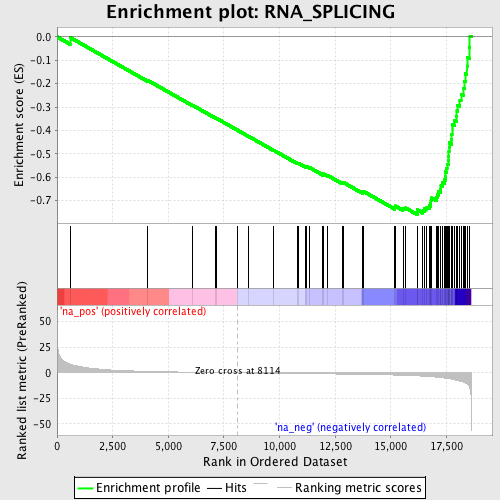
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | RNA_SPLICING |
| Enrichment Score (ES) | -0.7593775 |
| Normalized Enrichment Score (NES) | -2.4405663 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0 |
| FWER p-Value | 0.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PPARGC1A | 584 | 8.495 | -0.0026 | No | ||
| 2 | SFRS7 | 4053 | 1.412 | -0.1847 | No | ||
| 3 | PRPF4B | 6096 | 0.596 | -0.2927 | No | ||
| 4 | SRPK2 | 7114 | 0.291 | -0.3466 | No | ||
| 5 | NOL3 | 7142 | 0.283 | -0.3471 | No | ||
| 6 | CRNKL1 | 8096 | 0.004 | -0.3984 | No | ||
| 7 | GEMIN7 | 8619 | -0.135 | -0.4261 | No | ||
| 8 | ZNF638 | 9721 | -0.402 | -0.4841 | No | ||
| 9 | HNRPF | 10822 | -0.678 | -0.5411 | No | ||
| 10 | SFPQ | 10840 | -0.683 | -0.5397 | No | ||
| 11 | SNRPB2 | 11186 | -0.762 | -0.5557 | No | ||
| 12 | SFRS1 | 11226 | -0.769 | -0.5552 | No | ||
| 13 | SRRM1 | 11348 | -0.799 | -0.5590 | No | ||
| 14 | SF3B2 | 11949 | -0.956 | -0.5880 | No | ||
| 15 | NONO | 11953 | -0.957 | -0.5849 | No | ||
| 16 | SFRS2IP | 12157 | -1.014 | -0.5924 | No | ||
| 17 | SNRPN | 12824 | -1.207 | -0.6242 | No | ||
| 18 | DHX8 | 12854 | -1.215 | -0.6217 | No | ||
| 19 | SF3B4 | 13725 | -1.497 | -0.6635 | No | ||
| 20 | SNW1 | 13775 | -1.515 | -0.6609 | No | ||
| 21 | SFRS9 | 15184 | -2.148 | -0.7295 | No | ||
| 22 | SFRS8 | 15199 | -2.156 | -0.7229 | No | ||
| 23 | PTBP1 | 15547 | -2.383 | -0.7335 | No | ||
| 24 | SNRPG | 15665 | -2.464 | -0.7315 | No | ||
| 25 | DHX38 | 16184 | -2.954 | -0.7493 | Yes | ||
| 26 | SF3A3 | 16187 | -2.956 | -0.7394 | Yes | ||
| 27 | SFRS5 | 16445 | -3.226 | -0.7422 | Yes | ||
| 28 | SF3A2 | 16513 | -3.324 | -0.7345 | Yes | ||
| 29 | ASCC3L1 | 16620 | -3.478 | -0.7284 | Yes | ||
| 30 | DDX39 | 16730 | -3.654 | -0.7218 | Yes | ||
| 31 | PRPF8 | 16791 | -3.754 | -0.7123 | Yes | ||
| 32 | PRPF31 | 16804 | -3.767 | -0.7001 | Yes | ||
| 33 | SFRS10 | 16834 | -3.818 | -0.6887 | Yes | ||
| 34 | SIP1 | 17044 | -4.196 | -0.6857 | Yes | ||
| 35 | SNRPD3 | 17084 | -4.270 | -0.6732 | Yes | ||
| 36 | DHX16 | 17148 | -4.418 | -0.6616 | Yes | ||
| 37 | SNRPD2 | 17243 | -4.639 | -0.6509 | Yes | ||
| 38 | SFRS6 | 17254 | -4.671 | -0.6355 | Yes | ||
| 39 | RNPS1 | 17326 | -4.839 | -0.6229 | Yes | ||
| 40 | BCAS2 | 17405 | -5.039 | -0.6099 | Yes | ||
| 41 | SNRPF | 17457 | -5.204 | -0.5949 | Yes | ||
| 42 | DDX20 | 17463 | -5.217 | -0.5775 | Yes | ||
| 43 | NCBP2 | 17500 | -5.321 | -0.5613 | Yes | ||
| 44 | SF3B3 | 17528 | -5.426 | -0.5443 | Yes | ||
| 45 | SF3A1 | 17578 | -5.571 | -0.5279 | Yes | ||
| 46 | SYNCRIP | 17589 | -5.606 | -0.5094 | Yes | ||
| 47 | SFRS2 | 17611 | -5.660 | -0.4913 | Yes | ||
| 48 | SNRP70 | 17617 | -5.680 | -0.4722 | Yes | ||
| 49 | SNRPD1 | 17627 | -5.739 | -0.4531 | Yes | ||
| 50 | IVNS1ABP | 17729 | -6.121 | -0.4377 | Yes | ||
| 51 | CUGBP1 | 17743 | -6.188 | -0.4174 | Yes | ||
| 52 | TXNL4A | 17752 | -6.215 | -0.3966 | Yes | ||
| 53 | SNRPA1 | 17753 | -6.217 | -0.3755 | Yes | ||
| 54 | LSM4 | 17857 | -6.649 | -0.3584 | Yes | ||
| 55 | PRPF3 | 17953 | -7.160 | -0.3392 | Yes | ||
| 56 | PHF5A | 17973 | -7.243 | -0.3155 | Yes | ||
| 57 | FUSIP1 | 18008 | -7.442 | -0.2920 | Yes | ||
| 58 | U2AF1 | 18107 | -7.981 | -0.2701 | Yes | ||
| 59 | GEMIN5 | 18157 | -8.278 | -0.2446 | Yes | ||
| 60 | SNRPB | 18260 | -9.079 | -0.2192 | Yes | ||
| 61 | EFTUD2 | 18310 | -9.584 | -0.1892 | Yes | ||
| 62 | GEMIN6 | 18351 | -9.900 | -0.1576 | Yes | ||
| 63 | SRPK1 | 18424 | -10.862 | -0.1246 | Yes | ||
| 64 | USP39 | 18428 | -10.877 | -0.0877 | Yes | ||
| 65 | DBR1 | 18538 | -13.790 | -0.0466 | Yes | ||
| 66 | PPAN | 18557 | -14.912 | 0.0032 | Yes |
