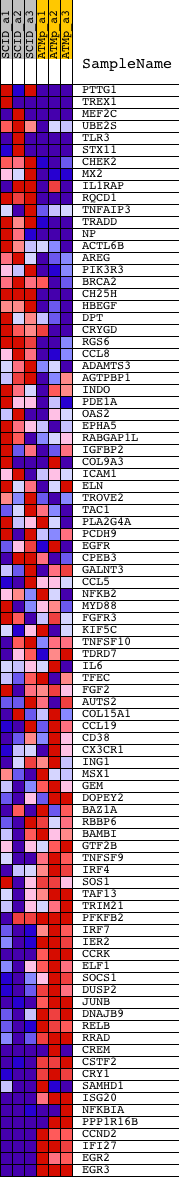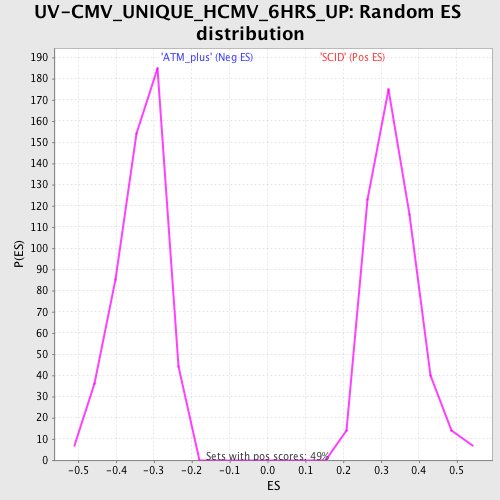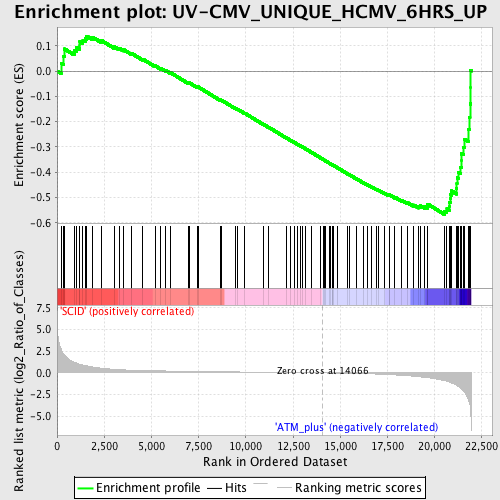
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus_repos |
| Phenotype | phenotype_SCID_versus_ATM_plus.cls#SCID_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | UV-CMV_UNIQUE_HCMV_6HRS_UP |
| Enrichment Score (ES) | -0.56721 |
| Normalized Enrichment Score (NES) | -1.6996583 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.16631301 |
| FWER p-Value | 0.904 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PTTG1 | 1419620_at 1424105_a_at 1438390_s_at | 214 | 2.722 | 0.0314 | No | ||
| 2 | TREX1 | 1450672_a_at 1459618_at | 353 | 2.155 | 0.0578 | No | ||
| 3 | MEF2C | 1421027_a_at 1421028_a_at 1424852_at 1439946_at 1445420_at 1446484_at 1451506_at 1451507_at 1458551_at | 392 | 2.064 | 0.0873 | No | ||
| 4 | UBE2S | 1416726_s_at 1430962_at | 911 | 1.217 | 0.0820 | No | ||
| 5 | TLR3 | 1422781_at 1422782_s_at | 1024 | 1.125 | 0.0940 | No | ||
| 6 | STX11 | 1423576_a_at 1451032_at 1453228_at 1457780_at | 1204 | 0.989 | 0.1008 | No | ||
| 7 | CHEK2 | 1422747_at | 1206 | 0.989 | 0.1157 | No | ||
| 8 | MX2 | 1419676_at | 1368 | 0.900 | 0.1220 | No | ||
| 9 | IL1RAP | 1421843_at 1439697_at 1442614_at 1449585_at | 1509 | 0.836 | 0.1282 | No | ||
| 10 | RQCD1 | 1450887_at | 1563 | 0.810 | 0.1381 | No | ||
| 11 | TNFAIP3 | 1433699_at 1450829_at | 1881 | 0.674 | 0.1338 | No | ||
| 12 | TRADD | 1429117_at 1439910_a_at 1452622_a_at | 2339 | 0.543 | 0.1211 | No | ||
| 13 | NP | 1416530_a_at | 3025 | 0.415 | 0.0960 | No | ||
| 14 | ACTL6B | 1422564_at | 3277 | 0.378 | 0.0903 | No | ||
| 15 | AREG | 1421134_at | 3520 | 0.349 | 0.0845 | No | ||
| 16 | PIK3R3 | 1445097_at 1447068_at 1456482_at | 3939 | 0.314 | 0.0701 | No | ||
| 17 | BRCA2 | 1419076_a_at | 4529 | 0.274 | 0.0473 | No | ||
| 18 | CH25H | 1449227_at | 5196 | 0.244 | 0.0205 | No | ||
| 19 | HBEGF | 1418349_at 1418350_at | 5500 | 0.231 | 0.0101 | No | ||
| 20 | DPT | 1418511_at | 5748 | 0.221 | 0.0022 | No | ||
| 21 | CRYGD | 1419141_at | 6022 | 0.211 | -0.0071 | No | ||
| 22 | RGS6 | 1427340_at 1452399_at | 6962 | 0.179 | -0.0474 | No | ||
| 23 | CCL8 | 1419684_at | 6987 | 0.179 | -0.0458 | No | ||
| 24 | ADAMTS3 | 1441693_at 1458143_at | 7435 | 0.166 | -0.0637 | No | ||
| 25 | AGTPBP1 | 1429933_at 1432385_a_at | 7462 | 0.165 | -0.0624 | No | ||
| 26 | INDO | 1420437_at | 8630 | 0.134 | -0.1138 | No | ||
| 27 | PDE1A | 1431625_at 1449298_a_at | 8731 | 0.131 | -0.1164 | No | ||
| 28 | OAS2 | 1425065_at | 9435 | 0.113 | -0.1469 | No | ||
| 29 | EPHA5 | 1420557_at 1437920_at 1444690_at | 9550 | 0.111 | -0.1504 | No | ||
| 30 | RABGAP1L | 1421488_at 1429196_at 1429197_s_at 1434062_at 1441202_at 1453365_at 1457178_at 1458155_at | 9944 | 0.101 | -0.1669 | No | ||
| 31 | IGFBP2 | 1454159_a_at | 10916 | 0.078 | -0.2101 | No | ||
| 32 | COL9A3 | 1420280_x_at 1460693_a_at 1460734_at | 11215 | 0.072 | -0.2227 | No | ||
| 33 | ICAM1 | 1424067_at | 12127 | 0.049 | -0.2636 | No | ||
| 34 | ELN | 1420854_at 1420855_at 1446221_at | 12377 | 0.044 | -0.2744 | No | ||
| 35 | TROVE2 | 1423433_at 1436533_at 1436534_at 1436535_at | 12563 | 0.039 | -0.2823 | No | ||
| 36 | TAC1 | 1416783_at 1419411_at 1431883_at | 12744 | 0.035 | -0.2900 | No | ||
| 37 | PLA2G4A | 1448558_a_at | 12887 | 0.032 | -0.2960 | No | ||
| 38 | PCDH9 | 1439322_at 1442659_at 1444215_at 1444724_at 1445052_at 1445061_at 1457973_at | 12998 | 0.029 | -0.3006 | No | ||
| 39 | EGFR | 1424932_at 1432647_at 1435888_at 1451530_at 1454313_at 1457563_at 1460420_a_at | 13159 | 0.025 | -0.3075 | No | ||
| 40 | CPEB3 | 1437765_at 1445868_at 1455372_at 1456048_at | 13460 | 0.016 | -0.3210 | No | ||
| 41 | GALNT3 | 1417588_at | 13959 | 0.003 | -0.3437 | No | ||
| 42 | CCL5 | 1418126_at | 14085 | -0.001 | -0.3495 | No | ||
| 43 | NFKB2 | 1425902_a_at 1429128_x_at | 14187 | -0.004 | -0.3540 | No | ||
| 44 | MYD88 | 1419272_at | 14229 | -0.005 | -0.3558 | No | ||
| 45 | FGFR3 | 1421841_at 1425796_a_at | 14435 | -0.012 | -0.3650 | No | ||
| 46 | KIF5C | 1422945_a_at 1427635_at 1447454_at 1450804_at 1455266_at 1457255_x_at | 14477 | -0.013 | -0.3667 | No | ||
| 47 | TNFSF10 | 1420412_at 1439680_at 1459913_at | 14582 | -0.016 | -0.3712 | No | ||
| 48 | TDRD7 | 1426716_at 1459216_at | 14587 | -0.016 | -0.3712 | No | ||
| 49 | IL6 | 1450297_at | 14615 | -0.017 | -0.3721 | No | ||
| 50 | TFEC | 1419537_at | 14846 | -0.025 | -0.3823 | No | ||
| 51 | FGF2 | 1449826_a_at | 15397 | -0.045 | -0.4068 | No | ||
| 52 | AUTS2 | 1427293_a_at 1438680_at 1441869_x_at 1442360_at 1443062_at 1443070_at 1443770_x_at 1444069_at 1445307_at 1445469_at 1445538_at 1446849_at 1452379_at 1457139_at 1458592_at 1459338_at | 15478 | -0.048 | -0.4097 | No | ||
| 53 | COL15A1 | 1448755_at | 15855 | -0.064 | -0.4260 | No | ||
| 54 | CCL19 | 1449277_at | 16233 | -0.082 | -0.4420 | No | ||
| 55 | CD38 | 1433741_at 1450136_at | 16430 | -0.094 | -0.4496 | No | ||
| 56 | CX3CR1 | 1450019_at 1450020_at | 16667 | -0.110 | -0.4587 | No | ||
| 57 | ING1 | 1416860_s_at 1433267_at 1448496_a_at 1454534_at | 16893 | -0.125 | -0.4671 | No | ||
| 58 | MSX1 | 1417127_at 1448601_s_at | 17047 | -0.138 | -0.4720 | No | ||
| 59 | GEM | 1426063_a_at | 17326 | -0.164 | -0.4823 | No | ||
| 60 | DOPEY2 | 1428330_at 1449904_at | 17584 | -0.188 | -0.4912 | No | ||
| 61 | BAZ1A | 1433599_at 1447930_at | 17586 | -0.188 | -0.4884 | No | ||
| 62 | RBBP6 | 1425114_at 1425115_at 1425421_at 1426487_a_at 1453611_at | 17893 | -0.223 | -0.4990 | No | ||
| 63 | BAMBI | 1423753_at | 18244 | -0.273 | -0.5109 | No | ||
| 64 | GTF2B | 1438066_at 1451135_at | 18568 | -0.317 | -0.5209 | No | ||
| 65 | TNFSF9 | 1422924_at | 18884 | -0.371 | -0.5297 | No | ||
| 66 | IRF4 | 1421173_at 1421174_at | 19146 | -0.427 | -0.5352 | No | ||
| 67 | SOS1 | 1421884_at 1421885_at 1421886_at | 19240 | -0.448 | -0.5326 | No | ||
| 68 | TAF13 | 1448707_at | 19478 | -0.502 | -0.5359 | No | ||
| 69 | TRIM21 | 1418077_at 1448940_at | 19614 | -0.542 | -0.5338 | No | ||
| 70 | PFKFB2 | 1422090_a_at 1422091_at 1422092_at 1429486_at 1431901_a_at | 19623 | -0.544 | -0.5260 | No | ||
| 71 | IRF7 | 1417244_a_at | 20525 | -0.882 | -0.5539 | Yes | ||
| 72 | IER2 | 1416442_at | 20648 | -0.946 | -0.5451 | Yes | ||
| 73 | CCRK | 1422494_s_at | 20767 | -1.044 | -0.5347 | Yes | ||
| 74 | ELF1 | 1417540_at 1439319_at | 20774 | -1.049 | -0.5191 | Yes | ||
| 75 | SOCS1 | 1440047_at 1450446_a_at | 20823 | -1.082 | -0.5049 | Yes | ||
| 76 | DUSP2 | 1450698_at | 20842 | -1.091 | -0.4892 | Yes | ||
| 77 | JUNB | 1415899_at | 20884 | -1.130 | -0.4739 | Yes | ||
| 78 | DNAJB9 | 1417191_at | 21159 | -1.428 | -0.4648 | Yes | ||
| 79 | RELB | 1417856_at | 21176 | -1.451 | -0.4436 | Yes | ||
| 80 | RRAD | 1422562_at | 21183 | -1.472 | -0.4216 | Yes | ||
| 81 | CREM | 1418322_at 1430598_at 1430847_a_at 1449037_at | 21261 | -1.567 | -0.4013 | Yes | ||
| 82 | CSTF2 | 1419644_at 1419645_at 1455523_at | 21357 | -1.704 | -0.3799 | Yes | ||
| 83 | CRY1 | 1433733_a_at | 21412 | -1.823 | -0.3547 | Yes | ||
| 84 | SAMHD1 | 1418131_at 1434438_at 1444064_at | 21417 | -1.831 | -0.3272 | Yes | ||
| 85 | ISG20 | 1419569_a_at | 21521 | -2.074 | -0.3005 | Yes | ||
| 86 | NFKBIA | 1420088_at 1420089_at 1438157_s_at 1448306_at 1449731_s_at | 21592 | -2.242 | -0.2697 | Yes | ||
| 87 | PPP1R16B | 1425414_at 1455080_at | 21805 | -3.184 | -0.2312 | Yes | ||
| 88 | CCND2 | 1416122_at 1416123_at 1416124_at 1430127_a_at 1434745_at 1448229_s_at 1455956_x_at | 21828 | -3.314 | -0.1820 | Yes | ||
| 89 | IFI27 | 1426278_at | 21869 | -3.596 | -0.1293 | Yes | ||
| 90 | EGR2 | 1427682_a_at 1427683_at | 21904 | -4.293 | -0.0658 | Yes | ||
| 91 | EGR3 | 1421486_at 1436329_at | 21908 | -4.438 | 0.0013 | Yes |