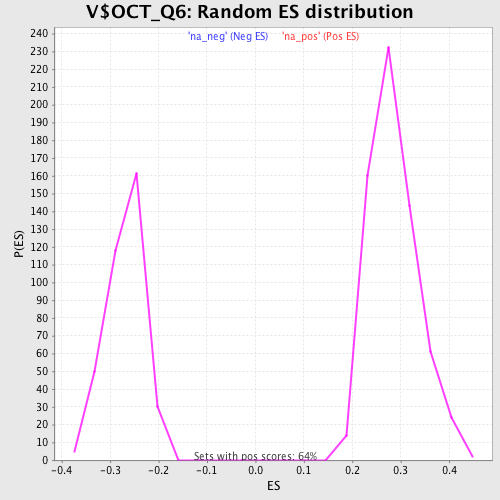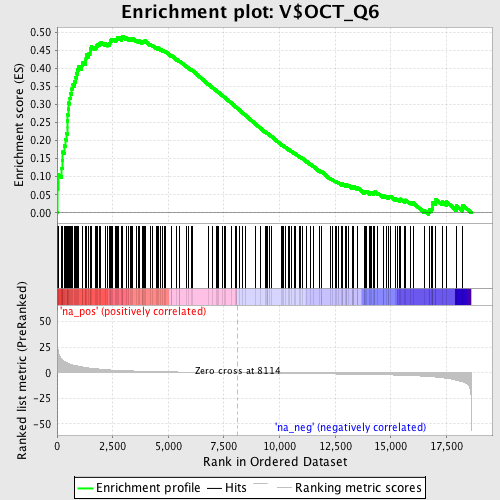
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | V$OCT_Q6 |
| Enrichment Score (ES) | 0.48919982 |
| Normalized Enrichment Score (NES) | 1.7129254 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.046031877 |
| FWER p-Value | 0.38 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | STAT4 | 6 | 35.873 | 0.0671 | Yes | ||
| 2 | HOXA7 | 40 | 22.177 | 0.1070 | Yes | ||
| 3 | VPREB3 | 210 | 13.301 | 0.1228 | Yes | ||
| 4 | GPRC5B | 234 | 12.715 | 0.1454 | Yes | ||
| 5 | BLNK | 244 | 12.553 | 0.1685 | Yes | ||
| 6 | CYR61 | 321 | 11.284 | 0.1856 | Yes | ||
| 7 | AXUD1 | 369 | 10.727 | 0.2032 | Yes | ||
| 8 | TCEAL1 | 411 | 10.196 | 0.2202 | Yes | ||
| 9 | RHOB | 456 | 9.755 | 0.2361 | Yes | ||
| 10 | TSGA14 | 457 | 9.754 | 0.2544 | Yes | ||
| 11 | DUSP6 | 481 | 9.497 | 0.2710 | Yes | ||
| 12 | SLC25A12 | 497 | 9.354 | 0.2878 | Yes | ||
| 13 | PDE4D | 515 | 9.182 | 0.3041 | Yes | ||
| 14 | SCML4 | 573 | 8.556 | 0.3171 | Yes | ||
| 15 | DLX1 | 603 | 8.339 | 0.3312 | Yes | ||
| 16 | HOXB4 | 664 | 7.921 | 0.3429 | Yes | ||
| 17 | PCYT1B | 710 | 7.648 | 0.3548 | Yes | ||
| 18 | MSI2 | 786 | 7.244 | 0.3643 | Yes | ||
| 19 | THRA | 838 | 6.988 | 0.3747 | Yes | ||
| 20 | SH3BGRL | 872 | 6.868 | 0.3858 | Yes | ||
| 21 | DMD | 908 | 6.710 | 0.3965 | Yes | ||
| 22 | ATF7IP | 969 | 6.391 | 0.4053 | Yes | ||
| 23 | FCHSD1 | 1118 | 5.758 | 0.4081 | Yes | ||
| 24 | H3F3B | 1142 | 5.607 | 0.4174 | Yes | ||
| 25 | SH3GL3 | 1266 | 5.222 | 0.4205 | Yes | ||
| 26 | SERTAD4 | 1296 | 5.117 | 0.4285 | Yes | ||
| 27 | THRAP5 | 1306 | 5.093 | 0.4376 | Yes | ||
| 28 | RRAS | 1428 | 4.766 | 0.4400 | Yes | ||
| 29 | DLGAP4 | 1481 | 4.635 | 0.4459 | Yes | ||
| 30 | TPD52 | 1486 | 4.625 | 0.4544 | Yes | ||
| 31 | DPYSL2 | 1527 | 4.522 | 0.4607 | Yes | ||
| 32 | PTEN | 1710 | 4.050 | 0.4584 | Yes | ||
| 33 | HIST1H2BM | 1759 | 3.939 | 0.4632 | Yes | ||
| 34 | KIF13A | 1830 | 3.795 | 0.4666 | Yes | ||
| 35 | HIST1H2BC | 1902 | 3.631 | 0.4696 | Yes | ||
| 36 | SREBF2 | 1968 | 3.502 | 0.4726 | Yes | ||
| 37 | ARHGAP4 | 2163 | 3.163 | 0.4680 | Yes | ||
| 38 | NFYB | 2280 | 2.975 | 0.4673 | Yes | ||
| 39 | ETV1 | 2339 | 2.859 | 0.4696 | Yes | ||
| 40 | EYA1 | 2380 | 2.802 | 0.4727 | Yes | ||
| 41 | LPL | 2393 | 2.780 | 0.4772 | Yes | ||
| 42 | NFIA | 2432 | 2.728 | 0.4803 | Yes | ||
| 43 | GRIA3 | 2509 | 2.650 | 0.4812 | Yes | ||
| 44 | HOXB6 | 2638 | 2.506 | 0.4789 | Yes | ||
| 45 | HIST1H2BK | 2684 | 2.447 | 0.4811 | Yes | ||
| 46 | HIST1H2BB | 2693 | 2.439 | 0.4852 | Yes | ||
| 47 | HOXC5 | 2779 | 2.353 | 0.4850 | Yes | ||
| 48 | HIST1H3B | 2912 | 2.212 | 0.4820 | Yes | ||
| 49 | TBXAS1 | 2924 | 2.198 | 0.4856 | Yes | ||
| 50 | HIST1H2AH | 2934 | 2.192 | 0.4892 | Yes | ||
| 51 | RAB26 | 3096 | 2.058 | 0.4843 | No | ||
| 52 | HIST1H2AB | 3212 | 1.962 | 0.4818 | No | ||
| 53 | ALK | 3304 | 1.893 | 0.4804 | No | ||
| 54 | MAB21L2 | 3350 | 1.851 | 0.4814 | No | ||
| 55 | PIGT | 3381 | 1.827 | 0.4833 | No | ||
| 56 | TCF12 | 3555 | 1.709 | 0.4771 | No | ||
| 57 | DPYSL3 | 3678 | 1.635 | 0.4735 | No | ||
| 58 | UPK3A | 3689 | 1.628 | 0.4761 | No | ||
| 59 | BTK | 3816 | 1.546 | 0.4721 | No | ||
| 60 | HIST2H2AC | 3835 | 1.536 | 0.4740 | No | ||
| 61 | DNAH5 | 3892 | 1.503 | 0.4738 | No | ||
| 62 | GAB2 | 3908 | 1.495 | 0.4758 | No | ||
| 63 | HIST2H2BE | 3952 | 1.465 | 0.4762 | No | ||
| 64 | CRISP1 | 4180 | 1.343 | 0.4665 | No | ||
| 65 | TLL2 | 4291 | 1.282 | 0.4629 | No | ||
| 66 | SILV | 4451 | 1.209 | 0.4565 | No | ||
| 67 | PAM | 4517 | 1.173 | 0.4552 | No | ||
| 68 | SLC6A15 | 4535 | 1.167 | 0.4565 | No | ||
| 69 | CRYZL1 | 4666 | 1.114 | 0.4515 | No | ||
| 70 | SPRR2B | 4752 | 1.084 | 0.4490 | No | ||
| 71 | ADORA2A | 4811 | 1.057 | 0.4478 | No | ||
| 72 | NXPH1 | 4869 | 1.034 | 0.4467 | No | ||
| 73 | ANXA2 | 5130 | 0.934 | 0.4343 | No | ||
| 74 | BAT2 | 5144 | 0.928 | 0.4353 | No | ||
| 75 | ADORA1 | 5387 | 0.834 | 0.4238 | No | ||
| 76 | HIST1H2AC | 5485 | 0.799 | 0.4200 | No | ||
| 77 | NRL | 5514 | 0.786 | 0.4200 | No | ||
| 78 | FGF12 | 5820 | 0.676 | 0.4047 | No | ||
| 79 | COL23A1 | 5909 | 0.645 | 0.4012 | No | ||
| 80 | TCF2 | 6021 | 0.614 | 0.3963 | No | ||
| 81 | LRRN1 | 6077 | 0.600 | 0.3944 | No | ||
| 82 | HOXB8 | 6805 | 0.381 | 0.3557 | No | ||
| 83 | ASCL3 | 6991 | 0.326 | 0.3463 | No | ||
| 84 | TSC1 | 7178 | 0.276 | 0.3367 | No | ||
| 85 | LYN | 7205 | 0.268 | 0.3358 | No | ||
| 86 | NEUROG1 | 7251 | 0.253 | 0.3339 | No | ||
| 87 | STX12 | 7415 | 0.204 | 0.3254 | No | ||
| 88 | POU2F3 | 7537 | 0.165 | 0.3192 | No | ||
| 89 | HPCAL4 | 7588 | 0.148 | 0.3167 | No | ||
| 90 | TMOD2 | 7826 | 0.082 | 0.3040 | No | ||
| 91 | HIST1H3C | 7999 | 0.031 | 0.2948 | No | ||
| 92 | VSNL1 | 8042 | 0.018 | 0.2925 | No | ||
| 93 | CNNM4 | 8072 | 0.011 | 0.2910 | No | ||
| 94 | PFKFB1 | 8179 | -0.019 | 0.2852 | No | ||
| 95 | ITPR3 | 8344 | -0.069 | 0.2765 | No | ||
| 96 | ARF6 | 8478 | -0.100 | 0.2695 | No | ||
| 97 | CLCA3 | 8908 | -0.207 | 0.2466 | No | ||
| 98 | SFRS14 | 9136 | -0.260 | 0.2348 | No | ||
| 99 | PRRX1 | 9347 | -0.312 | 0.2240 | No | ||
| 100 | PAX6 | 9397 | -0.326 | 0.2219 | No | ||
| 101 | HOXA10 | 9464 | -0.340 | 0.2190 | No | ||
| 102 | LIPG | 9472 | -0.342 | 0.2192 | No | ||
| 103 | TOP1 | 9529 | -0.354 | 0.2169 | No | ||
| 104 | EPHB3 | 9634 | -0.379 | 0.2119 | No | ||
| 105 | IRX4 | 10079 | -0.499 | 0.1888 | No | ||
| 106 | ALDH1A1 | 10135 | -0.511 | 0.1868 | No | ||
| 107 | FZD4 | 10175 | -0.521 | 0.1856 | No | ||
| 108 | HIST1H2AA | 10276 | -0.551 | 0.1812 | No | ||
| 109 | WNT6 | 10421 | -0.586 | 0.1745 | No | ||
| 110 | OTX2 | 10429 | -0.588 | 0.1753 | No | ||
| 111 | UBE2S | 10534 | -0.611 | 0.1708 | No | ||
| 112 | GPC4 | 10652 | -0.639 | 0.1656 | No | ||
| 113 | HDAC9 | 10695 | -0.647 | 0.1646 | No | ||
| 114 | FOXP1 | 10892 | -0.694 | 0.1552 | No | ||
| 115 | SRF | 10949 | -0.705 | 0.1535 | No | ||
| 116 | RPP21 | 11030 | -0.723 | 0.1505 | No | ||
| 117 | EGR2 | 11225 | -0.769 | 0.1415 | No | ||
| 118 | SLITRK6 | 11372 | -0.806 | 0.1351 | No | ||
| 119 | HOXD11 | 11519 | -0.847 | 0.1287 | No | ||
| 120 | MGAT5B | 11781 | -0.911 | 0.1163 | No | ||
| 121 | NDNL2 | 11883 | -0.938 | 0.1126 | No | ||
| 122 | GPR85 | 11894 | -0.942 | 0.1138 | No | ||
| 123 | FSTL5 | 12269 | -1.043 | 0.0955 | No | ||
| 124 | NRXN3 | 12368 | -1.068 | 0.0922 | No | ||
| 125 | GPR4 | 12497 | -1.108 | 0.0873 | No | ||
| 126 | MRAS | 12558 | -1.126 | 0.0862 | No | ||
| 127 | SYNPR | 12651 | -1.156 | 0.0834 | No | ||
| 128 | REL | 12776 | -1.191 | 0.0789 | No | ||
| 129 | SCOC | 12807 | -1.200 | 0.0795 | No | ||
| 130 | BDNF | 12829 | -1.208 | 0.0806 | No | ||
| 131 | TCERG1L | 12976 | -1.255 | 0.0751 | No | ||
| 132 | GPM6A | 12986 | -1.258 | 0.0769 | No | ||
| 133 | ZHX2 | 13005 | -1.264 | 0.0783 | No | ||
| 134 | MTUS1 | 13089 | -1.287 | 0.0763 | No | ||
| 135 | PTHLH | 13276 | -1.347 | 0.0687 | No | ||
| 136 | ITSN1 | 13278 | -1.348 | 0.0712 | No | ||
| 137 | CDX4 | 13305 | -1.356 | 0.0723 | No | ||
| 138 | FZD2 | 13325 | -1.363 | 0.0739 | No | ||
| 139 | CDX2 | 13481 | -1.416 | 0.0681 | No | ||
| 140 | SESN3 | 13502 | -1.425 | 0.0697 | No | ||
| 141 | FOXP2 | 13796 | -1.525 | 0.0567 | No | ||
| 142 | HIST1H2BA | 13834 | -1.541 | 0.0576 | No | ||
| 143 | SFRP2 | 13861 | -1.556 | 0.0591 | No | ||
| 144 | CSF3 | 13921 | -1.574 | 0.0588 | No | ||
| 145 | EHF | 14062 | -1.624 | 0.0543 | No | ||
| 146 | EAF2 | 14105 | -1.644 | 0.0551 | No | ||
| 147 | FBXO24 | 14147 | -1.658 | 0.0560 | No | ||
| 148 | FGF20 | 14237 | -1.693 | 0.0544 | No | ||
| 149 | POU3F4 | 14260 | -1.702 | 0.0564 | No | ||
| 150 | MAP2K6 | 14286 | -1.709 | 0.0582 | No | ||
| 151 | LHX6 | 14416 | -1.771 | 0.0546 | No | ||
| 152 | PRICKLE2 | 14657 | -1.882 | 0.0451 | No | ||
| 153 | FGF14 | 14660 | -1.883 | 0.0485 | No | ||
| 154 | PRDM1 | 14786 | -1.938 | 0.0454 | No | ||
| 155 | NRXN1 | 14903 | -2.001 | 0.0428 | No | ||
| 156 | MID1 | 14907 | -2.003 | 0.0464 | No | ||
| 157 | TRERF1 | 14999 | -2.046 | 0.0453 | No | ||
| 158 | SP6 | 15189 | -2.150 | 0.0391 | No | ||
| 159 | COL25A1 | 15294 | -2.213 | 0.0377 | No | ||
| 160 | LRCH4 | 15409 | -2.286 | 0.0358 | No | ||
| 161 | SEMA6C | 15444 | -2.317 | 0.0383 | No | ||
| 162 | DAPK3 | 15631 | -2.441 | 0.0328 | No | ||
| 163 | PCDH8 | 15668 | -2.466 | 0.0355 | No | ||
| 164 | BCDO2 | 15874 | -2.654 | 0.0293 | No | ||
| 165 | ATP2A2 | 16007 | -2.790 | 0.0274 | No | ||
| 166 | E2F3 | 16497 | -3.306 | 0.0071 | No | ||
| 167 | RTTN | 16715 | -3.619 | 0.0021 | No | ||
| 168 | TRAF3 | 16727 | -3.648 | 0.0084 | No | ||
| 169 | NR6A1 | 16824 | -3.805 | 0.0103 | No | ||
| 170 | LDB2 | 16861 | -3.859 | 0.0156 | No | ||
| 171 | DSCR1 | 16867 | -3.870 | 0.0226 | No | ||
| 172 | CDK2 | 16879 | -3.891 | 0.0293 | No | ||
| 173 | TWIST1 | 17022 | -4.143 | 0.0294 | No | ||
| 174 | NRAS | 17023 | -4.143 | 0.0372 | No | ||
| 175 | KCNN3 | 17310 | -4.792 | 0.0307 | No | ||
| 176 | CD86 | 17499 | -5.321 | 0.0305 | No | ||
| 177 | PIM2 | 17951 | -7.139 | 0.0195 | No | ||
| 178 | TSNAX | 18223 | -8.802 | 0.0213 | No |