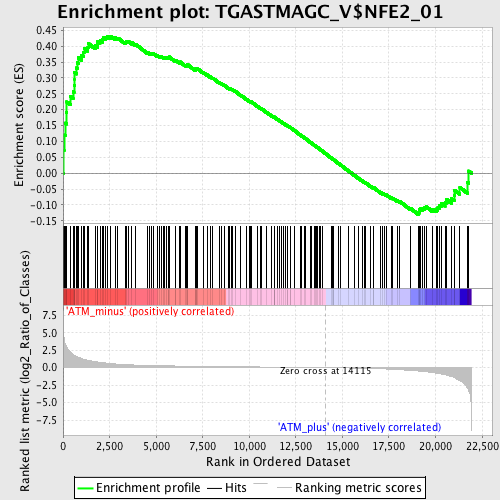
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_minus |
| GeneSet | TGASTMAGC_V$NFE2_01 |
| Enrichment Score (ES) | 0.43151912 |
| Normalized Enrichment Score (NES) | 1.4091326 |
| Nominal p-value | 0.016359918 |
| FDR q-value | 0.40480822 |
| FWER p-Value | 0.994 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ANGPTL4 | 1417130_s_at 1453410_at | 8 | 6.126 | 0.0723 | Yes | ||
| 2 | PSMD1 | 1419799_at 1429291_at | 52 | 4.209 | 0.1203 | Yes | ||
| 3 | SQSTM1 | 1440076_at 1442237_at 1444021_at 1450957_a_at | 112 | 3.412 | 0.1581 | Yes | ||
| 4 | ARHGAP8 | 1451463_at | 186 | 2.993 | 0.1902 | Yes | ||
| 5 | XPOT | 1428772_at 1428949_at 1441681_at 1441682_s_at | 202 | 2.930 | 0.2243 | Yes | ||
| 6 | ELA1 | 1423693_at | 409 | 2.190 | 0.2408 | Yes | ||
| 7 | BAG2 | 1423948_at | 536 | 1.887 | 0.2574 | Yes | ||
| 8 | ETV5 | 1420998_at 1428142_at 1450082_s_at | 588 | 1.801 | 0.2765 | Yes | ||
| 9 | SPATS2 | 1460677_at | 610 | 1.759 | 0.2964 | Yes | ||
| 10 | RBBP7 | 1415775_at 1456227_x_at | 615 | 1.752 | 0.3170 | Yes | ||
| 11 | ABCB6 | 1422524_at | 720 | 1.617 | 0.3314 | Yes | ||
| 12 | PSMD7 | 1432820_at 1451056_at 1454368_at | 758 | 1.577 | 0.3484 | Yes | ||
| 13 | PDE4D | 1443289_at 1458212_at 1459289_at 1459311_at | 815 | 1.494 | 0.3636 | Yes | ||
| 14 | PSMD12 | 1448492_a_at | 993 | 1.310 | 0.3710 | Yes | ||
| 15 | SKP1A | 1423148_at 1423149_at | 1093 | 1.219 | 0.3809 | Yes | ||
| 16 | ENO3 | 1417951_at 1449631_at | 1128 | 1.196 | 0.3935 | Yes | ||
| 17 | ABHD4 | 1416315_at 1439259_x_at | 1330 | 1.069 | 0.3970 | Yes | ||
| 18 | BLMH | 1438961_s_at 1439325_at 1452101_at | 1352 | 1.052 | 0.4085 | Yes | ||
| 19 | ALDOA | 1416921_x_at 1433604_x_at 1434799_x_at | 1722 | 0.867 | 0.4019 | Yes | ||
| 20 | GSTP1 | 1449575_a_at | 1846 | 0.811 | 0.4058 | Yes | ||
| 21 | EYA1 | 1421727_at 1443117_at 1457424_at | 1863 | 0.802 | 0.4146 | Yes | ||
| 22 | EIF4G1 | 1427036_a_at 1427037_at 1438686_at | 1981 | 0.754 | 0.4182 | Yes | ||
| 23 | E2F3 | 1427462_at 1434564_at | 2103 | 0.713 | 0.4211 | Yes | ||
| 24 | TIAL1 | 1421148_a_at 1426352_s_at 1446713_at 1452821_at 1455675_a_at 1455676_x_at | 2142 | 0.701 | 0.4277 | Yes | ||
| 25 | CD44 | 1423760_at 1434376_at 1443265_at 1452483_a_at | 2293 | 0.655 | 0.4286 | Yes | ||
| 26 | DLX1 | 1449470_at | 2391 | 0.625 | 0.4315 | Yes | ||
| 27 | SYTL1 | 1450240_a_at | 2559 | 0.581 | 0.4307 | No | ||
| 28 | ENO1 | 1419023_x_at | 2791 | 0.528 | 0.4264 | No | ||
| 29 | PSMA3 | 1448442_a_at | 2943 | 0.497 | 0.4254 | No | ||
| 30 | NEFH | 1424847_at | 3349 | 0.431 | 0.4119 | No | ||
| 31 | FOSL1 | 1417487_at 1417488_at | 3392 | 0.425 | 0.4150 | No | ||
| 32 | TTC1 | 1416994_at 1448537_at | 3503 | 0.410 | 0.4148 | No | ||
| 33 | DUSP13 | 1450198_at | 3687 | 0.388 | 0.4110 | No | ||
| 34 | ADRA1A | 1421659_at 1459351_at | 3895 | 0.365 | 0.4059 | No | ||
| 35 | ROM1 | 1448996_at | 4549 | 0.309 | 0.3796 | No | ||
| 36 | MYO10 | 1422544_at 1427363_at 1450650_at 1450651_at 1454731_at | 4664 | 0.302 | 0.3779 | No | ||
| 37 | SFTPC | 1418639_at 1438290_x_at | 4756 | 0.296 | 0.3772 | No | ||
| 38 | CDH23 | 1432346_a_at 1452028_a_at | 4848 | 0.291 | 0.3765 | No | ||
| 39 | SNCB | 1418053_at | 5069 | 0.279 | 0.3697 | No | ||
| 40 | PLEC1 | 1419835_s_at 1434610_at 1437554_at | 5167 | 0.275 | 0.3685 | No | ||
| 41 | LRRC2 | 1427388_at 1453628_s_at 1458224_at | 5276 | 0.269 | 0.3668 | No | ||
| 42 | FGF11 | 1421793_at 1439959_at | 5376 | 0.264 | 0.3654 | No | ||
| 43 | LRP1B | 1450211_at 1457235_at | 5446 | 0.261 | 0.3653 | No | ||
| 44 | RAB13 | 1424894_at | 5550 | 0.257 | 0.3636 | No | ||
| 45 | SNCG | 1417788_at | 5632 | 0.253 | 0.3629 | No | ||
| 46 | NRD1 | 1424391_at 1449716_s_at | 5645 | 0.252 | 0.3653 | No | ||
| 47 | BTK | 1422755_at | 5693 | 0.249 | 0.3661 | No | ||
| 48 | APBA2BP | 1431946_a_at 1447907_x_at | 6031 | 0.234 | 0.3534 | No | ||
| 49 | MMP19 | 1421976_at 1421977_at | 6058 | 0.232 | 0.3550 | No | ||
| 50 | SYT2 | 1420418_at 1440323_at 1449866_at | 6275 | 0.224 | 0.3477 | No | ||
| 51 | NUDT11 | 1426887_at | 6279 | 0.224 | 0.3503 | No | ||
| 52 | IL18RAP | 1421291_at 1456545_at | 6566 | 0.213 | 0.3397 | No | ||
| 53 | MFN2 | 1441424_at 1448131_at 1451900_at | 6624 | 0.211 | 0.3395 | No | ||
| 54 | USP13 | 1452442_at 1455473_at | 6670 | 0.209 | 0.3400 | No | ||
| 55 | PLCD1 | 1416675_s_at 1448432_at | 6691 | 0.208 | 0.3415 | No | ||
| 56 | SLITRK6 | 1437231_at | 7099 | 0.193 | 0.3251 | No | ||
| 57 | WDFY3 | 1427456_at 1428517_at 1445718_at 1446109_at 1447433_at 1458952_at 1459287_at 1459414_at | 7103 | 0.193 | 0.3273 | No | ||
| 58 | TLX2 | 1421770_a_at | 7154 | 0.191 | 0.3272 | No | ||
| 59 | ABCA2 | 1449302_at | 7167 | 0.191 | 0.3290 | No | ||
| 60 | DIRAS1 | 1425748_at 1451820_at 1455526_at | 7227 | 0.188 | 0.3285 | No | ||
| 61 | CRYBA2 | 1419011_at | 7530 | 0.178 | 0.3167 | No | ||
| 62 | PVALB | 1417653_at | 7734 | 0.172 | 0.3095 | No | ||
| 63 | CLDN15 | 1418920_at | 7895 | 0.167 | 0.3041 | No | ||
| 64 | GADD45G | 1453851_a_at | 8045 | 0.163 | 0.2992 | No | ||
| 65 | PRDM1 | 1420425_at 1444997_at | 8379 | 0.153 | 0.2857 | No | ||
| 66 | TNXB | 1450798_at | 8480 | 0.150 | 0.2829 | No | ||
| 67 | PHLDA2 | 1417837_at | 8662 | 0.145 | 0.2763 | No | ||
| 68 | CALB2 | 1451129_at | 8890 | 0.138 | 0.2675 | No | ||
| 69 | MT3 | 1420575_at | 8960 | 0.136 | 0.2660 | No | ||
| 70 | CRYGS | 1420453_at | 9020 | 0.135 | 0.2649 | No | ||
| 71 | LRRFIP2 | 1436378_at 1436510_a_at | 9081 | 0.133 | 0.2637 | No | ||
| 72 | P2RXL1 | 1450327_at | 9244 | 0.129 | 0.2578 | No | ||
| 73 | LIMK1 | 1417627_a_at 1425836_a_at 1456234_at 1458989_at | 9540 | 0.121 | 0.2457 | No | ||
| 74 | PSMD2 | 1415831_at | 9867 | 0.112 | 0.2321 | No | ||
| 75 | FLT1 | 1419300_at 1440926_at 1451756_at 1454037_a_at | 10018 | 0.108 | 0.2264 | No | ||
| 76 | MYBPH | 1419487_at | 10086 | 0.106 | 0.2246 | No | ||
| 77 | BNIP3 | 1422470_at | 10097 | 0.106 | 0.2254 | No | ||
| 78 | CPNE8 | 1430520_at 1430521_s_at 1431146_a_at | 10432 | 0.098 | 0.2112 | No | ||
| 79 | SH3BGRL2 | 1434109_at 1434211_at | 10606 | 0.093 | 0.2044 | No | ||
| 80 | SLC26A1 | 1451239_a_at 1458327_x_at | 10674 | 0.092 | 0.2024 | No | ||
| 81 | TUBA4 | 1417373_a_at 1417374_at 1417375_at | 10931 | 0.085 | 0.1917 | No | ||
| 82 | PPP2R2C | 1438054_x_at 1438671_at 1443804_at | 11196 | 0.078 | 0.1805 | No | ||
| 83 | FBXO2 | 1427004_at | 11327 | 0.075 | 0.1754 | No | ||
| 84 | TLL1 | 1420753_at | 11328 | 0.075 | 0.1763 | No | ||
| 85 | SMPX | 1418095_at | 11331 | 0.075 | 0.1771 | No | ||
| 86 | CPA6 | 1440617_at | 11502 | 0.070 | 0.1701 | No | ||
| 87 | SLC22A18 | 1417809_at 1427531_a_at | 11632 | 0.067 | 0.1650 | No | ||
| 88 | XLKD1 | 1429379_at 1453128_at | 11733 | 0.065 | 0.1612 | No | ||
| 89 | PLS3 | 1423725_at | 11830 | 0.062 | 0.1575 | No | ||
| 90 | HS3ST2 | 1438624_x_at 1440814_x_at 1454784_at | 11962 | 0.058 | 0.1522 | No | ||
| 91 | ADAM15 | 1416080_at 1425170_a_at 1438760_x_at 1454206_a_at | 12024 | 0.057 | 0.1501 | No | ||
| 92 | MDFI | 1420713_a_at 1425924_at | 12045 | 0.057 | 0.1498 | No | ||
| 93 | TUBB4 | 1423221_at | 12203 | 0.053 | 0.1432 | No | ||
| 94 | VAT1 | 1423726_at 1438165_x_at 1441930_x_at | 12224 | 0.053 | 0.1430 | No | ||
| 95 | ITPKC | 1440024_at 1454755_at | 12419 | 0.048 | 0.1346 | No | ||
| 96 | OMG | 1418212_at | 12769 | 0.039 | 0.1191 | No | ||
| 97 | PPP1R9B | 1447247_at 1447486_at | 12818 | 0.037 | 0.1173 | No | ||
| 98 | SNX10 | 1431055_a_at 1459764_x_at | 12944 | 0.034 | 0.1120 | No | ||
| 99 | ADCK4 | 1448490_at | 13030 | 0.032 | 0.1084 | No | ||
| 100 | MPP5 | 1421064_at | 13288 | 0.025 | 0.0969 | No | ||
| 101 | FBXO44 | 1430541_at 1434455_at | 13340 | 0.023 | 0.0949 | No | ||
| 102 | LAMA3 | 1444860_at | 13487 | 0.019 | 0.0884 | No | ||
| 103 | NR1D1 | 1421802_at 1426464_at | 13560 | 0.017 | 0.0853 | No | ||
| 104 | SLC16A6 | 1417884_at | 13601 | 0.016 | 0.0837 | No | ||
| 105 | DTNA | 1419223_a_at 1425292_at 1426066_a_at 1427588_a_at 1429768_at 1447280_at 1456069_at 1458908_at | 13636 | 0.015 | 0.0823 | No | ||
| 106 | LENEP | 1422309_a_at | 13769 | 0.010 | 0.0763 | No | ||
| 107 | SERPINB5 | 1421752_a_at 1424623_at 1438856_x_at 1441941_x_at | 13815 | 0.009 | 0.0744 | No | ||
| 108 | IL6 | 1450297_at | 13928 | 0.006 | 0.0693 | No | ||
| 109 | PROCR | 1420664_s_at | 14431 | -0.011 | 0.0464 | No | ||
| 110 | MMRN2 | 1437123_at | 14465 | -0.012 | 0.0450 | No | ||
| 111 | NUDT10 | 1435061_at | 14498 | -0.014 | 0.0437 | No | ||
| 112 | VDR | 1418175_at 1418176_at | 14514 | -0.015 | 0.0432 | No | ||
| 113 | FGF9 | 1420795_at 1438718_at 1445258_at | 14774 | -0.024 | 0.0316 | No | ||
| 114 | FBXO30 | 1429441_at 1436550_at 1453136_at 1453137_at | 14877 | -0.028 | 0.0273 | No | ||
| 115 | MAPRE3 | 1427079_at | 15350 | -0.047 | 0.0061 | No | ||
| 116 | RB1CC1 | 1418968_at 1431605_at 1449292_at | 15632 | -0.061 | -0.0060 | No | ||
| 117 | HDAC3 | 1426437_s_at | 15848 | -0.072 | -0.0150 | No | ||
| 118 | PSMD11 | 1429370_a_at 1432726_at 1456059_at 1456104_at | 16069 | -0.085 | -0.0241 | No | ||
| 119 | SYNPO | 1427045_at 1434089_at | 16183 | -0.092 | -0.0282 | No | ||
| 120 | IDS | 1421216_a_at 1427812_at 1427813_at 1434751_at 1450166_at 1455481_at | 16252 | -0.096 | -0.0302 | No | ||
| 121 | RAB3D | 1418890_a_at 1418891_a_at 1449259_at 1449260_at | 16512 | -0.113 | -0.0408 | No | ||
| 122 | NCDN | 1416852_a_at 1416853_at | 16665 | -0.125 | -0.0462 | No | ||
| 123 | MAP2K1 | 1416351_at | 16691 | -0.127 | -0.0459 | No | ||
| 124 | EIF4E | 1450909_at 1457489_at 1459371_at | 17066 | -0.160 | -0.0612 | No | ||
| 125 | AKT3 | 1422078_at 1435260_at 1435879_at 1443222_at 1460307_at | 17177 | -0.171 | -0.0642 | No | ||
| 126 | RABEP1 | 1421305_x_at 1426023_a_at 1427448_at 1443856_at 1444930_at 1459494_at | 17265 | -0.181 | -0.0660 | No | ||
| 127 | PAK6 | 1445475_at 1455200_at | 17358 | -0.192 | -0.0680 | No | ||
| 128 | MMP9 | 1416298_at 1448291_at | 17610 | -0.228 | -0.0768 | No | ||
| 129 | DTX2 | 1421720_a_at 1439429_x_at | 17707 | -0.241 | -0.0783 | No | ||
| 130 | RIT1 | 1420540_a_at 1428710_at | 17948 | -0.271 | -0.0861 | No | ||
| 131 | PFN1 | 1449018_at | 18085 | -0.291 | -0.0889 | No | ||
| 132 | SEC24D | 1426972_at 1458821_at | 18641 | -0.385 | -0.1098 | No | ||
| 133 | ABCD1 | 1418838_at | 19079 | -0.483 | -0.1242 | No | ||
| 134 | DDX17 | 1437773_x_at 1439037_at 1452155_a_at | 19116 | -0.492 | -0.1200 | No | ||
| 135 | MCART1 | 1433816_at 1454712_at | 19138 | -0.498 | -0.1150 | No | ||
| 136 | ABCF3 | 1426747_at 1426748_s_at 1452177_at | 19211 | -0.513 | -0.1123 | No | ||
| 137 | PPP2CA | 1417367_at 1444875_at 1456390_at | 19324 | -0.541 | -0.1110 | No | ||
| 138 | MDM2 | 1423605_a_at 1427718_a_at 1457929_at | 19427 | -0.568 | -0.1089 | No | ||
| 139 | PEA15 | 1416406_at 1416407_at 1459733_at | 19489 | -0.588 | -0.1047 | No | ||
| 140 | TMEM24 | 1428095_a_at 1457585_at | 19856 | -0.709 | -0.1131 | No | ||
| 141 | CMAS | 1426662_at | 20046 | -0.780 | -0.1126 | No | ||
| 142 | GADD45A | 1449519_at | 20130 | -0.816 | -0.1067 | No | ||
| 143 | CAMKK1 | 1418954_at | 20213 | -0.852 | -0.1003 | No | ||
| 144 | GABARAPL1 | 1416418_at 1416419_s_at 1442572_at | 20304 | -0.895 | -0.0938 | No | ||
| 145 | VASP | 1451097_at | 20512 | -1.003 | -0.0914 | No | ||
| 146 | RICS | 1438451_at 1447099_at | 20571 | -1.037 | -0.0818 | No | ||
| 147 | RTN3 | 1418101_a_at 1425539_a_at 1442365_at 1443220_at | 20853 | -1.260 | -0.0797 | No | ||
| 148 | BACH1 | 1440831_at 1449311_at | 21003 | -1.421 | -0.0697 | No | ||
| 149 | TUBA1 | 1418884_x_at | 21017 | -1.439 | -0.0532 | No | ||
| 150 | CLIC1 | 1416656_at 1458665_at | 21311 | -1.860 | -0.0446 | No | ||
| 151 | DHRS3 | 1448390_a_at | 21716 | -2.870 | -0.0291 | No | ||
| 152 | BAZ2A | 1427209_at 1427210_at 1438192_s_at | 21784 | -3.301 | 0.0070 | No |

