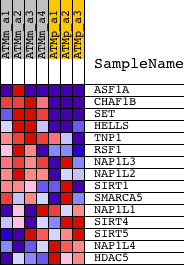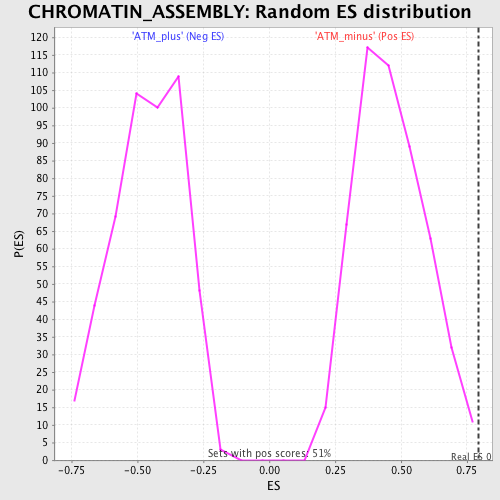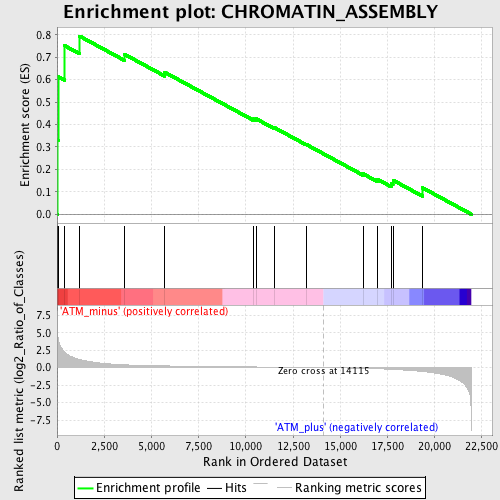
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_minus |
| GeneSet | CHROMATIN_ASSEMBLY |
| Enrichment Score (ES) | 0.7942132 |
| Normalized Enrichment Score (NES) | 1.7235199 |
| Nominal p-value | 0.0019762847 |
| FDR q-value | 0.1314193 |
| FWER p-Value | 0.676 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ASF1A | 1423511_at 1459882_at | 26 | 4.864 | 0.3315 | Yes | ||
| 2 | CHAF1B | 1423877_at 1431275_at | 56 | 4.137 | 0.6130 | Yes | ||
| 3 | SET | 1426853_at 1426854_a_at | 389 | 2.252 | 0.7519 | Yes | ||
| 4 | HELLS | 1417541_at 1430139_at 1453361_at | 1189 | 1.151 | 0.7942 | Yes | ||
| 5 | TNP1 | 1415924_at 1438632_x_at | 3578 | 0.401 | 0.7127 | No | ||
| 6 | RSF1 | 1438735_at | 5704 | 0.249 | 0.6327 | No | ||
| 7 | NAP1L3 | 1418500_at | 10407 | 0.098 | 0.4250 | No | ||
| 8 | NAP1L2 | 1418046_at | 10536 | 0.095 | 0.4256 | No | ||
| 9 | SIRT1 | 1418640_at 1458538_at | 11507 | 0.070 | 0.3862 | No | ||
| 10 | SMARCA5 | 1440048_at | 13200 | 0.027 | 0.3109 | No | ||
| 11 | NAP1L1 | 1420476_a_at 1420477_at 1420478_at 1420479_a_at 1429227_x_at 1437945_x_at 1440263_at 1452778_x_at | 16203 | -0.093 | 0.1803 | No | ||
| 12 | SIRT4 | 1426847_at | 16952 | -0.149 | 0.1563 | No | ||
| 13 | SIRT5 | 1428915_at 1428916_s_at 1432105_at | 17695 | -0.239 | 0.1388 | No | ||
| 14 | NAP1L4 | 1448476_at | 17793 | -0.251 | 0.1515 | No | ||
| 15 | HDAC5 | 1415743_at | 19351 | -0.547 | 0.1179 | No |