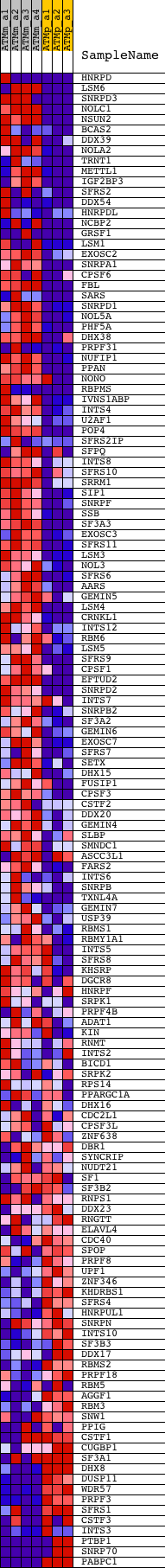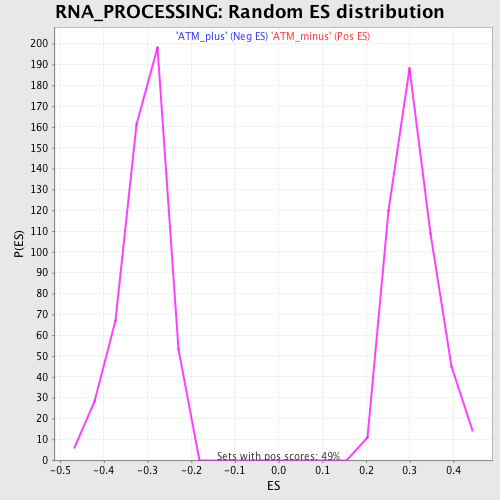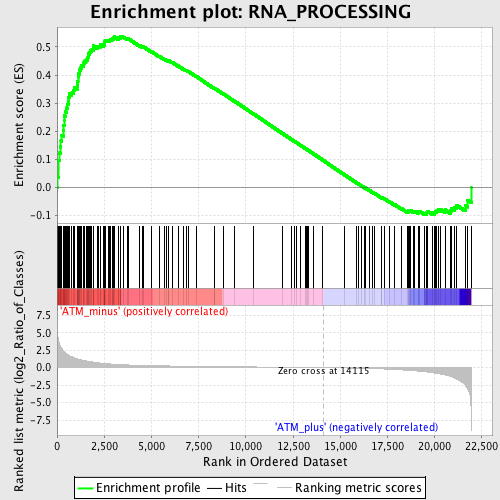
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_minus |
| GeneSet | RNA_PROCESSING |
| Enrichment Score (ES) | 0.5387197 |
| Normalized Enrichment Score (NES) | 1.7426705 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.18948118 |
| FWER p-Value | 0.571 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | HNRPD | 1425142_a_at 1446083_at | 33 | 4.652 | 0.0357 | Yes | ||
| 2 | LSM6 | 1431054_at 1437108_at 1437109_s_at 1455926_at | 69 | 3.895 | 0.0652 | Yes | ||
| 3 | SNRPD3 | 1422884_at 1422885_at 1449629_s_at | 70 | 3.889 | 0.0963 | Yes | ||
| 4 | NOLC1 | 1428869_at 1428870_at 1450087_a_at | 110 | 3.462 | 0.1222 | Yes | ||
| 5 | NSUN2 | 1423850_at 1453759_at | 175 | 3.066 | 0.1438 | Yes | ||
| 6 | BCAS2 | 1418234_s_at 1437262_x_at 1458219_at | 203 | 2.929 | 0.1660 | Yes | ||
| 7 | DDX39 | 1423643_at 1438168_x_at 1438287_x_at 1451065_a_at 1455814_x_at | 254 | 2.682 | 0.1852 | Yes | ||
| 8 | NOLA2 | 1416605_at 1416606_s_at | 322 | 2.444 | 0.2017 | Yes | ||
| 9 | TRNT1 | 1425561_at 1425562_s_at | 347 | 2.374 | 0.2195 | Yes | ||
| 10 | METTL1 | 1419122_at 1439155_at 1447683_x_at | 371 | 2.310 | 0.2370 | Yes | ||
| 11 | IGF2BP3 | 1422610_s_at 1422611_s_at 1433731_at 1433732_x_at 1436145_at 1450687_at 1453957_a_at 1459016_at | 377 | 2.290 | 0.2551 | Yes | ||
| 12 | SFRS2 | 1415807_s_at 1427504_s_at 1427816_at 1452439_s_at | 428 | 2.140 | 0.2699 | Yes | ||
| 13 | DDX54 | 1438853_x_at 1447882_x_at 1460396_at | 500 | 1.976 | 0.2824 | Yes | ||
| 14 | HNRPDL | 1420093_s_at 1420094_at 1424251_a_at 1424252_at 1428224_at 1428225_s_at 1449039_a_at 1456698_s_at | 554 | 1.864 | 0.2949 | Yes | ||
| 15 | NCBP2 | 1423045_at 1423046_s_at 1450847_at | 595 | 1.786 | 0.3074 | Yes | ||
| 16 | GRSF1 | 1433457_s_at 1433824_x_at 1437838_x_at 1439366_at | 619 | 1.748 | 0.3203 | Yes | ||
| 17 | LSM1 | 1423873_at 1441452_at | 648 | 1.707 | 0.3326 | Yes | ||
| 18 | EXOSC2 | 1426630_at 1446994_at | 787 | 1.531 | 0.3386 | Yes | ||
| 19 | SNRPA1 | 1417351_a_at 1417352_s_at 1417353_x_at | 882 | 1.425 | 0.3456 | Yes | ||
| 20 | CPSF6 | 1428231_at 1428232_at 1428233_at 1428234_at 1437372_at 1439188_at 1447381_at 1459687_x_at | 911 | 1.399 | 0.3556 | Yes | ||
| 21 | FBL | 1416684_at 1416685_s_at | 1059 | 1.245 | 0.3588 | Yes | ||
| 22 | SARS | 1426257_a_at 1452000_s_at | 1078 | 1.233 | 0.3678 | Yes | ||
| 23 | SNRPD1 | 1416336_s_at | 1086 | 1.226 | 0.3773 | Yes | ||
| 24 | NOL5A | 1426533_at 1455035_s_at | 1108 | 1.209 | 0.3860 | Yes | ||
| 25 | PHF5A | 1424170_at | 1109 | 1.206 | 0.3956 | Yes | ||
| 26 | DHX38 | 1426786_s_at 1453751_at | 1139 | 1.189 | 0.4038 | Yes | ||
| 27 | PRPF31 | 1448633_at 1453005_at 1457083_at | 1198 | 1.147 | 0.4103 | Yes | ||
| 28 | NUFIP1 | 1448802_at | 1208 | 1.143 | 0.4191 | Yes | ||
| 29 | PPAN | 1423703_at | 1260 | 1.107 | 0.4256 | Yes | ||
| 30 | NONO | 1415820_x_at 1431239_at 1448103_s_at | 1296 | 1.084 | 0.4326 | Yes | ||
| 31 | RBPMS | 1425652_s_at 1429359_s_at 1437161_x_at 1444765_at 1455936_a_at 1459078_at | 1382 | 1.023 | 0.4369 | Yes | ||
| 32 | IVNS1ABP | 1420961_a_at 1425718_a_at 1450084_s_at | 1418 | 1.007 | 0.4434 | Yes | ||
| 33 | INTS4 | 1423806_at | 1448 | 0.991 | 0.4500 | Yes | ||
| 34 | U2AF1 | 1422509_at | 1530 | 0.956 | 0.4539 | Yes | ||
| 35 | POP4 | 1448419_at | 1599 | 0.921 | 0.4582 | Yes | ||
| 36 | SFRS2IP | 1434691_at 1443085_at 1452884_at 1452885_at 1460445_at | 1629 | 0.906 | 0.4641 | Yes | ||
| 37 | SFPQ | 1423795_at 1423796_at 1436898_at 1438458_a_at 1438459_x_at 1439058_at | 1642 | 0.902 | 0.4707 | Yes | ||
| 38 | INTS8 | 1431096_at 1435250_at 1456775_at 1460609_at | 1673 | 0.891 | 0.4765 | Yes | ||
| 39 | SFRS10 | 1419543_a_at 1427432_a_at 1441466_at 1446496_at | 1734 | 0.860 | 0.4806 | Yes | ||
| 40 | SRRM1 | 1420934_a_at 1420935_a_at 1442330_at 1447447_s_at 1450045_at 1454689_at 1459828_at | 1760 | 0.848 | 0.4862 | Yes | ||
| 41 | SIP1 | 1451044_at | 1835 | 0.816 | 0.4894 | Yes | ||
| 42 | SNRPF | 1428672_at | 1907 | 0.783 | 0.4924 | Yes | ||
| 43 | SSB | 1416421_a_at 1416422_a_at 1416423_x_at 1453928_a_at | 1916 | 0.780 | 0.4982 | Yes | ||
| 44 | SF3A3 | 1423811_at | 1919 | 0.779 | 0.5044 | Yes | ||
| 45 | EXOSC3 | 1431502_a_at | 2125 | 0.705 | 0.5006 | Yes | ||
| 46 | SFRS11 | 1427269_at 1430077_at 1439095_at 1452371_at | 2213 | 0.680 | 0.5021 | Yes | ||
| 47 | LSM3 | 1448536_at | 2275 | 0.659 | 0.5045 | Yes | ||
| 48 | NOL3 | 1444786_at 1451503_at | 2308 | 0.648 | 0.5082 | Yes | ||
| 49 | SFRS6 | 1416720_at 1416721_s_at 1447898_s_at 1448454_at | 2442 | 0.612 | 0.5070 | Yes | ||
| 50 | AARS | 1423685_at 1451083_s_at | 2497 | 0.595 | 0.5093 | Yes | ||
| 51 | GEMIN5 | 1436310_at 1436311_at 1455453_at | 2500 | 0.595 | 0.5140 | Yes | ||
| 52 | LSM4 | 1448622_at | 2525 | 0.589 | 0.5176 | Yes | ||
| 53 | CRNKL1 | 1420849_at 1420850_at | 2535 | 0.588 | 0.5219 | Yes | ||
| 54 | INTS12 | 1428985_at 1452316_at | 2579 | 0.577 | 0.5245 | Yes | ||
| 55 | RBM6 | 1417213_a_at 1440223_at 1452466_a_at 1458912_at | 2724 | 0.543 | 0.5223 | Yes | ||
| 56 | LSM5 | 1418656_at | 2766 | 0.533 | 0.5247 | Yes | ||
| 57 | SFRS9 | 1417727_at | 2822 | 0.523 | 0.5263 | Yes | ||
| 58 | CPSF1 | 1417665_a_at | 2908 | 0.506 | 0.5265 | Yes | ||
| 59 | EFTUD2 | 1416557_a_at 1438835_a_at 1438836_at 1456107_x_at | 2933 | 0.499 | 0.5293 | Yes | ||
| 60 | SNRPD2 | 1452680_at | 2997 | 0.489 | 0.5304 | Yes | ||
| 61 | INTS7 | 1428531_at 1428532_at 1443045_at 1452822_at 1458590_at | 3009 | 0.487 | 0.5338 | Yes | ||
| 62 | SNRPB2 | 1426613_a_at 1452422_a_at | 3041 | 0.480 | 0.5362 | Yes | ||
| 63 | SF3A2 | 1450576_a_at 1455546_s_at | 3243 | 0.444 | 0.5305 | Yes | ||
| 64 | GEMIN6 | 1424300_at | 3267 | 0.441 | 0.5330 | Yes | ||
| 65 | EXOSC7 | 1448365_at | 3338 | 0.433 | 0.5332 | Yes | ||
| 66 | SFRS7 | 1424033_at 1424883_s_at 1436871_at | 3351 | 0.431 | 0.5361 | Yes | ||
| 67 | SETX | 1433794_at 1440459_at 1458080_at | 3370 | 0.429 | 0.5387 | Yes | ||
| 68 | DHX15 | 1416144_a_at 1416145_at | 3510 | 0.409 | 0.5356 | No | ||
| 69 | FUSIP1 | 1418526_at 1418527_a_at 1423982_at 1449121_at | 3740 | 0.383 | 0.5282 | No | ||
| 70 | CPSF3 | 1422855_at 1437328_x_at 1437852_x_at | 3760 | 0.380 | 0.5303 | No | ||
| 71 | CSTF2 | 1419644_at 1419645_at 1455523_at | 4368 | 0.323 | 0.5051 | No | ||
| 72 | DDX20 | 1416751_a_at | 4536 | 0.309 | 0.4999 | No | ||
| 73 | GEMIN4 | 1433622_at | 4567 | 0.307 | 0.5010 | No | ||
| 74 | SLBP | 1430058_at 1460168_at | 5017 | 0.282 | 0.4826 | No | ||
| 75 | SMNDC1 | 1429043_at 1452999_at | 5406 | 0.263 | 0.4669 | No | ||
| 76 | ASCC3L1 | 1454772_at 1460552_at | 5678 | 0.250 | 0.4565 | No | ||
| 77 | FARS2 | 1431354_a_at 1439406_x_at | 5791 | 0.245 | 0.4533 | No | ||
| 78 | INTS6 | 1423274_at 1423275_at 1436089_at 1443128_at | 5887 | 0.241 | 0.4509 | No | ||
| 79 | SNRPB | 1419260_a_at 1437193_s_at | 5920 | 0.240 | 0.4513 | No | ||
| 80 | TXNL4A | 1419179_at | 6102 | 0.231 | 0.4448 | No | ||
| 81 | GEMIN7 | 1435764_a_at | 6413 | 0.219 | 0.4324 | No | ||
| 82 | USP39 | 1437007_x_at 1460209_at | 6714 | 0.207 | 0.4203 | No | ||
| 83 | RBMS1 | 1418703_at 1434005_at 1441437_at 1458578_at 1459968_at | 6830 | 0.202 | 0.4166 | No | ||
| 84 | RBMY1A1 | 1420786_a_at | 6955 | 0.198 | 0.4125 | No | ||
| 85 | INTS5 | 1417887_at 1434011_a_at 1434012_at | 7360 | 0.184 | 0.3954 | No | ||
| 86 | SFRS8 | 1424997_at 1433677_at 1434966_at 1438674_a_at 1438675_at 1443905_at 1454683_at 1456880_at | 8315 | 0.155 | 0.3529 | No | ||
| 87 | KHSRP | 1428174_x_at 1436813_x_at 1452690_at | 8328 | 0.155 | 0.3536 | No | ||
| 88 | DGCR8 | 1422981_at 1435439_at 1446169_at 1455311_at | 8827 | 0.140 | 0.3319 | No | ||
| 89 | HNRPF | 1450963_at | 9383 | 0.125 | 0.3074 | No | ||
| 90 | SRPK1 | 1454042_a_at 1455107_at | 10417 | 0.098 | 0.2608 | No | ||
| 91 | PRPF4B | 1425497_a_at 1425498_at 1436427_at 1451732_at 1451909_a_at 1455696_a_at | 11921 | 0.060 | 0.1923 | No | ||
| 92 | ADAT1 | 1420748_a_at 1425390_at 1444049_at | 12437 | 0.048 | 0.1690 | No | ||
| 93 | KIN | 1427317_at | 12546 | 0.045 | 0.1644 | No | ||
| 94 | RNMT | 1436328_at 1440002_at 1446442_at 1453106_a_at | 12690 | 0.041 | 0.1582 | No | ||
| 95 | INTS2 | 1429461_at 1436247_at | 12894 | 0.036 | 0.1492 | No | ||
| 96 | BICD1 | 1427530_at 1437548_at 1438701_at 1451684_a_at | 13168 | 0.028 | 0.1369 | No | ||
| 97 | SRPK2 | 1417134_at 1417135_at 1417136_s_at 1431372_at 1435746_at 1440080_at 1442241_at 1442980_at 1447755_at 1448603_at | 13199 | 0.027 | 0.1357 | No | ||
| 98 | RPS14 | 1436586_x_at | 13283 | 0.025 | 0.1321 | No | ||
| 99 | PPARGC1A | 1434099_at 1434100_x_at 1437751_at 1456395_at 1460336_at | 13325 | 0.024 | 0.1304 | No | ||
| 100 | DHX16 | 1423925_at | 13558 | 0.017 | 0.1199 | No | ||
| 101 | CDC2L1 | 1418841_s_at | 14078 | 0.001 | 0.0961 | No | ||
| 102 | CPSF3L | 1423964_at | 15201 | -0.041 | 0.0449 | No | ||
| 103 | ZNF638 | 1417791_a_at 1417792_at 1447983_at 1458523_at | 15873 | -0.074 | 0.0147 | No | ||
| 104 | DBR1 | 1451641_at | 15959 | -0.079 | 0.0115 | No | ||
| 105 | SYNCRIP | 1422768_at 1422769_at 1426402_at 1428546_at 1450743_s_at | 16144 | -0.089 | 0.0037 | No | ||
| 106 | NUDT21 | 1417681_at 1428339_at 1437213_at 1455966_s_at | 16301 | -0.099 | -0.0026 | No | ||
| 107 | SF1 | 1422321_a_at 1423750_a_at 1423751_at 1459765_s_at 1459766_x_at | 16321 | -0.100 | -0.0027 | No | ||
| 108 | SF3B2 | 1429362_a_at 1433461_at 1438637_x_at 1453123_at 1456040_at 1456352_a_at 1459836_x_at | 16521 | -0.113 | -0.0109 | No | ||
| 109 | RNPS1 | 1437027_x_at 1437359_at 1437851_x_at 1438267_x_at 1448324_at | 16703 | -0.128 | -0.0182 | No | ||
| 110 | DDX23 | 1430050_at | 16810 | -0.137 | -0.0220 | No | ||
| 111 | RNGTT | 1425844_a_at 1448714_at | 17192 | -0.173 | -0.0381 | No | ||
| 112 | ELAVL4 | 1428741_at 1450258_a_at 1452894_at 1457399_at | 17201 | -0.174 | -0.0370 | No | ||
| 113 | CDC40 | 1442667_at 1445348_at 1453342_at 1457366_at | 17316 | -0.187 | -0.0408 | No | ||
| 114 | SPOP | 1416524_at 1416525_at 1444578_at 1447827_x_at 1455510_at 1458886_at | 17604 | -0.227 | -0.0521 | No | ||
| 115 | PRPF8 | 1422453_at | 17866 | -0.261 | -0.0620 | No | ||
| 116 | UPF1 | 1419685_at | 18235 | -0.315 | -0.0764 | No | ||
| 117 | ZNF346 | 1417088_at 1448581_at | 18558 | -0.368 | -0.0882 | No | ||
| 118 | KHDRBS1 | 1418628_at 1418630_at | 18592 | -0.374 | -0.0867 | No | ||
| 119 | SFRS4 | 1448778_at | 18616 | -0.379 | -0.0848 | No | ||
| 120 | HNRPUL1 | 1451984_at | 18655 | -0.387 | -0.0834 | No | ||
| 121 | SNRPN | 1415895_at 1415896_x_at 1432096_at 1435716_x_at | 18715 | -0.397 | -0.0830 | No | ||
| 122 | INTS10 | 1439712_at | 18875 | -0.435 | -0.0868 | No | ||
| 123 | SF3B3 | 1415985_at 1430075_at | 18945 | -0.450 | -0.0863 | No | ||
| 124 | DDX17 | 1437773_x_at 1439037_at 1452155_a_at | 19116 | -0.492 | -0.0902 | No | ||
| 125 | RBMS2 | 1423540_at 1433979_at | 19137 | -0.498 | -0.0871 | No | ||
| 126 | PRPF18 | 1429614_at | 19212 | -0.514 | -0.0864 | No | ||
| 127 | RBM5 | 1438069_a_at 1441319_at 1444618_at 1444971_at 1452186_at 1452187_at 1456262_at | 19453 | -0.579 | -0.0928 | No | ||
| 128 | AGGF1 | 1425354_a_at | 19557 | -0.609 | -0.0927 | No | ||
| 129 | RBM3 | 1429169_at | 19585 | -0.617 | -0.0890 | No | ||
| 130 | SNW1 | 1429002_at 1429003_at 1459026_at | 19636 | -0.633 | -0.0862 | No | ||
| 131 | PPIG | 1434475_at 1435341_at 1436505_at | 19857 | -0.709 | -0.0906 | No | ||
| 132 | CSTF1 | 1417117_at 1448597_at | 19987 | -0.755 | -0.0905 | No | ||
| 133 | CUGBP1 | 1423932_at 1425932_a_at 1426407_at 1426408_at 1427413_a_at | 20022 | -0.769 | -0.0859 | No | ||
| 134 | SF3A1 | 1419087_s_at 1445719_at 1446687_at 1449333_at | 20117 | -0.813 | -0.0837 | No | ||
| 135 | DHX8 | 1426629_at | 20210 | -0.852 | -0.0811 | No | ||
| 136 | DUSP11 | 1452594_at 1454785_at | 20311 | -0.899 | -0.0785 | No | ||
| 137 | WDR57 | 1428264_at 1452713_a_at | 20547 | -1.024 | -0.0811 | No | ||
| 138 | PRPF3 | 1424314_at 1457390_at | 20816 | -1.229 | -0.0836 | No | ||
| 139 | SFRS1 | 1428099_a_at 1428100_at 1430982_at 1434972_x_at 1452430_s_at 1453722_s_at 1457136_at | 20867 | -1.271 | -0.0757 | No | ||
| 140 | CSTF3 | 1424723_s_at 1443909_at | 21036 | -1.465 | -0.0717 | No | ||
| 141 | INTS3 | 1423921_at 1423922_s_at | 21162 | -1.646 | -0.0643 | No | ||
| 142 | PTBP1 | 1424874_a_at 1450443_at 1458284_at | 21611 | -2.519 | -0.0647 | No | ||
| 143 | SNRP70 | 1429009_at 1429010_at 1429011_x_at 1451104_a_at 1457047_at | 21750 | -3.026 | -0.0468 | No | ||
| 144 | PABPC1 | 1418883_a_at 1419500_at 1453840_at | 21933 | -6.913 | 0.0001 | No |