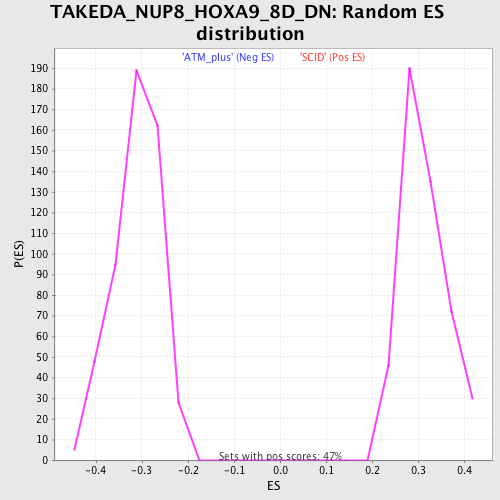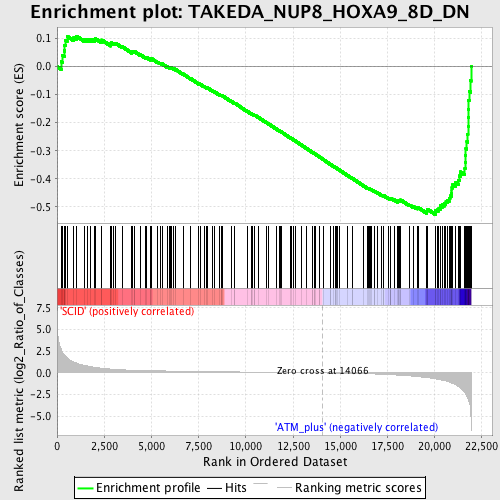
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus_repos |
| Phenotype | phenotype_SCID_versus_ATM_plus.cls#SCID_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | TAKEDA_NUP8_HOXA9_8D_DN |
| Enrichment Score (ES) | -0.5267119 |
| Normalized Enrichment Score (NES) | -1.7019506 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.17411889 |
| FWER p-Value | 0.897 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | GPR97 | 1417894_at | 215 | 2.720 | 0.0176 | No | ||
| 2 | ENC1 | 1420964_at 1420965_a_at 1450061_at | 273 | 2.403 | 0.0393 | No | ||
| 3 | NRP1 | 1418084_at 1448943_at 1448944_at 1457198_at | 379 | 2.098 | 0.0557 | No | ||
| 4 | S100B | 1419383_at 1434342_at | 405 | 2.034 | 0.0751 | No | ||
| 5 | KCNMA1 | 1424848_at 1425987_a_at 1440728_at 1444973_at 1445774_at 1457018_at | 435 | 1.963 | 0.0936 | No | ||
| 6 | MMP9 | 1416298_at 1448291_at | 529 | 1.771 | 0.1072 | No | ||
| 7 | PRNP | 1416130_at 1446507_at 1448233_at | 889 | 1.229 | 0.1031 | No | ||
| 8 | CTSB | 1417490_at 1417491_at 1417492_at 1444987_at 1448732_at | 1033 | 1.118 | 0.1079 | No | ||
| 9 | EXT1 | 1417730_at 1439877_at 1440011_at 1440092_at 1456655_at 1458054_at 1458296_at | 1458 | 0.859 | 0.0971 | No | ||
| 10 | EMP1 | 1416529_at 1459171_at | 1632 | 0.776 | 0.0970 | No | ||
| 11 | RAP2A | 1421574_at 1426965_at 1440473_at | 1793 | 0.707 | 0.0968 | No | ||
| 12 | ETNK1 | 1419884_at 1430996_at 1433514_at 1433515_s_at 1439972_at 1449650_at 1454633_at | 1970 | 0.647 | 0.0952 | No | ||
| 13 | CD9 | 1416066_at | 2024 | 0.628 | 0.0991 | No | ||
| 14 | CST7 | 1419202_at 1445263_at | 2357 | 0.539 | 0.0893 | No | ||
| 15 | ADAM19 | 1418402_at 1418403_at | 2373 | 0.536 | 0.0941 | No | ||
| 16 | CLU | 1418626_a_at 1437458_x_at 1437689_x_at 1454849_x_at | 2841 | 0.445 | 0.0771 | No | ||
| 17 | IL10 | 1450330_at | 2846 | 0.444 | 0.0814 | No | ||
| 18 | C1QA | 1417381_at | 2871 | 0.439 | 0.0848 | No | ||
| 19 | PLA2G7 | 1430700_a_at | 2997 | 0.421 | 0.0833 | No | ||
| 20 | EPAS1 | 1435436_at 1449888_at | 3108 | 0.401 | 0.0823 | No | ||
| 21 | MMP14 | 1416572_at 1448383_at | 3458 | 0.357 | 0.0699 | No | ||
| 22 | C9ORF19 | 1428492_at 1452803_at | 3962 | 0.312 | 0.0499 | No | ||
| 23 | CXCL1 | 1419209_at 1441855_x_at 1457644_s_at | 3966 | 0.311 | 0.0529 | No | ||
| 24 | CREB5 | 1440089_at 1442576_at 1457222_at | 4076 | 0.302 | 0.0510 | No | ||
| 25 | C5AR1 | 1422190_at 1439902_at | 4101 | 0.301 | 0.0529 | No | ||
| 26 | ARHGAP18 | 1426952_at 1431133_at 1438882_at | 4410 | 0.281 | 0.0416 | No | ||
| 27 | CD14 | 1417268_at | 4702 | 0.266 | 0.0309 | No | ||
| 28 | ST8SIA1 | 1419694_at 1419695_at 1455695_at | 4749 | 0.264 | 0.0315 | No | ||
| 29 | CLEC7A | 1420699_at | 4946 | 0.255 | 0.0251 | No | ||
| 30 | LGMN | 1448883_at | 4957 | 0.254 | 0.0272 | No | ||
| 31 | CDC42 | 1415724_a_at 1435807_at 1449574_a_at 1460708_s_at | 5007 | 0.252 | 0.0275 | No | ||
| 32 | OLIG1 | 1416149_at | 5335 | 0.238 | 0.0149 | No | ||
| 33 | GPNMB | 1439345_at 1448303_at | 5470 | 0.232 | 0.0111 | No | ||
| 34 | ANTXR1 | 1431995_at 1451446_at 1458657_at | 5563 | 0.228 | 0.0092 | No | ||
| 35 | SLC11A1 | 1420361_at | 5820 | 0.219 | -0.0004 | No | ||
| 36 | MAF | 1435828_at 1437473_at 1444073_at 1447849_s_at 1447945_at 1456060_at | 5957 | 0.213 | -0.0045 | No | ||
| 37 | GPR34 | 1422542_at | 6021 | 0.211 | -0.0052 | No | ||
| 38 | PPBP | 1418480_at | 6043 | 0.210 | -0.0041 | No | ||
| 39 | RAB30 | 1426452_a_at | 6189 | 0.205 | -0.0087 | No | ||
| 40 | HTR2C | 1435513_at 1450477_at | 6286 | 0.202 | -0.0110 | No | ||
| 41 | SLC41A2 | 1452445_at | 6672 | 0.189 | -0.0268 | No | ||
| 42 | SFRS6 | 1416720_at 1416721_s_at 1447898_s_at 1448454_at | 7077 | 0.176 | -0.0436 | No | ||
| 43 | HK3 | 1435490_at 1442798_x_at | 7504 | 0.164 | -0.0615 | No | ||
| 44 | F5 | 1449269_at | 7581 | 0.161 | -0.0633 | No | ||
| 45 | CLEC5A | 1421366_at | 7792 | 0.154 | -0.0714 | No | ||
| 46 | HRH4 | 1426099_at | 7923 | 0.151 | -0.0759 | No | ||
| 47 | RPS6KA2 | 1417542_at 1417543_at 1441311_at | 7944 | 0.150 | -0.0753 | No | ||
| 48 | HTRA1 | 1416749_at 1438251_x_at | 8249 | 0.143 | -0.0878 | No | ||
| 49 | TCERG1 | 1421033_a_at 1434434_s_at 1450100_a_at | 8345 | 0.140 | -0.0907 | No | ||
| 50 | CD200R1 | 1422191_at 1439703_at | 8605 | 0.134 | -0.1012 | No | ||
| 51 | COLEC12 | 1419693_at 1446535_at | 8690 | 0.132 | -0.1038 | No | ||
| 52 | RAPGEF2 | 1452833_at | 8715 | 0.131 | -0.1035 | No | ||
| 53 | UHMK1 | 1446120_at 1454260_at | 8734 | 0.131 | -0.1030 | No | ||
| 54 | CDH8 | 1422052_at 1441690_at 1455365_at 1456221_at | 9236 | 0.118 | -0.1249 | No | ||
| 55 | ST6GALNAC5 | 1419420_at 1446059_at 1449468_at | 9411 | 0.114 | -0.1317 | No | ||
| 56 | UTS2 | 1421799_at | 9413 | 0.114 | -0.1306 | No | ||
| 57 | IL5RA | 1421620_at | 10059 | 0.098 | -0.1592 | No | ||
| 58 | MS4A2 | 1421475_at 1443264_at | 10272 | 0.093 | -0.1680 | No | ||
| 59 | MARCO | 1449498_at 1458297_s_at | 10306 | 0.093 | -0.1686 | No | ||
| 60 | MMP12 | 1449153_at | 10362 | 0.092 | -0.1702 | No | ||
| 61 | SRPX | 1451939_a_at | 10440 | 0.090 | -0.1728 | No | ||
| 62 | PTCD1 | 1416391_at 1425096_a_at | 10478 | 0.089 | -0.1736 | No | ||
| 63 | STAC | 1418957_at 1445344_at | 10667 | 0.085 | -0.1814 | No | ||
| 64 | PPP1R1B | 1451331_at | 11087 | 0.074 | -0.1999 | No | ||
| 65 | CAMK1D | 1426389_at 1438643_at 1452050_at | 11214 | 0.072 | -0.2049 | No | ||
| 66 | NEK1 | 1434267_at 1438592_at 1453612_at 1458211_at | 11603 | 0.063 | -0.2221 | No | ||
| 67 | ATP6V0D2 | 1434798_at 1444176_at | 11804 | 0.057 | -0.2307 | No | ||
| 68 | DACH1 | 1420694_a_at 1420695_at 1433743_at 1447174_at 1459381_at | 11813 | 0.057 | -0.2305 | No | ||
| 69 | PDK4 | 1417273_at | 11877 | 0.055 | -0.2328 | No | ||
| 70 | SLCO2B1 | 1433933_s_at 1454777_at 1458171_at | 12366 | 0.044 | -0.2548 | No | ||
| 71 | SLC25A27 | 1431201_at 1440090_at 1454230_a_at 1458119_at | 12417 | 0.043 | -0.2567 | No | ||
| 72 | TSPAN2 | 1424567_at 1424568_at 1432417_a_at | 12521 | 0.040 | -0.2610 | No | ||
| 73 | FPR1 | 1450808_at | 12605 | 0.038 | -0.2644 | No | ||
| 74 | ABP1 | 1420709_s_at 1424600_at | 12957 | 0.030 | -0.2802 | No | ||
| 75 | DOCK4 | 1431114_at 1436405_at 1441462_at 1441469_at 1446283_at 1457146_at 1459279_at | 13206 | 0.024 | -0.2914 | No | ||
| 76 | TEX15 | 1420719_at | 13508 | 0.015 | -0.3050 | No | ||
| 77 | ALDH2 | 1434987_at 1434988_x_at 1437410_at 1448143_at | 13533 | 0.014 | -0.3060 | No | ||
| 78 | GJA4 | 1416255_at | 13639 | 0.011 | -0.3107 | No | ||
| 79 | ALOX5 | 1441962_at | 13651 | 0.011 | -0.3111 | No | ||
| 80 | RNASE1 | 1416523_at | 13690 | 0.010 | -0.3127 | No | ||
| 81 | CXCL5 | 1419728_at | 13874 | 0.005 | -0.3211 | No | ||
| 82 | ANKRD29 | 1436294_at 1438756_at | 13893 | 0.005 | -0.3219 | No | ||
| 83 | ADAM9 | 1416094_at | 14117 | -0.002 | -0.3321 | No | ||
| 84 | DAB2 | 1420498_a_at 1423805_at 1429693_at 1430604_a_at | 14484 | -0.013 | -0.3488 | No | ||
| 85 | JUN | 1417409_at 1448694_at | 14497 | -0.014 | -0.3492 | No | ||
| 86 | SLC16A7 | 1448502_at | 14631 | -0.018 | -0.3551 | No | ||
| 87 | SLPI | 1448377_at | 14745 | -0.021 | -0.3601 | No | ||
| 88 | IL1R2 | 1419532_at 1457108_at | 14757 | -0.022 | -0.3604 | No | ||
| 89 | PNPLA1 | 1457448_at | 14807 | -0.023 | -0.3624 | No | ||
| 90 | NCF2 | 1448561_at | 14861 | -0.025 | -0.3646 | No | ||
| 91 | SLC30A7 | 1450697_at | 14954 | -0.028 | -0.3685 | No | ||
| 92 | CD163 | 1419144_at | 15369 | -0.044 | -0.3871 | No | ||
| 93 | C1QC | 1449401_at | 15631 | -0.053 | -0.3985 | No | ||
| 94 | P2RY10 | 1452815_at | 15647 | -0.054 | -0.3987 | No | ||
| 95 | ITGAM | 1422046_at | 16240 | -0.082 | -0.4250 | No | ||
| 96 | RAPH1 | 1434302_at 1434303_at 1444586_at 1456918_at | 16455 | -0.095 | -0.4339 | No | ||
| 97 | C1QB | 1417063_at 1434366_x_at 1437726_x_at 1457322_at | 16476 | -0.097 | -0.4338 | No | ||
| 98 | TRPA1 | 1457164_at | 16539 | -0.102 | -0.4356 | No | ||
| 99 | ZDHHC19 | 1438607_at | 16594 | -0.105 | -0.4370 | No | ||
| 100 | FCGR2B | 1435476_a_at 1435477_s_at 1447622_at 1451941_a_at 1455332_x_at | 16627 | -0.107 | -0.4374 | No | ||
| 101 | ATP10D | 1436544_at | 16815 | -0.120 | -0.4448 | No | ||
| 102 | CD24 | 1416034_at 1437502_x_at 1448182_a_at | 16822 | -0.120 | -0.4439 | No | ||
| 103 | IL7R | 1448575_at 1448576_at | 16982 | -0.133 | -0.4498 | No | ||
| 104 | SFRS4 | 1448778_at | 17201 | -0.152 | -0.4583 | No | ||
| 105 | SLC7A7 | 1417392_a_at 1447181_s_at | 17267 | -0.158 | -0.4597 | No | ||
| 106 | C1ORF24 | 1422567_at 1454942_at | 17269 | -0.158 | -0.4581 | No | ||
| 107 | NPL | 1424265_at | 17530 | -0.183 | -0.4682 | No | ||
| 108 | ELOVL7 | 1424097_at 1424098_at 1440354_at 1441891_x_at | 17656 | -0.197 | -0.4720 | No | ||
| 109 | SLC16A6 | 1417884_at | 17660 | -0.198 | -0.4701 | No | ||
| 110 | ENTPD1 | 1423326_at 1450939_at 1453586_at | 17679 | -0.199 | -0.4689 | No | ||
| 111 | SLC26A11 | 1436865_at 1460115_at | 17853 | -0.218 | -0.4747 | No | ||
| 112 | SLAMF8 | 1425294_at | 18046 | -0.245 | -0.4810 | No | ||
| 113 | CYBB | 1422978_at 1436778_at 1436779_at | 18055 | -0.246 | -0.4789 | No | ||
| 114 | PYGL | 1417741_at | 18096 | -0.251 | -0.4782 | No | ||
| 115 | IL3RA | 1419712_at | 18157 | -0.261 | -0.4783 | No | ||
| 116 | CHI3L1 | 1451537_at | 18176 | -0.263 | -0.4765 | No | ||
| 117 | ARPC5 | 1444568_at 1448129_at | 18205 | -0.267 | -0.4751 | No | ||
| 118 | LGALS12 | 1417686_at 1417687_at | 18670 | -0.332 | -0.4930 | No | ||
| 119 | C7ORF10 | 1424597_at 1451404_at | 18855 | -0.366 | -0.4978 | No | ||
| 120 | AMFR | 1450102_a_at | 19062 | -0.408 | -0.5031 | No | ||
| 121 | CAPZA2 | 1423057_at 1423058_at | 19115 | -0.419 | -0.5013 | No | ||
| 122 | TRAT1 | 1427532_at 1437561_at | 19578 | -0.532 | -0.5171 | No | ||
| 123 | NIPBL | 1430309_at 1437158_at 1440690_at 1442103_at 1443992_at 1447408_at 1453576_at | 19594 | -0.536 | -0.5124 | No | ||
| 124 | TM6SF1 | 1451353_at | 19618 | -0.543 | -0.5079 | No | ||
| 125 | TGFBI | 1415871_at 1437463_x_at 1448123_s_at 1456250_x_at | 20028 | -0.669 | -0.5200 | Yes | ||
| 126 | RHOH | 1429319_at 1441385_at 1443015_at | 20029 | -0.669 | -0.5132 | Yes | ||
| 127 | CORO2A | 1445034_at 1452367_at | 20127 | -0.706 | -0.5105 | Yes | ||
| 128 | GPR137B | 1443208_at 1449670_x_at 1449671_at 1450881_s_at | 20215 | -0.739 | -0.5070 | Yes | ||
| 129 | NDRG2 | 1448154_at | 20288 | -0.776 | -0.5025 | Yes | ||
| 130 | SGK | 1416041_at | 20303 | -0.782 | -0.4952 | Yes | ||
| 131 | MARCH1 | 1431367_at 1434955_at 1440209_at | 20416 | -0.831 | -0.4920 | Yes | ||
| 132 | TDO2 | 1419093_at 1449337_at | 20518 | -0.879 | -0.4877 | Yes | ||
| 133 | LCP2 | 1418641_at 1418642_at | 20596 | -0.919 | -0.4820 | Yes | ||
| 134 | RAB3C | 1432415_at 1432432_a_at 1449494_at | 20662 | -0.962 | -0.4752 | Yes | ||
| 135 | HVCN1 | 1424032_at | 20762 | -1.041 | -0.4693 | Yes | ||
| 136 | HSD11B1 | 1449038_at | 20811 | -1.070 | -0.4607 | Yes | ||
| 137 | AP1S2 | 1437998_at 1447903_x_at 1452657_at 1456252_x_at 1460036_at | 20871 | -1.116 | -0.4521 | Yes | ||
| 138 | BCL2 | 1422938_at 1427818_at 1437122_at 1440770_at 1443837_x_at 1457687_at | 20911 | -1.157 | -0.4422 | Yes | ||
| 139 | PCSK5 | 1424605_at 1437339_s_at 1438248_at 1451406_a_at 1452475_at 1459124_at | 20913 | -1.159 | -0.4305 | Yes | ||
| 140 | FYB | 1452117_a_at | 20961 | -1.211 | -0.4205 | Yes | ||
| 141 | RAP1A | 1424139_at | 21116 | -1.376 | -0.4136 | Yes | ||
| 142 | PPP1R3B | 1436590_at | 21271 | -1.578 | -0.4048 | Yes | ||
| 143 | CUTL1 | 1424667_a_at 1424668_a_at 1425611_a_at 1425612_at 1434683_at 1437686_x_at 1440465_at 1447295_at 1451435_at | 21299 | -1.623 | -0.3896 | Yes | ||
| 144 | LFNG | 1420643_at 1449943_at | 21368 | -1.723 | -0.3753 | Yes | ||
| 145 | ABI1 | 1423177_a_at 1423178_at 1450890_a_at | 21599 | -2.263 | -0.3630 | Yes | ||
| 146 | NKTR | 1423248_at 1423249_at 1437660_at 1440804_at 1458676_at | 21610 | -2.316 | -0.3401 | Yes | ||
| 147 | HIPK1 | 1421298_a_at 1424540_at 1457515_at | 21623 | -2.346 | -0.3169 | Yes | ||
| 148 | ETS1 | 1422027_a_at 1422028_a_at 1426725_s_at 1452163_at | 21652 | -2.465 | -0.2933 | Yes | ||
| 149 | AKAP13 | 1430445_at 1433722_at 1436933_at 1439254_at 1440392_at 1440977_at 1443923_at 1445311_at 1447663_at | 21705 | -2.721 | -0.2682 | Yes | ||
| 150 | BCL6 | 1421818_at 1450381_a_at | 21747 | -2.870 | -0.2411 | Yes | ||
| 151 | PRDM2 | 1429251_at 1444123_at 1453068_at | 21763 | -2.922 | -0.2122 | Yes | ||
| 152 | ITM2A | 1423608_at 1451047_at | 21772 | -2.980 | -0.1825 | Yes | ||
| 153 | IGF2R | 1424111_at 1424112_at 1440979_at 1445966_at 1457356_at | 21786 | -3.055 | -0.1522 | Yes | ||
| 154 | IFNGR1 | 1443602_at 1448167_at | 21802 | -3.165 | -0.1209 | Yes | ||
| 155 | KLF2 | 1448890_at | 21824 | -3.304 | -0.0885 | Yes | ||
| 156 | S100A9 | 1448756_at | 21895 | -4.073 | -0.0505 | Yes | ||
| 157 | S100A8 | 1419394_s_at | 21925 | -5.183 | 0.0005 | Yes |