | SV2B | 1434800_at 1435687_at | 6743 | 0.187 | 0.0326 | No | ||
| 50 | TITF1 | 1422346_at | 6856 | 0.183 | 0.0291 | No | ||
| 51 | UBXD3 | 1426007_a_at 1451906_at 1456385_x_at 1457581_at | 7058 | 0.177 | 0.0215 | No | ||
| 52 | SFRS6 | 1416720_at 1416721_s_at 1447898_s_at 1448454_at | 7077 | 0.176 | 0.0223 | No | ||
| 53 | CDH13 | 1423551_at 1431824_at 1454015_a_at 1459535_at | 7175 | 0.173 | 0.0194 | No | ||
| 54 | RNF19 | 1417508_at 1417509_at | 7184 | 0.173 | 0.0206 | No | ||
| 55 | SSTR5 | 1445459_at | 7323 | 0.169 | 0.0158 | No | ||
| 56 | HOXB3 | 1427605_at 1456229_at | 7398 | 0.167 | 0.0139 | No | ||
| 57 | PLEK | 1417523_at 1448748_at 1448749_at | 7428 | 0.166 | 0.0141 | No | ||
| 58 | FIGN | 1450268_at | 7499 | 0.164 | 0.0124 | No | ||
| 59 | CDH16 | 1448906_at | 7520 | 0.163 | 0.0130 | No | ||
| 60 | CLDN16 | 1420434_at | 7831 | 0.153 | 0.0001 | No | ||
| 61 | LSAMP | 1443754_x_at 1455636_at 1460092_at | 8099 | 0.147 | -0.0108 | No | ||
| 62 | OLFM2 | 1435790_at | 8407 | 0.139 | -0.0237 | No | ||
| 63 | SHOX2 | 1420559_a_at 1438042_at | 8521 | 0.136 | -0.0276 | No | ||
| 64 | KCTD4 | 1420537_at 1441801_at | 8652 | 0.133 | -0.0324 | No | ||
| 65 | NEUROG1 | 1438551_at 1450836_at | 8727 | 0.131 | -0.0346 | No | ||
| 66 | PURG | 1424970_at 1452029_a_at | 8751 | 0.130 | -0.0345 | No | ||
| 67 | NEUROD1 | 1426412_at 1426413_at | 8767 | 0.130 | -0.0340 | No | ||
| 68 | KCNQ4 | 1435721_at 1442909_at | 8914 | 0.126 | -0.0395 | No | ||
| 69 | ATOH1 | 1449822_at | 9016 | 0.124 | -0.0431 | No | ||
| 70 | GNGT2 | 1428733_at | 9037 | 0.123 | -0.0428 | No | ||
| 71 | MYOCD | 1425808_a_at 1425978_at | 9321 | 0.116 | -0.0548 | No | ||
| 72 | HAGH | 1424171_a_at 1424172_at 1437519_x_at | 9373 | 0.115 | -0.0561 | No | ||
| 73 | TNF | 1419607_at | 9455 | 0.113 | -0.0588 | No | ||
| 74 | NUDT10 | 1435061_at | 9554 | 0.111 | -0.0623 | No | ||
| 75 | YTHDF1 | 1415767_at 1437102_at 1437456_x_at | 9690 | 0.107 | -0.0675 | No | ||
| 76 | HOXA10 | 1431475_a_at 1446408_at | 9730 | 0.106 | -0.0683 | No | ||
| 77 | ADM | 1416077_at 1447839_x_at | 10131 | 0.097 | -0.0859 | No | ||
| 78 | CACNB1 | 1425777_at 1426108_s_at 1440940_at 1451834_at | 10138 | 0.097 | -0.0853 | No | ||
| 79 | IL13 | 1420802_at | 10244 | 0.094 | -0.0892 | No | ||
| 80 | MAMDC1 | 1442206_at 1442561_at 1445372_at 1458147_at | 10294 | 0.093 | -0.0906 | No | ||
| 81 | ACLY | 1425326_at 1438389_x_at 1439445_x_at 1439459_x_at 1446315_at 1451666_at | 10308 | 0.093 | -0.0904 | No | ||
| 82 | NR3C1 | 1421866_at 1421867_at 1453860_s_at 1457635_s_at 1460303_at | 10372 | 0.091 | -0.0924 | No | ||
| 83 | KCNK2 | 1445929_at 1449158_at | 10596 | 0.086 | -0.1019 | No | ||
| 84 | NUDT3 | 1451575_a_at | 10719 | 0.084 | -0.1068 | No | ||
| 85 | RORA | 1420583_a_at 1424034_at 1424035_at 1436325_at 1436326_at 1441085_at 1443511_at 1443647_at 1446360_at 1455165_at 1457177_at 1458129_at | 10762 | 0.082 | -0.1079 | No | ||
| 86 | ELAVL2 | 1421881_a_at 1421882_a_at 1421883_at 1433052_at | 10772 | 0.082 | -0.1076 | No | ||
| 87 | SEMA3A | 1420416_at 1420417_at 1449865_at | 10890 | 0.079 | -0.1123 | No | ||
| 88 | SLC31A1 | 1433750_at 1455285_at | 10920 | 0.078 | -0.1129 | No | ||
| 89 | VAX1 | 1421774_at | 10974 | 0.077 | -0.1146 | No | ||
| 90 | NFATC4 | 1423380_s_at 1432821_at 1454369_a_at | 10981 | 0.077 | -0.1142 | No | ||
| 91 | A2BP1 | 1418314_a_at 1438217_at 1447735_x_at 1455358_at 1456921_at | 11030 | 0.075 | -0.1157 | No | ||
| 92 | LMO3 | 1455754_at | 11248 | 0.071 | -0.1250 | No | ||
| 93 | ZCCHC5 | 1437355_at | 11281 | 0.071 | -0.1259 | No | ||
| 94 | SOX5 | 1423500_a_at 1432189_a_at 1432190_at 1440827_x_at 1446461_at 1452511_at 1459261_at | 11524 | 0.065 | -0.1364 | No | ||
| 95 | ERG | 1425370_a_at 1440244_at | 11676 | 0.061 | -0.1428 | No | ||
| 96 | HOXB6 | 1451660_a_at | 11693 | 0.060 | -0.1430 | No | ||
| 97 | OBFC1 | 1434737_at 1441896_x_at 1445544_x_at 1456679_at 1458998_at | 11918 | 0.054 | -0.1528 | No | ||
| 98 | VSNL1 | 1420955_at 1450055_at | 11945 | 0.054 | -0.1535 | No | ||
| 99 | SLC35A2 | 1422827_x_at 1432533_a_at 1436664_a_at 1439433_a_at 1453858_at 1457024_x_at | 11983 | 0.053 | -0.1547 | No | ||
| 100 | JARID2 | 1422698_s_at 1446082_at 1450710_at 1456661_at | 12069 | 0.051 | -0.1582 | No | ||
| 101 | PPM1E | 1434990_at 1445741_at | 12076 | 0.050 | -0.1580 | No | ||
| 102 | SOX2 | 1416967_at | 12260 | 0.046 | -0.1660 | No | ||
| 103 | HAND2 | 1422221_at 1436041_at 1443854_at 1446812_at | 12388 | 0.044 | -0.1714 | No | ||
| 104 | LRFN5 | 1441502_at 1454687_at | 12486 | 0.041 | -0.1755 | No | ||
| 105 | OTX1 | 1437601_at | 12770 | 0.035 | -0.1882 | No | ||
| 106 | EVI1 | 1423011_at 1438325_at | 12919 | 0.031 | -0.1947 | No | ||
| 107 | ARHGAP6 | 1417704_a_at 1451867_x_at 1456333_a_at | 12965 | 0.030 | -0.1965 | No | ||
| 108 | PCDH9 | 1439322_at 1442659_at 1444215_at 1444724_at 1445052_at 1445061_at 1457973_at | 12998 | 0.029 | -0.1977 | No | ||
| 109 | ARHGAP5 | 1423194_at 1450896_at 1450897_at 1457410_at | 13009 | 0.029 | -0.1979 | No | ||
| 110 | NPAS2 | 1421036_at 1421037_at | 13184 | 0.024 | -0.2057 | No | ||
| 111 | PER2 | 1417602_at 1417603_at 1445892_at 1457350_at | 13291 | 0.021 | -0.2104 | No | ||
| 112 | BNC2 | 1438861_at 1440593_at 1459132_at | 13352 | 0.019 | -0.2130 | No | ||
| 113 | NKX2-2 | 1421112_at | 13361 | 0.019 | -0.2132 | No | ||
| 114 | SAMSN1 | 1421457_a_at 1431539_at | 13388 | 0.018 | -0.2142 | No | ||
| 115 | FOSB | 1422134_at | 13390 | 0.018 | -0.2141 | No | ||
| 116 | PRELP | 1416321_s_at 1416322_at | 13432 | 0.017 | -0.2158 | No | ||
| 117 | WRN | 1425074_at 1425982_a_at 1450163_a_at | 13473 | 0.016 | -0.2175 | No | ||
| 118 | DLL1 | 1419204_at | 13645 | 0.011 | -0.2253 | No | ||
| 119 | HEY1 | 1415999_at | 13675 | 0.010 | -0.2265 | No | ||
| 120 | WNT1 | 1425377_at | 13758 | 0.008 | -0.2302 | No | ||
| 121 | ST5 | 1428372_at | 14095 | -0.001 | -0.2456 | No | ||
| 122 | EDNRA | 1433525_at 1440093_at 1451691_at 1460513_a_at | 14216 | -0.005 | -0.2511 | No | ||
| 123 | CACNG2 | 1420596_at 1440210_at 1453363_at | 14231 | -0.005 | -0.2517 | No | ||
| 124 | CACNA2D3 | 1419225_at | 14240 | -0.005 | -0.2520 | No | ||
| 125 | SUMO2 | 1415782_at | 14318 | -0.007 | -0.2555 | No | ||
| 126 | GCNT2 | 1421415_s_at 1425503_at 1430826_s_at 1437607_at 1438660_at 1450238_at 1451733_at 1453674_at | 14367 | -0.009 | -0.2576 | No | ||
| 127 | TNFSF10 | 1420412_at 1439680_at 1459913_at | 14582 | -0.016 | -0.2673 | No | ||
| 128 | GNAZ | 1426517_at 1435268_at | 14658 | -0.019 | -0.2706 | No | ||
| 129 | GRIN2B | 1422223_at 1431700_at | 14738 | -0.021 | -0.2741 | No | ||
| 130 | SLC24A4 | 1435206_at | 14839 | -0.024 | -0.2784 | No | ||
| 131 | ENAM | 1421699_at | 14970 | -0.029 | -0.2842 | No | ||
| 132 | IGFBP5 | 1422313_a_at 1452114_s_at | 14975 | -0.029 | -0.2841 | No | ||
| 133 | LTBP1 | 1419786_at 1440678_at 1446232_at 1447547_at 1448870_at 1457852_at 1458739_at | 15313 | -0.041 | -0.2992 | No | ||
| 134 | DDX52 | 1434607_at 1434608_at | 15314 | -0.041 | -0.2988 | No | ||
| 135 | TNMD | 1417979_at | 15427 | -0.046 | -0.3036 | No | ||
| 136 | RIMS1 | 1435667_at 1438305_at 1444393_at | 15693 | -0.056 | -0.3152 | No | ||
| 137 | PEX16 | 1425021_a_at 1436253_at 1447801_x_at | 15912 | -0.066 | -0.3247 | No | ||
| 138 | SEPX1 | 1418888_a_at | 16272 | -0.084 | -0.3404 | No | ||
| 139 | HPCAL1 | 1448812_at | 16305 | -0.085 | -0.3411 | No | ||
| 140 | SPINK5 | 1430567_at | 16315 | -0.086 | -0.3407 | No | ||
| 141 | NCDN | 1416852_a_at 1416853_at | 16457 | -0.095 | -0.3463 | No | ||
| 142 | CCDC6 | 1428311_at 1459159_a_at | 16614 | -0.106 | -0.3525 | No | ||
| 143 | ZFHX1B | 1422748_at 1434298_at 1442393_at 1445518_at 1454200_at 1456389_at | 16812 | -0.119 | -0.3605 | No | ||
| 144 | GSTA4 | 1416368_at | 16843 | -0.121 | -0.3608 | No | ||
| 145 | TACSTD2 | 1423323_at | 16871 | -0.124 | -0.3609 | No | ||
| 146 | CASK | 1422518_at 1422519_at 1427692_a_at 1445152_at | 16955 | -0.130 | -0.3635 | No | ||
| 147 | DAPK1 | 1426915_at 1427358_a_at 1445032_at | 17131 | -0.145 | -0.3703 | No | ||
| 148 | LBX1 | 1422323_a_at 1427400_at | 17313 | -0.162 | -0.3771 | No | ||
| 149 | PAK1IP1 | 1423766_at 1430875_a_at | 17568 | -0.186 | -0.3871 | No | ||
| 150 | POLR3F | 1429686_at 1430328_at 1431999_at 1439075_at 1451773_s_at | 17695 | -0.200 | -0.3910 | No | ||
| 151 | CHCHD7 | 1440158_x_at 1444318_at 1446844_at 1454640_at | 17703 | -0.201 | -0.3895 | No | ||
| 152 | LRRTM1 | 1437746_at 1452624_at 1455883_a_at | 17809 | -0.213 | -0.3924 | No | ||
| 153 | SMAD5 | 1421047_at 1433641_at 1451873_a_at | 17862 | -0.219 | -0.3928 | No | ||
| 154 | ANGPT2 | 1422415_at 1446903_at 1448831_at | 18431 | -0.298 | -0.4162 | No | ||
| 155 | RGS3 | 1425701_a_at 1427648_at 1449516_a_at 1454026_a_at | 18473 | -0.303 | -0.4153 | No | ||
| 156 | TPM1 | 1423049_a_at 1423721_at 1447713_at 1456623_at | 18508 | -0.307 | -0.4141 | No | ||
| 157 | MNT | 1418192_at | 18590 | -0.320 | -0.4149 | No | ||
| 158 | TGIF | 1422286_a_at | 18672 | -0.332 | -0.4156 | No | ||
| 159 | BCAR3 | 1415936_at 1431564_at | 18956 | -0.386 | -0.4251 | No | ||
| 160 | LEMD2 | 1424326_at | 18970 | -0.388 | -0.4221 | No | ||
| 161 | TOP1 | 1423474_at 1438495_at 1457573_at 1459672_at | 19100 | -0.415 | -0.4242 | No | ||
| 162 | NRAS | 1422688_a_at 1454060_a_at | 19135 | -0.424 | -0.4219 | No | ||
| 163 | CHD6 | 1427384_at 1431047_at | 19393 | -0.483 | -0.4293 | No | ||
| 164 | CPEB4 | 1420617_at 1420618_at 1449931_at | 19768 | -0.585 | -0.4412 | Yes | ||
| 165 | ZBTB9 | 1429125_at | 19789 | -0.590 | -0.4367 | Yes | ||
| 166 | HS3ST3B1 | 1421331_at 1421332_at 1433977_at 1442816_at 1444055_at | 19819 | -0.601 | -0.4326 | Yes | ||
| 167 | HDAC4 | 1436758_at 1447566_at 1454693_at | 20113 | -0.702 | -0.4396 | Yes | ||
| 168 | PDE3B | 1433694_at | 20125 | -0.705 | -0.4337 | Yes | ||
| 169 | DOCK9 | 1445232_at 1446803_at 1446833_at 1446927_at | 20128 | -0.707 | -0.4273 | Yes | ||
| 170 | PHOX2B | 1422232_at 1455907_x_at | 20380 | -0.812 | -0.4314 | Yes | ||
| 171 | EPC2 | 1455317_at | 20395 | -0.823 | -0.4246 | Yes | ||
| 172 | ARRDC4 | 1424759_at | 20520 | -0.880 | -0.4222 | Yes | ||
| 173 | ABI3 | 1446191_at 1452928_at | 20625 | -0.934 | -0.4185 | Yes | ||
| 174 | CENTD1 | 1436895_at 1446649_at 1452291_at 1456337_at 1458173_at | 20747 | -1.027 | -0.4146 | Yes | ||
| 175 | RAMP1 | 1417481_at 1447474_at | 20780 | -1.051 | -0.4065 | Yes | ||
| 176 | ANGPTL2 | 1421002_at 1431848_at | 20782 | -1.051 | -0.3969 | Yes | ||
| 177 | FLI1 | 1422024_at 1433512_at 1441584_at | 20787 | -1.055 | -0.3875 | Yes | ||
| 178 | PURA | 1420628_at 1438219_at 1449934_at 1453783_at 1456898_at | 20835 | -1.087 | -0.3797 | Yes | ||
| 179 | DUSP2 | 1450698_at | 20842 | -1.091 | -0.3700 | Yes | ||
| 180 | FOXP1 | 1421140_a_at 1421141_a_at 1421142_s_at 1435222_at 1438802_at 1443258_at 1446280_at 1447209_at 1455242_at 1456851_at | 21031 | -1.284 | -0.3669 | Yes | ||
| 181 | USP32 | 1430403_at 1436159_at 1439857_at 1440399_at 1443689_at 1459857_at | 21033 | -1.285 | -0.3552 | Yes | ||
| 182 | SNTB2 | 1420371_at 1420372_at 1436986_at 1449840_at | 21056 | -1.304 | -0.3443 | Yes | ||
| 183 | MTSS1 | 1424826_s_at 1434036_at 1440847_at 1443729_at 1446284_at 1451496_at | 21124 | -1.383 | -0.3348 | Yes | ||
| 184 | MYLIP | 1424988_at | 21493 | -2.022 | -0.3332 | Yes | ||
| 185 | IHPK2 | 1428373_at 1435319_at | 21544 | -2.129 | -0.3160 | Yes | ||
| 186 | FUT8 | 1441015_at 1442483_at 1443656_at 1456367_at 1460319_at | 21554 | -2.160 | -0.2967 | Yes | ||
| 187 | PPP2CA | 1417367_at 1444875_at 1456390_at | 21563 | -2.190 | -0.2771 | Yes | ||
| 188 | REST | 1425564_at 1425565_at 1428227_at | 21648 | -2.452 | -0.2585 | Yes | ||
| 189 | TBL1X | 1434643_at 1434644_at 1441435_at 1446736_at 1455042_at | 21686 | -2.601 | -0.2364 | Yes | ||
| 190 | FGF13 | 1418497_at 1418498_at | 21695 | -2.635 | -0.2127 | Yes | ||
| 191 | ATRX | 1420945_at 1420946_at 1420947_at 1438750_at 1441014_at 1441771_at 1450051_at 1453734_at 1456832_at | 21709 | -2.755 | -0.1882 | Yes | ||
| 192 | NEO1 | 1423513_at 1434931_at 1440789_at 1444295_at 1447693_s_at 1447694_x_at 1459485_at | 21731 | -2.823 | -0.1633 | Yes | ||
| 193 | SSBP3 | 1425940_a_at 1427917_s_at 1439136_at 1443554_at 1457560_at 1458749_at | 21778 | -3.006 | -0.1379 | Yes | ||
| 194 | HBP1 | 1432143_a_at 1438430_at 1446269_at 1460367_at | 21819 | -3.274 | -0.1099 | Yes | ||
| 195 | S100A9 | 1448756_at | 21895 | -4.073 | -0.0761 | Yes | ||
| 196 | TEC | 1441524_at 1460204_at | 21901 | -4.227 | -0.0377 | Yes | ||
| 197 | EGR2 | 1427682_a_at 1427683_at | 21904 | -4.293 | 0.0015 | Yes |