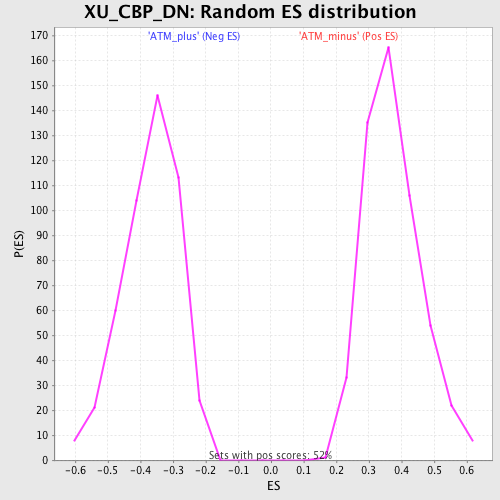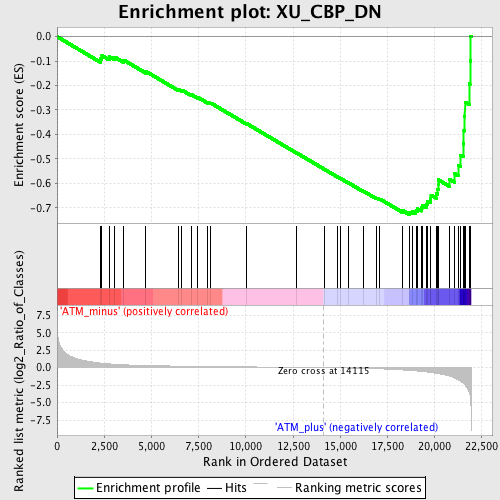
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_01_ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus.phenotype_ATM_minus_versus_ATM_plus.cls #ATM_minus_versus_ATM_plus_repos |
| Phenotype | phenotype_ATM_minus_versus_ATM_plus.cls#ATM_minus_versus_ATM_plus_repos |
| Upregulated in class | ATM_plus |
| GeneSet | XU_CBP_DN |
| Enrichment Score (ES) | -0.727614 |
| Normalized Enrichment Score (NES) | -1.9794196 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.008656621 |
| FWER p-Value | 0.0090 |

| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | ATP1A1 | 1423653_at 1435919_at 1435920_x_at 1451071_a_at 1458868_at | 2318 | 0.645 | -0.0903 | No | ||
| 2 | SLC25A3 | 1416300_a_at 1440302_at | 2374 | 0.631 | -0.0775 | No | ||
| 3 | ANP32A | 1421918_at 1434555_at 1450407_a_at | 2759 | 0.533 | -0.0822 | No | ||
| 4 | PER2 | 1417602_at 1417603_at 1445892_at 1457350_at | 3040 | 0.480 | -0.0834 | No | ||
| 5 | TCTE2 | 1425009_at 1430773_a_at 1458672_at | 3541 | 0.405 | -0.0964 | No | ||
| 6 | ANKRD17 | 1415726_at 1421220_at 1436775_a_at 1444608_at 1445826_at 1454763_at 1459952_at | 4693 | 0.300 | -0.1417 | No | ||
| 7 | LAT2 | 1426169_a_at | 6453 | 0.218 | -0.2168 | No | ||
| 8 | NUCB2 | 1418355_at | 6605 | 0.212 | -0.2186 | No | ||
| 9 | KRTAP8-1 | 1427211_at | 7104 | 0.193 | -0.2367 | No | ||
| 10 | ZFP444 | 1430851_at 1452376_at 1456154_at | 7437 | 0.181 | -0.2475 | No | ||
| 11 | ERCC2 | 1460727_at | 7989 | 0.165 | -0.2687 | No | ||
| 12 | ALDH1A3 | 1417642_at 1427395_a_at 1448789_at | 8139 | 0.160 | -0.2716 | No | ||
| 13 | GABRG3 | 1422187_at | 10025 | 0.107 | -0.3551 | No | ||
| 14 | CIITA | 1421210_at 1421211_a_at 1447075_at | 12699 | 0.041 | -0.4762 | No | ||
| 15 | IGH-1A | 1425385_a_at 1426196_at 1451632_a_at 1452535_at 1452538_at 1455530_at | 14160 | -0.002 | -0.5429 | No | ||
| 16 | FRAT2 | 1455220_at | 14829 | -0.027 | -0.5728 | No | ||
| 17 | 5830457O10RIK | 1424028_at | 15017 | -0.033 | -0.5805 | No | ||
| 18 | COTL1 | 1416001_a_at 1416002_x_at 1425801_x_at 1436236_x_at 1436838_x_at 1437811_x_at | 15417 | -0.050 | -0.5975 | No | ||
| 19 | EPHA4 | 1421928_at 1421929_at 1429021_at 1439757_s_at 1456863_at | 16244 | -0.095 | -0.6329 | No | ||
| 20 | FCER2A | 1422122_at 1451713_a_at | 16925 | -0.146 | -0.6604 | No | ||
| 21 | ARHGEF2 | 1421042_at 1421043_s_at 1427646_a_at | 17098 | -0.163 | -0.6644 | No | ||
| 22 | IGH-VJ558 | 1427839_at 1427850_x_at 1427851_x_at 1427852_x_at 1432520_at 1437346_x_at 1452574_x_at | 18285 | -0.323 | -0.7107 | No | ||
| 23 | CCR6 | 1450357_a_at | 18656 | -0.387 | -0.7182 | Yes | ||
| 24 | 9430098E02RIK | 1425191_at | 18817 | -0.421 | -0.7154 | Yes | ||
| 25 | TBC1D10A | 1448587_at | 19008 | -0.466 | -0.7128 | Yes | ||
| 26 | SDF2 | 1416857_at | 19090 | -0.485 | -0.7047 | Yes | ||
| 27 | TPR | 1426948_at 1426949_s_at 1456112_at 1456651_a_at | 19280 | -0.530 | -0.7005 | Yes | ||
| 28 | NR1H2 | 1416353_at | 19328 | -0.541 | -0.6896 | Yes | ||
| 29 | RAB14 | 1415686_at 1419243_at 1419244_a_at 1419245_at 1419246_s_at 1439687_at 1441992_at | 19555 | -0.609 | -0.6852 | Yes | ||
| 30 | LTA | 1420353_at | 19620 | -0.625 | -0.6730 | Yes | ||
| 31 | CASP9 | 1426125_a_at 1437537_at 1457816_at | 19786 | -0.679 | -0.6641 | Yes | ||
| 32 | ZFP574 | 1434128_a_at 1435202_at 1456486_at | 19801 | -0.686 | -0.6481 | Yes | ||
| 33 | MS4A4C | 1420671_x_at 1450291_s_at | 20081 | -0.796 | -0.6416 | Yes | ||
| 34 | CD97 | 1418394_a_at | 20140 | -0.821 | -0.6244 | Yes | ||
| 35 | ARF6 | 1418822_a_at 1418823_at 1418824_at | 20186 | -0.844 | -0.6060 | Yes | ||
| 36 | HSD11B1 | 1449038_at | 20191 | -0.846 | -0.5857 | Yes | ||
| 37 | RELB | 1417856_at | 20776 | -1.198 | -0.5834 | Yes | ||
| 38 | GSN | 1415812_at 1436991_x_at 1437171_x_at 1441225_at 1456312_x_at 1456568_at 1456569_x_at | 21034 | -1.462 | -0.5598 | Yes | ||
| 39 | PSCD1 | 1418183_a_at 1442189_at 1447786_at 1448997_at 1457095_at | 21246 | -1.778 | -0.5264 | Yes | ||
| 40 | ADAR | 1425405_a_at 1434268_at 1439276_at 1445596_at | 21357 | -1.927 | -0.4848 | Yes | ||
| 41 | IGH-6 | 1427329_a_at 1427351_s_at | 21520 | -2.233 | -0.4382 | Yes | ||
| 42 | DIAP1 | 1421143_at 1437821_at | 21547 | -2.283 | -0.3842 | Yes | ||
| 43 | ID3 | 1416630_at | 21590 | -2.418 | -0.3276 | Yes | ||
| 44 | ETS1 | 1422027_a_at 1422028_a_at 1426725_s_at 1452163_at | 21603 | -2.483 | -0.2681 | Yes | ||
| 45 | TGFB1 | 1420653_at 1445360_at | 21829 | -3.528 | -0.1931 | Yes | ||
| 46 | VAV2 | 1421272_at 1435065_x_at 1435244_at 1458758_at | 21871 | -3.956 | -0.0992 | Yes | ||
| 47 | CCR7 | 1423466_at | 21889 | -4.223 | 0.0021 | Yes |