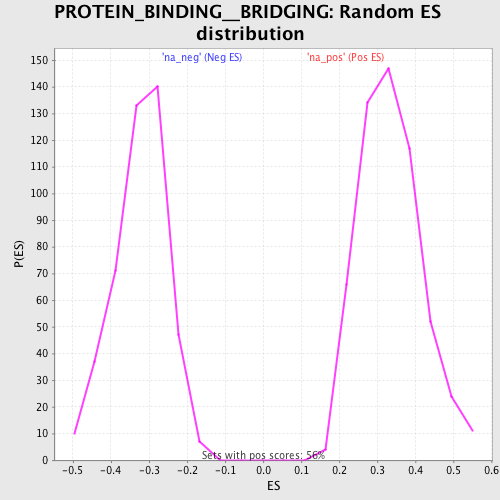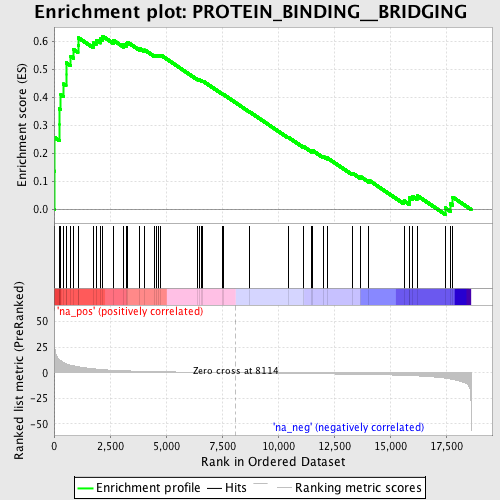
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | PROTEIN_BINDING__BRIDGING |
| Enrichment Score (ES) | 0.61800057 |
| Normalized Enrichment Score (NES) | 1.8499155 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.062338747 |
| FWER p-Value | 0.378 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | VAV3 | 12 | 30.494 | 0.1374 | Yes | ||
| 2 | GAB1 | 22 | 26.495 | 0.2568 | Yes | ||
| 3 | GRB7 | 241 | 12.611 | 0.3021 | Yes | ||
| 4 | BLNK | 244 | 12.553 | 0.3588 | Yes | ||
| 5 | LASP1 | 301 | 11.682 | 0.4087 | Yes | ||
| 6 | LAT | 424 | 10.053 | 0.4476 | Yes | ||
| 7 | SOCS2 | 532 | 8.965 | 0.4824 | Yes | ||
| 8 | SH2D2A | 533 | 8.965 | 0.5230 | Yes | ||
| 9 | SH3BP2 | 742 | 7.475 | 0.5457 | Yes | ||
| 10 | SH3BGRL | 872 | 6.868 | 0.5698 | Yes | ||
| 11 | ANXA1 | 1067 | 5.961 | 0.5863 | Yes | ||
| 12 | STAM | 1088 | 5.872 | 0.6118 | Yes | ||
| 13 | SH2D3C | 1755 | 3.947 | 0.5938 | Yes | ||
| 14 | CRKL | 1876 | 3.698 | 0.6041 | Yes | ||
| 15 | SHB | 2075 | 3.302 | 0.6084 | Yes | ||
| 16 | ARHGAP4 | 2163 | 3.163 | 0.6180 | Yes | ||
| 17 | CHN2 | 2649 | 2.487 | 0.6031 | No | ||
| 18 | ARHGAP1 | 3104 | 2.051 | 0.5880 | No | ||
| 19 | ABI2 | 3220 | 1.958 | 0.5906 | No | ||
| 20 | IRS4 | 3283 | 1.913 | 0.5959 | No | ||
| 21 | ARHGAP6 | 3820 | 1.542 | 0.5741 | No | ||
| 22 | GRAP2 | 4040 | 1.421 | 0.5687 | No | ||
| 23 | EVPL | 4499 | 1.184 | 0.5494 | No | ||
| 24 | GRB14 | 4588 | 1.148 | 0.5498 | No | ||
| 25 | GRB2 | 4679 | 1.109 | 0.5500 | No | ||
| 26 | RUSC1 | 4758 | 1.081 | 0.5507 | No | ||
| 27 | ITSN2 | 6414 | 0.495 | 0.4638 | No | ||
| 28 | CHN1 | 6480 | 0.474 | 0.4625 | No | ||
| 29 | LOR | 6588 | 0.444 | 0.4587 | No | ||
| 30 | SPRR1B | 6639 | 0.430 | 0.4580 | No | ||
| 31 | SH3BGR | 7519 | 0.169 | 0.4114 | No | ||
| 32 | ARPC4 | 7574 | 0.152 | 0.4092 | No | ||
| 33 | SNX9 | 8719 | -0.160 | 0.3483 | No | ||
| 34 | SRC | 10441 | -0.590 | 0.2583 | No | ||
| 35 | CNKSR1 | 11115 | -0.743 | 0.2254 | No | ||
| 36 | SLA | 11503 | -0.843 | 0.2083 | No | ||
| 37 | SKAP2 | 11542 | -0.853 | 0.2102 | No | ||
| 38 | SPRR1A | 12012 | -0.973 | 0.1893 | No | ||
| 39 | IVL | 12191 | -1.022 | 0.1843 | No | ||
| 40 | GRB10 | 13328 | -1.365 | 0.1293 | No | ||
| 41 | CNTNAP1 | 13697 | -1.489 | 0.1162 | No | ||
| 42 | EPS8 | 14048 | -1.618 | 0.1047 | No | ||
| 43 | KHDRBS1 | 15619 | -2.432 | 0.0312 | No | ||
| 44 | BCL3 | 15860 | -2.642 | 0.0302 | No | ||
| 45 | GRAP | 15873 | -2.654 | 0.0416 | No | ||
| 46 | RAD50 | 16004 | -2.788 | 0.0472 | No | ||
| 47 | STUB1 | 16203 | -2.973 | 0.0500 | No | ||
| 48 | TOB1 | 17459 | -5.211 | 0.0060 | No | ||
| 49 | FKBP4 | 17696 | -5.983 | 0.0203 | No | ||
| 50 | ST13 | 17794 | -6.443 | 0.0443 | No |