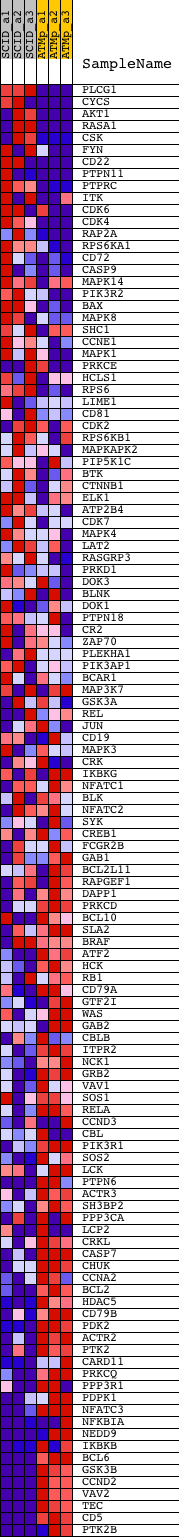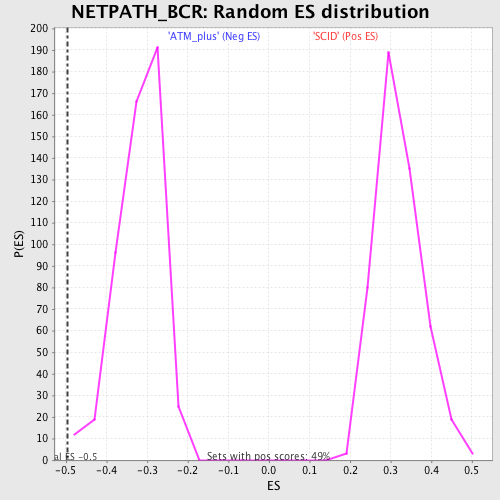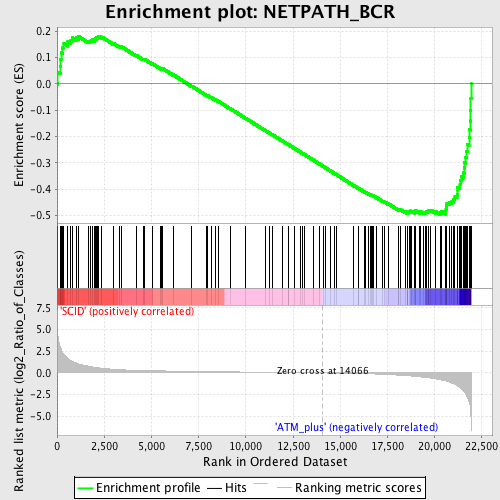
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus.phenotype_SCID_versus_ATM_plus.cls #SCID_versus_ATM_plus_repos |
| Phenotype | phenotype_SCID_versus_ATM_plus.cls#SCID_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | NETPATH_BCR |
| Enrichment Score (ES) | -0.49631324 |
| Normalized Enrichment Score (NES) | -1.5563899 |
| Nominal p-value | 0.0019646366 |
| FDR q-value | 0.3496294 |
| FWER p-Value | 1.0 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | PLCG1 | 1435149_at 1450360_at | 28 | 4.786 | 0.0437 | No | ||
| 2 | CYCS | 1422484_at 1445484_at | 164 | 3.037 | 0.0661 | No | ||
| 3 | AKT1 | 1416657_at 1425711_a_at 1440950_at 1442759_at | 182 | 2.933 | 0.0929 | No | ||
| 4 | RASA1 | 1426476_at 1426477_at 1426478_at 1438998_at | 209 | 2.769 | 0.1177 | No | ||
| 5 | CSK | 1423518_at 1439744_at | 289 | 2.344 | 0.1361 | No | ||
| 6 | FYN | 1417558_at 1441647_at 1448765_at | 360 | 2.138 | 0.1530 | No | ||
| 7 | CD22 | 1419768_at 1419769_at 1445222_at | 571 | 1.679 | 0.1592 | No | ||
| 8 | PTPN11 | 1421196_at 1427699_a_at 1451225_at | 732 | 1.415 | 0.1651 | No | ||
| 9 | PTPRC | 1422124_a_at 1440165_at | 795 | 1.337 | 0.1748 | No | ||
| 10 | ITK | 1417171_at 1430833_at 1452518_a_at 1456836_at 1457120_at | 1018 | 1.129 | 0.1753 | No | ||
| 11 | CDK6 | 1435338_at 1440040_at 1455287_at 1460291_at | 1148 | 1.022 | 0.1790 | No | ||
| 12 | CDK4 | 1422441_x_at | 1669 | 0.757 | 0.1623 | No | ||
| 13 | RAP2A | 1421574_at 1426965_at 1440473_at | 1793 | 0.707 | 0.1633 | No | ||
| 14 | RPS6KA1 | 1416896_at | 1849 | 0.689 | 0.1672 | No | ||
| 15 | CD72 | 1426112_a_at | 1975 | 0.646 | 0.1676 | No | ||
| 16 | CASP9 | 1426125_a_at 1437537_at 1457816_at | 2014 | 0.632 | 0.1718 | No | ||
| 17 | MAPK14 | 1416703_at 1416704_at 1426104_at 1442364_at 1451927_a_at 1459617_at | 2079 | 0.612 | 0.1746 | No | ||
| 18 | PIK3R2 | 1418463_at | 2146 | 0.593 | 0.1771 | No | ||
| 19 | BAX | 1416837_at | 2180 | 0.583 | 0.1811 | No | ||
| 20 | MAPK8 | 1420931_at 1420932_at 1437045_at 1440856_at 1457936_at | 2363 | 0.538 | 0.1778 | No | ||
| 21 | SHC1 | 1422853_at 1422854_at | 2961 | 0.426 | 0.1545 | No | ||
| 22 | CCNE1 | 1416492_at 1441910_x_at | 3304 | 0.375 | 0.1423 | No | ||
| 23 | MAPK1 | 1419568_at 1426585_s_at 1442876_at 1453104_at | 3406 | 0.363 | 0.1411 | No | ||
| 24 | PRKCE | 1437860_at 1437861_s_at 1442609_at 1444649_at 1449956_at 1452878_at | 4192 | 0.295 | 0.1079 | No | ||
| 25 | HCLS1 | 1418842_at | 4553 | 0.273 | 0.0940 | No | ||
| 26 | RPS6 | 1453466_at | 4629 | 0.270 | 0.0931 | No | ||
| 27 | LIME1 | 1416869_x_at 1437589_x_at 1448500_a_at | 5043 | 0.251 | 0.0765 | No | ||
| 28 | CD81 | 1416330_at | 5493 | 0.231 | 0.0581 | No | ||
| 29 | CDK2 | 1416873_a_at 1447617_at | 5549 | 0.229 | 0.0577 | No | ||
| 30 | RPS6KB1 | 1428849_at 1446376_at 1454956_at 1457562_at 1459951_at 1460705_at | 5573 | 0.228 | 0.0588 | No | ||
| 31 | MAPKAPK2 | 1426648_at | 6162 | 0.206 | 0.0338 | No | ||
| 32 | PIP5K1C | 1450228_a_at | 7133 | 0.175 | -0.0090 | No | ||
| 33 | BTK | 1422755_at | 7914 | 0.151 | -0.0434 | No | ||
| 34 | CTNNB1 | 1420811_a_at 1430533_a_at 1450008_a_at | 7984 | 0.149 | -0.0451 | No | ||
| 35 | ELK1 | 1421896_at 1421897_at 1446390_at | 8178 | 0.145 | -0.0526 | No | ||
| 36 | ATP2B4 | 1445701_at | 8375 | 0.140 | -0.0603 | No | ||
| 37 | CDK7 | 1424257_at 1439511_at 1451741_a_at 1458987_at | 8539 | 0.136 | -0.0665 | No | ||
| 38 | MAPK4 | 1435367_at | 9165 | 0.120 | -0.0940 | No | ||
| 39 | LAT2 | 1426169_a_at | 9991 | 0.100 | -0.1309 | No | ||
| 40 | RASGRP3 | 1438030_at 1438031_at | 11021 | 0.076 | -0.1773 | No | ||
| 41 | PRKD1 | 1422673_at | 11259 | 0.071 | -0.1875 | No | ||
| 42 | DOK3 | 1418096_at | 11402 | 0.068 | -0.1934 | No | ||
| 43 | BLNK | 1451780_at | 11936 | 0.054 | -0.2173 | No | ||
| 44 | DOK1 | 1417790_at | 12230 | 0.047 | -0.2303 | No | ||
| 45 | PTPN18 | 1419125_at 1444894_at | 12266 | 0.046 | -0.2315 | No | ||
| 46 | CR2 | 1425289_a_at | 12566 | 0.039 | -0.2448 | No | ||
| 47 | ZAP70 | 1422701_at 1439749_at 1440178_x_at | 12867 | 0.033 | -0.2583 | No | ||
| 48 | PLEKHA1 | 1417965_at 1443111_at | 13003 | 0.029 | -0.2642 | No | ||
| 49 | PIK3AP1 | 1421285_at 1429831_at | 13081 | 0.027 | -0.2674 | No | ||
| 50 | BCAR1 | 1439388_s_at 1450622_at | 13601 | 0.012 | -0.2911 | No | ||
| 51 | MAP3K7 | 1419988_at 1425795_a_at 1426627_at 1449693_at 1455441_at 1458787_at | 13897 | 0.005 | -0.3046 | No | ||
| 52 | GSK3A | 1435638_at | 14111 | -0.002 | -0.3144 | No | ||
| 53 | REL | 1420710_at | 14232 | -0.005 | -0.3198 | No | ||
| 54 | JUN | 1417409_at 1448694_at | 14497 | -0.014 | -0.3318 | No | ||
| 55 | CD19 | 1450570_a_at | 14678 | -0.020 | -0.3398 | No | ||
| 56 | MAPK3 | 1427060_at | 14783 | -0.022 | -0.3444 | No | ||
| 57 | CRK | 1416201_at 1425855_a_at 1436835_at 1448248_at 1460176_at | 15694 | -0.056 | -0.3856 | No | ||
| 58 | IKBKG | 1421208_at 1421209_s_at 1435646_at 1435647_at 1450161_at 1454690_at | 15968 | -0.069 | -0.3975 | No | ||
| 59 | NFATC1 | 1417621_at 1425761_a_at 1428479_at 1447084_at 1447085_s_at | 15972 | -0.069 | -0.3969 | No | ||
| 60 | BLK | 1422775_at | 16285 | -0.085 | -0.4105 | No | ||
| 61 | NFATC2 | 1425901_at 1425990_a_at 1426031_a_at 1426032_at 1439205_at 1440426_at | 16317 | -0.086 | -0.4111 | No | ||
| 62 | SYK | 1418261_at 1418262_at 1425797_a_at 1457239_at | 16497 | -0.099 | -0.4183 | No | ||
| 63 | CREB1 | 1421582_a_at 1421583_at 1423402_at 1428755_at 1452529_a_at 1452901_at | 16598 | -0.105 | -0.4219 | No | ||
| 64 | FCGR2B | 1435476_a_at 1435477_s_at 1447622_at 1451941_a_at 1455332_x_at | 16627 | -0.107 | -0.4222 | No | ||
| 65 | GAB1 | 1417693_a_at 1417694_at 1448814_at | 16677 | -0.110 | -0.4234 | No | ||
| 66 | BCL2L11 | 1426334_a_at 1435448_at 1435449_at 1456005_a_at 1456006_at 1459794_at | 16782 | -0.118 | -0.4271 | No | ||
| 67 | RAPGEF1 | 1421146_at 1427006_at 1441386_at | 16931 | -0.128 | -0.4327 | No | ||
| 68 | DAPP1 | 1421936_at 1421937_at 1447337_at | 17236 | -0.155 | -0.4451 | No | ||
| 69 | PRKCD | 1422847_a_at 1442256_at | 17365 | -0.167 | -0.4495 | No | ||
| 70 | BCL10 | 1418970_a_at 1418971_x_at 1418972_at 1443524_x_at | 17549 | -0.185 | -0.4561 | No | ||
| 71 | SLA2 | 1437504_at 1451492_at | 18091 | -0.251 | -0.4785 | No | ||
| 72 | BRAF | 1425693_at 1435434_at 1435480_at 1442749_at 1445786_at 1447940_a_at 1447941_x_at 1456505_at 1458641_at | 18093 | -0.251 | -0.4762 | No | ||
| 73 | ATF2 | 1426582_at 1426583_at 1427559_a_at 1437777_at 1452116_s_at 1459237_at | 18187 | -0.264 | -0.4780 | No | ||
| 74 | HCK | 1446553_at 1449455_at | 18436 | -0.299 | -0.4866 | No | ||
| 75 | RB1 | 1417850_at 1441494_at 1444400_at 1444454_at 1457447_at | 18543 | -0.312 | -0.4885 | No | ||
| 76 | CD79A | 1418830_at | 18548 | -0.313 | -0.4857 | No | ||
| 77 | GTF2I | 1431675_a_at 1431676_x_at | 18563 | -0.316 | -0.4834 | No | ||
| 78 | WAS | 1419631_at | 18685 | -0.334 | -0.4858 | No | ||
| 79 | GAB2 | 1419829_a_at 1420785_at 1439786_at 1446860_at | 18692 | -0.335 | -0.4829 | No | ||
| 80 | CBLB | 1437304_at 1455082_at 1458469_at | 18786 | -0.353 | -0.4839 | No | ||
| 81 | ITPR2 | 1421678_at 1424833_at 1424834_s_at 1427287_s_at 1427693_at 1444418_at 1444551_at 1447328_at | 18943 | -0.384 | -0.4874 | No | ||
| 82 | NCK1 | 1421487_a_at 1424543_at 1447271_at | 18972 | -0.389 | -0.4851 | No | ||
| 83 | GRB2 | 1418508_a_at 1449111_a_at | 18989 | -0.393 | -0.4821 | No | ||
| 84 | VAV1 | 1422932_a_at | 19214 | -0.442 | -0.4882 | No | ||
| 85 | SOS1 | 1421884_at 1421885_at 1421886_at | 19240 | -0.448 | -0.4852 | No | ||
| 86 | RELA | 1419536_a_at | 19403 | -0.485 | -0.4880 | No | ||
| 87 | CCND3 | 1415907_at 1437584_at 1444323_at 1446567_at 1447434_at 1457549_at | 19489 | -0.507 | -0.4872 | No | ||
| 88 | CBL | 1434829_at 1446608_at 1450457_at 1455886_at | 19555 | -0.527 | -0.4852 | No | ||
| 89 | PIK3R1 | 1425514_at 1425515_at 1438682_at 1444591_at 1451737_at | 19644 | -0.549 | -0.4841 | No | ||
| 90 | SOS2 | 1452281_at | 19678 | -0.557 | -0.4803 | No | ||
| 91 | LCK | 1425396_a_at 1439145_at 1439146_s_at 1457917_at | 19798 | -0.593 | -0.4802 | No | ||
| 92 | PTPN6 | 1456694_x_at 1460188_at | 20040 | -0.672 | -0.4849 | No | ||
| 93 | ACTR3 | 1426392_a_at 1434968_a_at 1452051_at | 20289 | -0.777 | -0.4890 | Yes | ||
| 94 | SH3BP2 | 1448328_at | 20365 | -0.805 | -0.4849 | Yes | ||
| 95 | PPP3CA | 1426401_at 1438478_a_at 1440051_at 1452056_s_at 1458495_at | 20580 | -0.908 | -0.4862 | Yes | ||
| 96 | LCP2 | 1418641_at 1418642_at | 20596 | -0.919 | -0.4782 | Yes | ||
| 97 | CRKL | 1421953_at 1421954_at 1425604_at 1436950_at | 20621 | -0.933 | -0.4705 | Yes | ||
| 98 | CASP7 | 1426062_a_at 1448659_at | 20638 | -0.942 | -0.4624 | Yes | ||
| 99 | CHUK | 1417091_at 1428210_s_at 1451383_a_at | 20646 | -0.945 | -0.4539 | Yes | ||
| 100 | CCNA2 | 1417910_at 1417911_at | 20779 | -1.050 | -0.4500 | Yes | ||
| 101 | BCL2 | 1422938_at 1427818_at 1437122_at 1440770_at 1443837_x_at 1457687_at | 20911 | -1.157 | -0.4452 | Yes | ||
| 102 | HDAC5 | 1415743_at | 21007 | -1.250 | -0.4378 | Yes | ||
| 103 | CD79B | 1417640_at | 21072 | -1.317 | -0.4283 | Yes | ||
| 104 | PDK2 | 1448825_at | 21182 | -1.471 | -0.4195 | Yes | ||
| 105 | ACTR2 | 1441660_at 1452587_at | 21217 | -1.518 | -0.4068 | Yes | ||
| 106 | PTK2 | 1423059_at 1430827_a_at 1439198_at 1440082_at 1441475_at 1443384_at 1445137_at | 21220 | -1.521 | -0.3926 | Yes | ||
| 107 | CARD11 | 1430257_at 1435996_at | 21337 | -1.678 | -0.3821 | Yes | ||
| 108 | PRKCQ | 1426044_a_at | 21352 | -1.700 | -0.3668 | Yes | ||
| 109 | PPP3R1 | 1421786_at 1433591_at 1450368_a_at | 21405 | -1.812 | -0.3521 | Yes | ||
| 110 | PDPK1 | 1415729_at 1416501_at 1459775_at 1459776_x_at | 21531 | -2.098 | -0.3381 | Yes | ||
| 111 | NFATC3 | 1419976_s_at 1452497_a_at | 21574 | -2.204 | -0.3193 | Yes | ||
| 112 | NFKBIA | 1420088_at 1420089_at 1438157_s_at 1448306_at 1449731_s_at | 21592 | -2.242 | -0.2990 | Yes | ||
| 113 | NEDD9 | 1422818_at 1437132_x_at 1447885_x_at 1450767_at | 21649 | -2.458 | -0.2785 | Yes | ||
| 114 | IKBKB | 1426207_at 1426333_a_at 1432275_at 1445141_at 1454184_a_at | 21683 | -2.583 | -0.2557 | Yes | ||
| 115 | BCL6 | 1421818_at 1450381_a_at | 21747 | -2.870 | -0.2316 | Yes | ||
| 116 | GSK3B | 1434439_at 1437001_at 1439931_at 1439949_at 1451020_at 1454958_at | 21817 | -3.254 | -0.2042 | Yes | ||
| 117 | CCND2 | 1416122_at 1416123_at 1416124_at 1430127_a_at 1434745_at 1448229_s_at 1455956_x_at | 21828 | -3.314 | -0.1735 | Yes | ||
| 118 | VAV2 | 1421272_at 1435065_x_at 1435244_at 1458758_at | 21875 | -3.671 | -0.1411 | Yes | ||
| 119 | TEC | 1441524_at 1460204_at | 21901 | -4.227 | -0.1025 | Yes | ||
| 120 | CD5 | 1418353_at | 21918 | -5.017 | -0.0561 | Yes | ||
| 121 | PTK2B | 1434653_at 1442437_at 1442927_at | 21933 | -6.050 | 0.0001 | Yes |