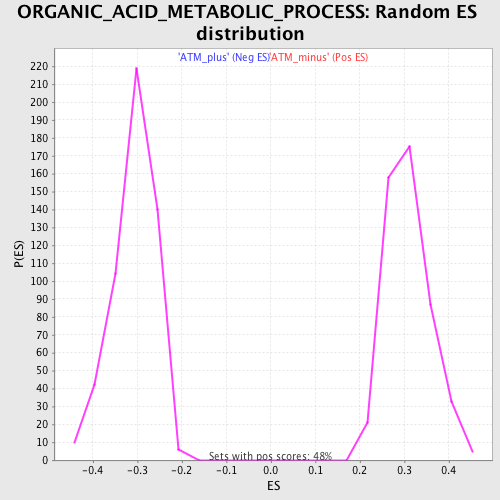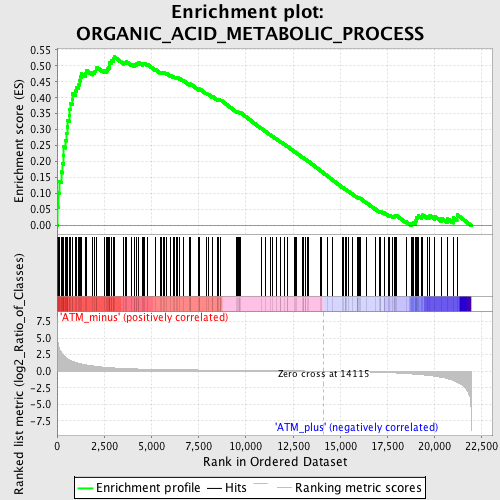
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_minus |
| GeneSet | ORGANIC_ACID_METABOLIC_PROCESS |
| Enrichment Score (ES) | 0.5277756 |
| Normalized Enrichment Score (NES) | 1.7158014 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.11407215 |
| FWER p-Value | 0.728 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ACLY | 1425326_at 1438389_x_at 1439445_x_at 1439459_x_at 1446315_at 1451666_at | 20 | 5.102 | 0.0590 | Yes | ||
| 2 | ARG1 | 1419549_at | 67 | 3.938 | 0.1031 | Yes | ||
| 3 | CPT1A | 1434866_x_at 1438156_x_at 1460409_at | 130 | 3.275 | 0.1387 | Yes | ||
| 4 | BCAT1 | 1430111_a_at 1450871_a_at | 220 | 2.860 | 0.1682 | Yes | ||
| 5 | ACOT12 | 1419395_at 1433064_at 1449457_at | 299 | 2.513 | 0.1941 | Yes | ||
| 6 | GOT2 | 1417715_a_at 1417716_at | 348 | 2.373 | 0.2198 | Yes | ||
| 7 | AGPAT2 | 1428821_at | 359 | 2.340 | 0.2468 | Yes | ||
| 8 | PYCR1 | 1424556_at | 449 | 2.082 | 0.2672 | Yes | ||
| 9 | GSTZ1 | 1427552_a_at 1458134_at | 506 | 1.961 | 0.2876 | Yes | ||
| 10 | CYP39A1 | 1418780_at | 526 | 1.916 | 0.3093 | Yes | ||
| 11 | QDPR | 1423664_at 1437993_x_at | 551 | 1.872 | 0.3301 | Yes | ||
| 12 | DARS | 1423800_at | 650 | 1.705 | 0.3457 | Yes | ||
| 13 | GAD1 | 1416561_at 1416562_at | 676 | 1.672 | 0.3641 | Yes | ||
| 14 | SLC7A5 | 1418326_at | 688 | 1.654 | 0.3831 | Yes | ||
| 15 | NFS1 | 1416373_at 1431431_a_at 1442017_at | 803 | 1.517 | 0.3956 | Yes | ||
| 16 | ACOT1 | 1449065_at | 823 | 1.488 | 0.4122 | Yes | ||
| 17 | GOT1 | 1450970_at | 968 | 1.333 | 0.4213 | Yes | ||
| 18 | PTGES3 | 1417998_at 1460221_at | 1031 | 1.279 | 0.4334 | Yes | ||
| 19 | MIF | 1416335_at 1433109_at 1454481_at | 1153 | 1.178 | 0.4417 | Yes | ||
| 20 | MTHFD2 | 1419253_at 1419254_at | 1200 | 1.145 | 0.4530 | Yes | ||
| 21 | SCYE1 | 1416486_at | 1226 | 1.129 | 0.4652 | Yes | ||
| 22 | RARS | 1416312_at | 1297 | 1.083 | 0.4747 | Yes | ||
| 23 | YARS | 1460638_at | 1507 | 0.963 | 0.4764 | Yes | ||
| 24 | SLC25A10 | 1416954_at 1416955_at | 1531 | 0.956 | 0.4865 | Yes | ||
| 25 | ECHS1 | 1452341_at | 1870 | 0.799 | 0.4804 | Yes | ||
| 26 | NANP | 1417941_at | 2003 | 0.744 | 0.4831 | Yes | ||
| 27 | SLC25A15 | 1420966_at 1420967_at | 2059 | 0.726 | 0.4891 | Yes | ||
| 28 | AS3MT | 1431980_a_at 1457201_at | 2106 | 0.712 | 0.4953 | Yes | ||
| 29 | AARS | 1423685_at 1451083_s_at | 2497 | 0.595 | 0.4844 | Yes | ||
| 30 | BRCA1 | 1424629_at 1424630_a_at 1427698_at 1443763_at 1451417_at | 2612 | 0.570 | 0.4859 | Yes | ||
| 31 | ACADS | 1460216_at | 2666 | 0.558 | 0.4900 | Yes | ||
| 32 | FADS2 | 1419031_at 1443838_x_at 1449325_at | 2705 | 0.547 | 0.4947 | Yes | ||
| 33 | GSS | 1441931_x_at 1448273_at | 2761 | 0.533 | 0.4984 | Yes | ||
| 34 | ASL | 1448350_at | 2767 | 0.532 | 0.5044 | Yes | ||
| 35 | PECI | 1431012_a_at | 2770 | 0.532 | 0.5106 | Yes | ||
| 36 | GLUD1 | 1416209_at 1448253_at | 2861 | 0.515 | 0.5125 | Yes | ||
| 37 | ACN9 | 1456614_at | 2898 | 0.508 | 0.5168 | Yes | ||
| 38 | PEPD | 1416712_at | 3002 | 0.488 | 0.5178 | Yes | ||
| 39 | ASRGL1 | 1424395_at 1424396_a_at 1460360_at | 3003 | 0.488 | 0.5235 | Yes | ||
| 40 | ALDH1L1 | 1424400_a_at 1424401_at 1456551_at | 3035 | 0.481 | 0.5278 | Yes | ||
| 41 | OXSM | 1419973_at 1429699_at 1441514_at 1449684_at 1449685_s_at 1455395_at | 3530 | 0.407 | 0.5099 | No | ||
| 42 | ACOT2 | 1422996_at 1439478_at | 3595 | 0.399 | 0.5116 | No | ||
| 43 | GCK | 1419146_a_at 1425303_at | 3680 | 0.389 | 0.5123 | No | ||
| 44 | HPD | 1424618_at | 3964 | 0.358 | 0.5035 | No | ||
| 45 | ACOX2 | 1420673_a_at | 4082 | 0.347 | 0.5022 | No | ||
| 46 | GAMT | 1422558_at | 4116 | 0.343 | 0.5047 | No | ||
| 47 | IGF1 | 1419519_at 1436497_at 1437401_at 1452014_a_at | 4190 | 0.338 | 0.5054 | No | ||
| 48 | SLC7A6 | 1433467_at 1460541_at | 4223 | 0.335 | 0.5078 | No | ||
| 49 | PPARD | 1425703_at 1439797_at | 4309 | 0.329 | 0.5078 | No | ||
| 50 | ASPA | 1418472_at | 4331 | 0.328 | 0.5107 | No | ||
| 51 | PTGES | 1439747_at 1449449_at 1449450_at | 4532 | 0.309 | 0.5051 | No | ||
| 52 | ME3 | 1429071_at | 4556 | 0.308 | 0.5077 | No | ||
| 53 | HSD3B7 | 1416968_a_at 1442912_at | 4627 | 0.304 | 0.5080 | No | ||
| 54 | BDH2 | 1453011_at | 4768 | 0.296 | 0.5051 | No | ||
| 55 | GLS2 | 1435245_at | 5210 | 0.272 | 0.4880 | No | ||
| 56 | CYP7B1 | 1421074_at 1421075_s_at 1446995_at | 5495 | 0.259 | 0.4780 | No | ||
| 57 | ALOX12 | 1422699_at 1422700_at | 5529 | 0.257 | 0.4795 | No | ||
| 58 | FPGS | 1460673_at | 5621 | 0.253 | 0.4783 | No | ||
| 59 | SMS | 1428699_at 1434190_at | 5689 | 0.249 | 0.4782 | No | ||
| 60 | FARS2 | 1431354_a_at 1439406_x_at | 5791 | 0.245 | 0.4764 | No | ||
| 61 | INDO | 1420437_at | 6002 | 0.235 | 0.4695 | No | ||
| 62 | SLC3A1 | 1448741_at | 6146 | 0.229 | 0.4657 | No | ||
| 63 | HADHB | 1437172_x_at | 6230 | 0.226 | 0.4645 | No | ||
| 64 | ALDH18A1 | 1415836_at 1435855_x_at 1437325_x_at 1437333_x_at 1437620_x_at | 6332 | 0.222 | 0.4625 | No | ||
| 65 | FTCD | 1419670_at 1457056_at | 6396 | 0.220 | 0.4622 | No | ||
| 66 | BCKDK | 1443813_x_at 1460644_at | 6504 | 0.216 | 0.4598 | No | ||
| 67 | FADS1 | 1423680_at 1440444_at | 6699 | 0.208 | 0.4533 | No | ||
| 68 | SDS | 1424744_at | 7036 | 0.195 | 0.4402 | No | ||
| 69 | NR1H4 | 1419105_at | 7058 | 0.194 | 0.4415 | No | ||
| 70 | GATM | 1423569_at | 7074 | 0.194 | 0.4431 | No | ||
| 71 | MTHFR | 1422132_at 1434087_at 1450498_at | 7502 | 0.179 | 0.4256 | No | ||
| 72 | GCLM | 1418627_at | 7529 | 0.178 | 0.4265 | No | ||
| 73 | DCT | 1418028_at | 7553 | 0.177 | 0.4275 | No | ||
| 74 | BCKDHA | 1416647_at | 7889 | 0.168 | 0.4141 | No | ||
| 75 | AGXT | 1418833_at | 7997 | 0.165 | 0.4111 | No | ||
| 76 | SLC27A5 | 1449112_at | 8216 | 0.158 | 0.4029 | No | ||
| 77 | SLC6A6 | 1420148_at 1421346_a_at 1437149_at 1449751_at | 8254 | 0.157 | 0.4031 | No | ||
| 78 | GBA2 | 1419980_at 1434271_at | 8506 | 0.149 | 0.3933 | No | ||
| 79 | GLYAT | 1424867_a_at 1424868_at 1451515_s_at | 8509 | 0.149 | 0.3950 | No | ||
| 80 | ACOT4 | 1422076_at 1422077_at | 8528 | 0.149 | 0.3959 | No | ||
| 81 | ARPP-19 | 1422609_at 1441455_at 1450685_at | 8636 | 0.145 | 0.3927 | No | ||
| 82 | BCKDHB | 1427153_at | 8679 | 0.144 | 0.3925 | No | ||
| 83 | PTGS1 | 1423414_at 1436448_a_at 1457652_x_at | 9524 | 0.121 | 0.3551 | No | ||
| 84 | DDAH1 | 1429298_at 1429299_at 1434519_at 1438879_at 1454995_at 1455400_at | 9570 | 0.120 | 0.3545 | No | ||
| 85 | PPARA | 1449051_at | 9580 | 0.120 | 0.3555 | No | ||
| 86 | PTGDS | 1423859_a_at 1423860_at | 9653 | 0.118 | 0.3535 | No | ||
| 87 | GNE | 1448810_at 1455583_at | 9684 | 0.117 | 0.3535 | No | ||
| 88 | SARS2 | 1448522_at | 9700 | 0.116 | 0.3542 | No | ||
| 89 | PFKFB1 | 1427213_at | 10817 | 0.088 | 0.3040 | No | ||
| 90 | CDO1 | 1448842_at | 11049 | 0.082 | 0.2944 | No | ||
| 91 | HGD | 1452986_at | 11284 | 0.076 | 0.2845 | No | ||
| 92 | CPT1B | 1418328_at | 11395 | 0.073 | 0.2803 | No | ||
| 93 | GGTLA1 | 1418216_at 1439420_x_at 1455747_at | 11635 | 0.067 | 0.2701 | No | ||
| 94 | SLC7A9 | 1448783_at | 11821 | 0.063 | 0.2624 | No | ||
| 95 | TYR | 1417717_a_at 1448821_at 1456095_at | 11840 | 0.062 | 0.2623 | No | ||
| 96 | MPST | 1418356_at | 12019 | 0.057 | 0.2548 | No | ||
| 97 | ALDH5A1 | 1453065_at | 12186 | 0.054 | 0.2478 | No | ||
| 98 | AKR1D1 | 1425771_at 1455100_at | 12207 | 0.053 | 0.2475 | No | ||
| 99 | MSRA | 1442385_at 1448856_a_at 1458923_at | 12566 | 0.044 | 0.2316 | No | ||
| 100 | GLDC | 1416049_at 1459959_at | 12602 | 0.043 | 0.2305 | No | ||
| 101 | SLC6A14 | 1420503_at 1420504_at | 12671 | 0.042 | 0.2278 | No | ||
| 102 | HPGD | 1419905_s_at 1419906_at | 12992 | 0.033 | 0.2135 | No | ||
| 103 | ACADSB | 1455446_x_at | 13029 | 0.032 | 0.2122 | No | ||
| 104 | MLYCD | 1449964_a_at | 13070 | 0.031 | 0.2108 | No | ||
| 105 | PRG3 | 1449924_at | 13148 | 0.028 | 0.2076 | No | ||
| 106 | SLC27A2 | 1416316_at | 13174 | 0.028 | 0.2067 | No | ||
| 107 | FAH | 1417220_at | 13255 | 0.026 | 0.2034 | No | ||
| 108 | PPARGC1A | 1434099_at 1434100_x_at 1437751_at 1456395_at 1460336_at | 13325 | 0.024 | 0.2005 | No | ||
| 109 | HAO1 | 1420420_at | 13949 | 0.005 | 0.1719 | No | ||
| 110 | ACO1 | 1423644_at 1456728_x_at | 13995 | 0.004 | 0.1699 | No | ||
| 111 | LYPLA3 | 1422341_s_at 1423704_at | 14338 | -0.008 | 0.1543 | No | ||
| 112 | BAAT | 1419001_at 1419002_s_at | 14566 | -0.017 | 0.1441 | No | ||
| 113 | IDH3B | 1418885_a_at 1418886_s_at 1441271_at | 15101 | -0.037 | 0.1200 | No | ||
| 114 | SLC7A8 | 1417929_at | 15143 | -0.039 | 0.1186 | No | ||
| 115 | GAD2 | 1421978_at | 15281 | -0.044 | 0.1128 | No | ||
| 116 | BBOX1 | 1419618_at | 15284 | -0.044 | 0.1132 | No | ||
| 117 | PAH | 1454638_a_at | 15338 | -0.047 | 0.1113 | No | ||
| 118 | MAT1A | 1423147_at | 15450 | -0.052 | 0.1068 | No | ||
| 119 | ABCD2 | 1419748_at 1438431_at 1439835_x_at 1446962_at 1456812_at | 15635 | -0.061 | 0.0991 | No | ||
| 120 | MCCC2 | 1428021_at 1432472_a_at 1454840_at | 15912 | -0.076 | 0.0873 | No | ||
| 121 | IDH1 | 1419821_s_at 1422433_s_at | 15922 | -0.077 | 0.0878 | No | ||
| 122 | GNPAT | 1417456_at | 15931 | -0.077 | 0.0884 | No | ||
| 123 | DDAH2 | 1416457_at | 15976 | -0.079 | 0.0873 | No | ||
| 124 | TST | 1448609_at | 16019 | -0.082 | 0.0863 | No | ||
| 125 | DEGS1 | 1423345_at 1423346_at | 16064 | -0.085 | 0.0853 | No | ||
| 126 | ATF4 | 1438992_x_at 1439258_at 1448135_at | 16381 | -0.105 | 0.0720 | No | ||
| 127 | HACL1 | 1449047_at | 16841 | -0.139 | 0.0526 | No | ||
| 128 | SLC7A4 | 1426068_at 1426069_s_at 1436776_x_at | 17079 | -0.162 | 0.0436 | No | ||
| 129 | ACOX3 | 1420684_at 1437352_at | 17101 | -0.164 | 0.0445 | No | ||
| 130 | SC4MOL | 1423078_a_at 1459627_at | 17140 | -0.167 | 0.0448 | No | ||
| 131 | CD74 | 1425519_a_at | 17323 | -0.188 | 0.0386 | No | ||
| 132 | ALDH4A1 | 1452375_at | 17329 | -0.188 | 0.0406 | No | ||
| 133 | SLC7A2 | 1422648_at 1426008_a_at 1436555_at 1440506_at 1450703_at | 17573 | -0.223 | 0.0320 | No | ||
| 134 | ADI1 | 1449076_x_at | 17615 | -0.228 | 0.0328 | No | ||
| 135 | ACADVL | 1424184_at | 17765 | -0.247 | 0.0289 | No | ||
| 136 | HDC | 1451796_s_at 1454713_s_at | 17848 | -0.259 | 0.0282 | No | ||
| 137 | PLOD1 | 1416289_at 1445893_at | 17870 | -0.262 | 0.0303 | No | ||
| 138 | SCLY | 1417671_at 1441192_at | 17933 | -0.269 | 0.0306 | No | ||
| 139 | CROT | 1450966_at | 17986 | -0.275 | 0.0314 | No | ||
| 140 | AGPAT1 | 1421024_at 1421025_at | 18491 | -0.355 | 0.0125 | No | ||
| 141 | GCLC | 1424296_at | 18760 | -0.407 | 0.0050 | No | ||
| 142 | GGT1 | 1448485_at | 18803 | -0.418 | 0.0079 | No | ||
| 143 | ACOT9 | 1418073_at | 18873 | -0.434 | 0.0099 | No | ||
| 144 | ADIPOR2 | 1434329_s_at 1437864_at | 18995 | -0.463 | 0.0098 | No | ||
| 145 | UGDH | 1416308_at 1447411_at | 19002 | -0.465 | 0.0149 | No | ||
| 146 | ADIPOR1 | 1424312_at 1439017_x_at 1451311_a_at | 19017 | -0.468 | 0.0198 | No | ||
| 147 | ECH1 | 1448491_at | 19018 | -0.469 | 0.0253 | No | ||
| 148 | SLC7A7 | 1417392_a_at 1447181_s_at | 19105 | -0.489 | 0.0271 | No | ||
| 149 | ALDH6A1 | 1448104_at | 19125 | -0.495 | 0.0320 | No | ||
| 150 | ACADM | 1415984_at | 19318 | -0.540 | 0.0295 | No | ||
| 151 | MAT2B | 1448196_at | 19374 | -0.553 | 0.0335 | No | ||
| 152 | ACO2 | 1436934_s_at 1442102_at 1451002_at | 19611 | -0.623 | 0.0300 | No | ||
| 153 | LTA4H | 1426807_at 1434790_a_at 1453528_at | 19745 | -0.669 | 0.0318 | No | ||
| 154 | WARS | 1415694_at 1425106_a_at 1434813_x_at 1437832_x_at | 20003 | -0.763 | 0.0289 | No | ||
| 155 | NPC1 | 1423086_at | 20366 | -0.919 | 0.0231 | No | ||
| 156 | PTS | 1450660_at 1458402_at | 20666 | -1.106 | 0.0223 | No | ||
| 157 | FAAH | 1421969_a_at 1434091_at | 20973 | -1.383 | 0.0245 | No | ||
| 158 | TNFRSF1A | 1417291_at | 21181 | -1.673 | 0.0347 | No |