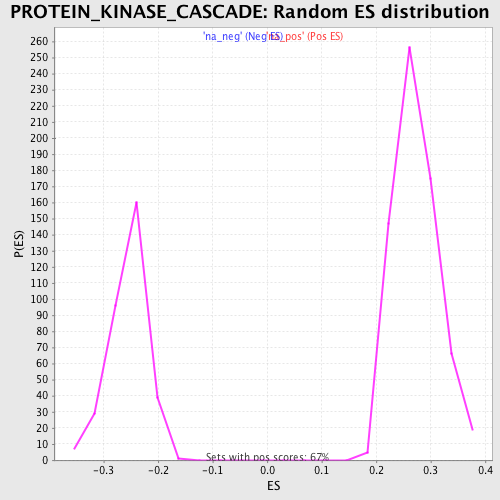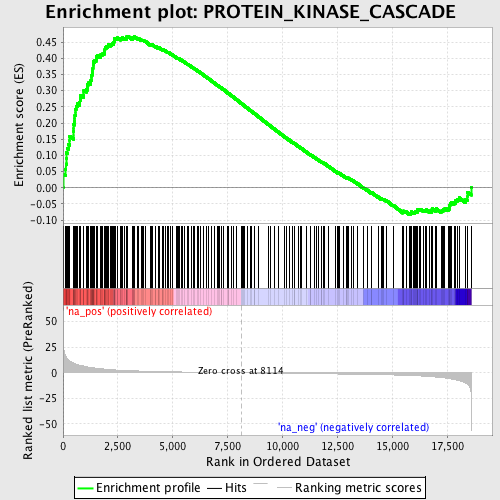
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | PROTEIN_KINASE_CASCADE |
| Enrichment Score (ES) | 0.46860275 |
| Normalized Enrichment Score (NES) | 1.7168514 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.22943498 |
| FWER p-Value | 0.946 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | STAT4 | 6 | 35.873 | 0.0425 | Yes | ||
| 2 | CD81 | 119 | 16.052 | 0.0555 | Yes | ||
| 3 | EDG5 | 131 | 15.570 | 0.0735 | Yes | ||
| 4 | CD27 | 146 | 15.080 | 0.0907 | Yes | ||
| 5 | TSPAN6 | 154 | 14.813 | 0.1080 | Yes | ||
| 6 | SQSTM1 | 189 | 13.689 | 0.1225 | Yes | ||
| 7 | MAPKAPK2 | 249 | 12.454 | 0.1341 | Yes | ||
| 8 | IFNAR2 | 295 | 11.771 | 0.1457 | Yes | ||
| 9 | STAT1 | 300 | 11.691 | 0.1594 | Yes | ||
| 10 | TNFSF10 | 470 | 9.581 | 0.1617 | Yes | ||
| 11 | PINK1 | 471 | 9.566 | 0.1731 | Yes | ||
| 12 | DUSP6 | 481 | 9.497 | 0.1839 | Yes | ||
| 13 | LITAF | 485 | 9.443 | 0.1950 | Yes | ||
| 14 | LGALS9 | 508 | 9.255 | 0.2049 | Yes | ||
| 15 | SOCS2 | 532 | 8.965 | 0.2143 | Yes | ||
| 16 | MAPK9 | 537 | 8.927 | 0.2247 | Yes | ||
| 17 | TNFAIP3 | 572 | 8.559 | 0.2331 | Yes | ||
| 18 | FYN | 575 | 8.548 | 0.2432 | Yes | ||
| 19 | TNFRSF1A | 593 | 8.427 | 0.2523 | Yes | ||
| 20 | MAP3K3 | 642 | 8.089 | 0.2593 | Yes | ||
| 21 | RPS6KA2 | 735 | 7.507 | 0.2633 | Yes | ||
| 22 | NOD1 | 769 | 7.328 | 0.2702 | Yes | ||
| 23 | STAT3 | 790 | 7.215 | 0.2777 | Yes | ||
| 24 | MAP3K11 | 801 | 7.169 | 0.2857 | Yes | ||
| 25 | IL12A | 937 | 6.582 | 0.2862 | Yes | ||
| 26 | RPS6KA5 | 945 | 6.565 | 0.2937 | Yes | ||
| 27 | STAT5A | 947 | 6.564 | 0.3015 | Yes | ||
| 28 | IKBKG | 1055 | 6.009 | 0.3028 | Yes | ||
| 29 | MAP4K2 | 1094 | 5.837 | 0.3077 | Yes | ||
| 30 | STAMBP | 1109 | 5.785 | 0.3138 | Yes | ||
| 31 | TRAF6 | 1114 | 5.768 | 0.3205 | Yes | ||
| 32 | CASP8 | 1171 | 5.513 | 0.3240 | Yes | ||
| 33 | WNK1 | 1250 | 5.279 | 0.3261 | Yes | ||
| 34 | MAP4K3 | 1264 | 5.226 | 0.3316 | Yes | ||
| 35 | RPS6KA4 | 1284 | 5.176 | 0.3368 | Yes | ||
| 36 | DUSP2 | 1286 | 5.168 | 0.3429 | Yes | ||
| 37 | RIPK1 | 1297 | 5.115 | 0.3484 | Yes | ||
| 38 | MAP3K7IP2 | 1320 | 5.047 | 0.3532 | Yes | ||
| 39 | ICK | 1339 | 4.991 | 0.3582 | Yes | ||
| 40 | LGALS1 | 1353 | 4.954 | 0.3634 | Yes | ||
| 41 | MAP3K5 | 1357 | 4.952 | 0.3692 | Yes | ||
| 42 | RGS3 | 1364 | 4.932 | 0.3747 | Yes | ||
| 43 | MAPK14 | 1366 | 4.922 | 0.3805 | Yes | ||
| 44 | HCLS1 | 1375 | 4.905 | 0.3859 | Yes | ||
| 45 | MAP4K4 | 1385 | 4.885 | 0.3913 | Yes | ||
| 46 | NDFIP2 | 1444 | 4.717 | 0.3938 | Yes | ||
| 47 | STK17B | 1501 | 4.561 | 0.3961 | Yes | ||
| 48 | CASP1 | 1513 | 4.546 | 0.4010 | Yes | ||
| 49 | PIAS1 | 1518 | 4.536 | 0.4062 | Yes | ||
| 50 | STK3 | 1581 | 4.367 | 0.4080 | Yes | ||
| 51 | PTEN | 1710 | 4.050 | 0.4059 | Yes | ||
| 52 | MAST1 | 1715 | 4.038 | 0.4105 | Yes | ||
| 53 | SH2D3C | 1755 | 3.947 | 0.4130 | Yes | ||
| 54 | SOD1 | 1813 | 3.837 | 0.4145 | Yes | ||
| 55 | TICAM2 | 1868 | 3.722 | 0.4160 | Yes | ||
| 56 | CRKL | 1876 | 3.698 | 0.4200 | Yes | ||
| 57 | SPRED2 | 1896 | 3.640 | 0.4234 | Yes | ||
| 58 | MAP3K4 | 1905 | 3.628 | 0.4272 | Yes | ||
| 59 | BIRC2 | 1910 | 3.619 | 0.4313 | Yes | ||
| 60 | CXCR4 | 1928 | 3.584 | 0.4347 | Yes | ||
| 61 | MAP3K12 | 1988 | 3.477 | 0.4356 | Yes | ||
| 62 | IKBKE | 2033 | 3.390 | 0.4373 | Yes | ||
| 63 | SOCS3 | 2048 | 3.357 | 0.4405 | Yes | ||
| 64 | PAK1 | 2078 | 3.295 | 0.4429 | Yes | ||
| 65 | F2 | 2173 | 3.151 | 0.4415 | Yes | ||
| 66 | BST2 | 2225 | 3.074 | 0.4424 | Yes | ||
| 67 | TBK1 | 2248 | 3.039 | 0.4448 | Yes | ||
| 68 | STAT2 | 2298 | 2.933 | 0.4457 | Yes | ||
| 69 | HIPK2 | 2305 | 2.926 | 0.4488 | Yes | ||
| 70 | ATP6AP2 | 2321 | 2.892 | 0.4515 | Yes | ||
| 71 | GJA1 | 2333 | 2.878 | 0.4543 | Yes | ||
| 72 | MKNK2 | 2345 | 2.853 | 0.4571 | Yes | ||
| 73 | GHRL | 2348 | 2.851 | 0.4604 | Yes | ||
| 74 | EDG2 | 2368 | 2.814 | 0.4627 | Yes | ||
| 75 | MAP3K7IP1 | 2473 | 2.687 | 0.4603 | Yes | ||
| 76 | FGF13 | 2478 | 2.679 | 0.4632 | Yes | ||
| 77 | TEC | 2604 | 2.546 | 0.4595 | Yes | ||
| 78 | DUSP16 | 2626 | 2.520 | 0.4613 | Yes | ||
| 79 | WNK3 | 2664 | 2.470 | 0.4623 | Yes | ||
| 80 | CLCF1 | 2689 | 2.444 | 0.4639 | Yes | ||
| 81 | DUSP4 | 2783 | 2.350 | 0.4616 | Yes | ||
| 82 | EGF | 2866 | 2.266 | 0.4598 | Yes | ||
| 83 | PRDX4 | 2878 | 2.254 | 0.4619 | Yes | ||
| 84 | LTBR | 2894 | 2.237 | 0.4638 | Yes | ||
| 85 | STK4 | 2902 | 2.227 | 0.4661 | Yes | ||
| 86 | SNF1LK2 | 2932 | 2.193 | 0.4671 | Yes | ||
| 87 | CCM2 | 2953 | 2.175 | 0.4686 | Yes | ||
| 88 | TLR6 | 3167 | 2.003 | 0.4594 | No | ||
| 89 | MARK2 | 3176 | 1.997 | 0.4613 | No | ||
| 90 | FPR1 | 3188 | 1.985 | 0.4631 | No | ||
| 91 | OTUD7B | 3198 | 1.979 | 0.4650 | No | ||
| 92 | TNFRSF10B | 3231 | 1.951 | 0.4656 | No | ||
| 93 | PKN1 | 3255 | 1.936 | 0.4666 | No | ||
| 94 | BCL10 | 3391 | 1.822 | 0.4614 | No | ||
| 95 | TNFSF14 | 3427 | 1.799 | 0.4617 | No | ||
| 96 | FKBP1A | 3557 | 1.708 | 0.4567 | No | ||
| 97 | C5AR1 | 3597 | 1.688 | 0.4566 | No | ||
| 98 | IL20 | 3649 | 1.659 | 0.4558 | No | ||
| 99 | RHOA | 3738 | 1.602 | 0.4529 | No | ||
| 100 | HGS | 3972 | 1.452 | 0.4420 | No | ||
| 101 | CAMKK2 | 4001 | 1.437 | 0.4422 | No | ||
| 102 | NF1 | 4013 | 1.433 | 0.4433 | No | ||
| 103 | MAPK13 | 4055 | 1.411 | 0.4427 | No | ||
| 104 | TRIP6 | 4205 | 1.331 | 0.4362 | No | ||
| 105 | MAP4K5 | 4210 | 1.327 | 0.4376 | No | ||
| 106 | MADD | 4336 | 1.256 | 0.4323 | No | ||
| 107 | ITGB1BP1 | 4347 | 1.248 | 0.4332 | No | ||
| 108 | F2R | 4379 | 1.236 | 0.4330 | No | ||
| 109 | ECT2 | 4521 | 1.172 | 0.4267 | No | ||
| 110 | PROK2 | 4542 | 1.163 | 0.4270 | No | ||
| 111 | IFNAR1 | 4558 | 1.157 | 0.4276 | No | ||
| 112 | RELA | 4669 | 1.113 | 0.4229 | No | ||
| 113 | WNK4 | 4776 | 1.071 | 0.4184 | No | ||
| 114 | SOCS6 | 4806 | 1.058 | 0.4181 | No | ||
| 115 | NEK6 | 4914 | 1.016 | 0.4135 | No | ||
| 116 | LATS2 | 4987 | 0.982 | 0.4107 | No | ||
| 117 | FAF1 | 5173 | 0.917 | 0.4018 | No | ||
| 118 | RHOC | 5219 | 0.900 | 0.4004 | No | ||
| 119 | NDFIP1 | 5237 | 0.894 | 0.4005 | No | ||
| 120 | MAPK8IP3 | 5293 | 0.869 | 0.3986 | No | ||
| 121 | MAP3K13 | 5377 | 0.837 | 0.3950 | No | ||
| 122 | CARD10 | 5440 | 0.817 | 0.3926 | No | ||
| 123 | MAP3K6 | 5537 | 0.776 | 0.3883 | No | ||
| 124 | STAT5B | 5666 | 0.724 | 0.3822 | No | ||
| 125 | IRAK1BP1 | 5718 | 0.709 | 0.3803 | No | ||
| 126 | STK38 | 5840 | 0.669 | 0.3745 | No | ||
| 127 | SHC1 | 5949 | 0.636 | 0.3694 | No | ||
| 128 | SLC20A1 | 5982 | 0.628 | 0.3684 | No | ||
| 129 | TAOK2 | 6110 | 0.593 | 0.3622 | No | ||
| 130 | SCG2 | 6171 | 0.572 | 0.3596 | No | ||
| 131 | EPGN | 6253 | 0.545 | 0.3559 | No | ||
| 132 | GNG3 | 6377 | 0.508 | 0.3498 | No | ||
| 133 | TDGF1 | 6528 | 0.460 | 0.3422 | No | ||
| 134 | ADRB3 | 6619 | 0.435 | 0.3378 | No | ||
| 135 | FYB | 6752 | 0.396 | 0.3311 | No | ||
| 136 | CHRNA7 | 6899 | 0.356 | 0.3235 | No | ||
| 137 | MAP3K10 | 7039 | 0.311 | 0.3163 | No | ||
| 138 | ADORA2B | 7076 | 0.303 | 0.3147 | No | ||
| 139 | SRPK2 | 7114 | 0.291 | 0.3131 | No | ||
| 140 | CCR2 | 7204 | 0.268 | 0.3086 | No | ||
| 141 | LYN | 7205 | 0.268 | 0.3089 | No | ||
| 142 | TRIM13 | 7320 | 0.232 | 0.3029 | No | ||
| 143 | CCL2 | 7511 | 0.172 | 0.2928 | No | ||
| 144 | DAPK1 | 7533 | 0.166 | 0.2919 | No | ||
| 145 | NMI | 7550 | 0.162 | 0.2912 | No | ||
| 146 | TRAF2 | 7662 | 0.129 | 0.2853 | No | ||
| 147 | FASLG | 7752 | 0.101 | 0.2806 | No | ||
| 148 | DUSP8 | 7882 | 0.066 | 0.2736 | No | ||
| 149 | HTR2B | 8109 | 0.001 | 0.2613 | No | ||
| 150 | DUSP9 | 8164 | -0.013 | 0.2584 | No | ||
| 151 | ADRA2A | 8206 | -0.026 | 0.2562 | No | ||
| 152 | NRTN | 8250 | -0.040 | 0.2539 | No | ||
| 153 | MAP2K4 | 8407 | -0.083 | 0.2455 | No | ||
| 154 | SGK2 | 8521 | -0.111 | 0.2395 | No | ||
| 155 | NOD2 | 8573 | -0.126 | 0.2369 | No | ||
| 156 | TGFA | 8710 | -0.158 | 0.2297 | No | ||
| 157 | NLK | 8711 | -0.158 | 0.2298 | No | ||
| 158 | GADD45G | 8887 | -0.200 | 0.2206 | No | ||
| 159 | MAP2K7 | 9344 | -0.312 | 0.1961 | No | ||
| 160 | MARK1 | 9435 | -0.334 | 0.1916 | No | ||
| 161 | DAXX | 9640 | -0.380 | 0.1809 | No | ||
| 162 | EDA2R | 9832 | -0.432 | 0.1711 | No | ||
| 163 | NEK11 | 10101 | -0.503 | 0.1571 | No | ||
| 164 | IL22RA2 | 10192 | -0.526 | 0.1528 | No | ||
| 165 | TPD52L1 | 10295 | -0.556 | 0.1479 | No | ||
| 166 | SRC | 10441 | -0.590 | 0.1407 | No | ||
| 167 | RHOH | 10540 | -0.611 | 0.1361 | No | ||
| 168 | ADRA1B | 10546 | -0.614 | 0.1366 | No | ||
| 169 | AKAP11 | 10739 | -0.657 | 0.1269 | No | ||
| 170 | STK38L | 10831 | -0.681 | 0.1228 | No | ||
| 171 | MAPK10 | 10886 | -0.694 | 0.1207 | No | ||
| 172 | MDFI | 11086 | -0.736 | 0.1107 | No | ||
| 173 | RPL17 | 11266 | -0.779 | 0.1019 | No | ||
| 174 | MAP3K7 | 11282 | -0.782 | 0.1020 | No | ||
| 175 | MYD88 | 11464 | -0.833 | 0.0931 | No | ||
| 176 | MAP3K9 | 11530 | -0.849 | 0.0906 | No | ||
| 177 | CXXC5 | 11661 | -0.881 | 0.0846 | No | ||
| 178 | CARTPT | 11791 | -0.913 | 0.0787 | No | ||
| 179 | ECM1 | 11793 | -0.914 | 0.0797 | No | ||
| 180 | FGF2 | 11888 | -0.939 | 0.0757 | No | ||
| 181 | DUSP10 | 11933 | -0.952 | 0.0744 | No | ||
| 182 | SECTM1 | 12094 | -0.997 | 0.0669 | No | ||
| 183 | IL31RA | 12404 | -1.080 | 0.0514 | No | ||
| 184 | FGFR1 | 12522 | -1.114 | 0.0463 | No | ||
| 185 | GPS2 | 12540 | -1.120 | 0.0468 | No | ||
| 186 | PLCE1 | 12606 | -1.142 | 0.0446 | No | ||
| 187 | REL | 12776 | -1.191 | 0.0368 | No | ||
| 188 | NF2 | 12930 | -1.238 | 0.0299 | No | ||
| 189 | SPRED1 | 12934 | -1.240 | 0.0313 | No | ||
| 190 | RGS4 | 12983 | -1.257 | 0.0301 | No | ||
| 191 | MINK1 | 13010 | -1.265 | 0.0302 | No | ||
| 192 | CANT1 | 13143 | -1.304 | 0.0246 | No | ||
| 193 | EDARADD | 13218 | -1.327 | 0.0222 | No | ||
| 194 | PPM1A | 13439 | -1.401 | 0.0119 | No | ||
| 195 | MBIP | 13709 | -1.491 | -0.0010 | No | ||
| 196 | GRM4 | 13852 | -1.552 | -0.0069 | No | ||
| 197 | TNFSF15 | 14047 | -1.618 | -0.0155 | No | ||
| 198 | MAPK8 | 14071 | -1.628 | -0.0148 | No | ||
| 199 | FGFR3 | 14358 | -1.747 | -0.0283 | No | ||
| 200 | TRAF7 | 14499 | -1.810 | -0.0338 | No | ||
| 201 | MAP3K2 | 14560 | -1.841 | -0.0348 | No | ||
| 202 | ADRA1A | 14609 | -1.860 | -0.0352 | No | ||
| 203 | UBE2N | 14737 | -1.915 | -0.0399 | No | ||
| 204 | ATP2C1 | 15042 | -2.070 | -0.0539 | No | ||
| 205 | MAPK8IP2 | 15488 | -2.341 | -0.0754 | No | ||
| 206 | IRAK2 | 15516 | -2.362 | -0.0740 | No | ||
| 207 | CFLAR | 15527 | -2.371 | -0.0717 | No | ||
| 208 | DAPK3 | 15631 | -2.441 | -0.0744 | No | ||
| 209 | CD40 | 15769 | -2.557 | -0.0788 | No | ||
| 210 | VAPA | 15839 | -2.623 | -0.0795 | No | ||
| 211 | BCL3 | 15860 | -2.642 | -0.0774 | No | ||
| 212 | HMOX1 | 15861 | -2.642 | -0.0743 | No | ||
| 213 | PRKAG3 | 15961 | -2.741 | -0.0764 | No | ||
| 214 | MKNK1 | 16023 | -2.805 | -0.0764 | No | ||
| 215 | NUP62 | 16047 | -2.825 | -0.0742 | No | ||
| 216 | MAPK11 | 16111 | -2.878 | -0.0742 | No | ||
| 217 | ADAM9 | 16136 | -2.907 | -0.0721 | No | ||
| 218 | SNF1LK | 16147 | -2.922 | -0.0691 | No | ||
| 219 | ERC1 | 16163 | -2.939 | -0.0664 | No | ||
| 220 | GSG2 | 16256 | -3.030 | -0.0678 | No | ||
| 221 | SCOTIN | 16289 | -3.060 | -0.0659 | No | ||
| 222 | PPP5C | 16423 | -3.196 | -0.0694 | No | ||
| 223 | ADRB2 | 16512 | -3.321 | -0.0702 | No | ||
| 224 | FLNA | 16550 | -3.370 | -0.0682 | No | ||
| 225 | UBE2V1 | 16694 | -3.590 | -0.0717 | No | ||
| 226 | DBNL | 16799 | -3.764 | -0.0729 | No | ||
| 227 | RIPK2 | 16805 | -3.768 | -0.0686 | No | ||
| 228 | DUSP22 | 16813 | -3.779 | -0.0645 | No | ||
| 229 | SOCS1 | 16975 | -4.049 | -0.0684 | No | ||
| 230 | PLK2 | 17000 | -4.104 | -0.0649 | No | ||
| 231 | DAPK2 | 17227 | -4.598 | -0.0717 | No | ||
| 232 | CD24 | 17278 | -4.710 | -0.0688 | No | ||
| 233 | IRAK1 | 17337 | -4.869 | -0.0661 | No | ||
| 234 | GADD45B | 17402 | -5.031 | -0.0636 | No | ||
| 235 | ZDHHC13 | 17552 | -5.503 | -0.0652 | No | ||
| 236 | CHUK | 17594 | -5.618 | -0.0607 | No | ||
| 237 | GPR89A | 17604 | -5.643 | -0.0544 | No | ||
| 238 | ADRA2C | 17636 | -5.763 | -0.0493 | No | ||
| 239 | MALT1 | 17693 | -5.961 | -0.0452 | No | ||
| 240 | PIK3CB | 17817 | -6.520 | -0.0441 | No | ||
| 241 | GPS1 | 17896 | -6.857 | -0.0402 | No | ||
| 242 | EEF1D | 17960 | -7.198 | -0.0350 | No | ||
| 243 | FADD | 18058 | -7.765 | -0.0310 | No | ||
| 244 | MAP4K1 | 18350 | -9.884 | -0.0351 | No | ||
| 245 | TRAF5 | 18418 | -10.781 | -0.0259 | No | ||
| 246 | SRPK1 | 18424 | -10.862 | -0.0132 | No | ||
| 247 | BLK | 18596 | -19.781 | 0.0011 | No |