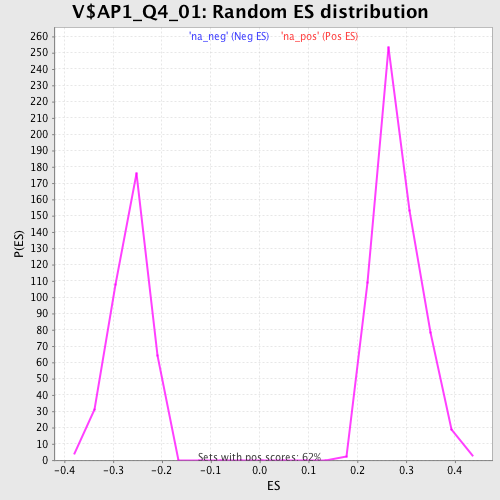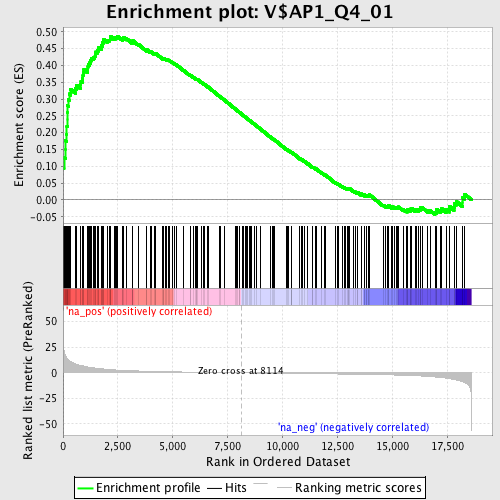
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | V$AP1_Q4_01 |
| Enrichment Score (ES) | 0.48729005 |
| Normalized Enrichment Score (NES) | 1.7317394 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.05486509 |
| FWER p-Value | 0.318 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | EMP1 | 0 | 60.375 | 0.0946 | Yes | ||
| 2 | PPP1R9B | 41 | 21.764 | 0.1266 | Yes | ||
| 3 | CDKN1A | 100 | 17.229 | 0.1504 | Yes | ||
| 4 | SYTL1 | 104 | 16.860 | 0.1767 | Yes | ||
| 5 | ELK3 | 166 | 14.407 | 0.1960 | Yes | ||
| 6 | BCL9L | 173 | 14.167 | 0.2179 | Yes | ||
| 7 | SQSTM1 | 189 | 13.689 | 0.2385 | Yes | ||
| 8 | ITM2B | 190 | 13.683 | 0.2600 | Yes | ||
| 9 | HSPB8 | 197 | 13.504 | 0.2808 | Yes | ||
| 10 | PSTPIP1 | 236 | 12.693 | 0.2986 | Yes | ||
| 11 | GADD45A | 276 | 12.024 | 0.3154 | Yes | ||
| 12 | BACH1 | 342 | 11.060 | 0.3292 | Yes | ||
| 13 | PKP3 | 560 | 8.734 | 0.3311 | Yes | ||
| 14 | DDIT3 | 618 | 8.219 | 0.3409 | Yes | ||
| 15 | ETV5 | 784 | 7.256 | 0.3433 | Yes | ||
| 16 | CLIC1 | 812 | 7.082 | 0.3529 | Yes | ||
| 17 | ATP1B1 | 875 | 6.834 | 0.3603 | Yes | ||
| 18 | LAMC1 | 903 | 6.725 | 0.3694 | Yes | ||
| 19 | RAB3D | 914 | 6.690 | 0.3793 | Yes | ||
| 20 | PEA15 | 946 | 6.565 | 0.3879 | Yes | ||
| 21 | RIT1 | 1112 | 5.776 | 0.3880 | Yes | ||
| 22 | RNF34 | 1131 | 5.681 | 0.3959 | Yes | ||
| 23 | H3F3B | 1142 | 5.607 | 0.4042 | Yes | ||
| 24 | ITPKC | 1218 | 5.364 | 0.4085 | Yes | ||
| 25 | ADAM15 | 1259 | 5.240 | 0.4146 | Yes | ||
| 26 | RPS6KA4 | 1284 | 5.176 | 0.4214 | Yes | ||
| 27 | HCLS1 | 1375 | 4.905 | 0.4242 | Yes | ||
| 28 | SNX10 | 1451 | 4.700 | 0.4275 | Yes | ||
| 29 | MYB | 1483 | 4.633 | 0.4331 | Yes | ||
| 30 | AKT3 | 1488 | 4.622 | 0.4401 | Yes | ||
| 31 | RGS2 | 1552 | 4.449 | 0.4436 | Yes | ||
| 32 | PADI4 | 1597 | 4.328 | 0.4480 | Yes | ||
| 33 | CMAS | 1624 | 4.247 | 0.4533 | Yes | ||
| 34 | ABHD4 | 1743 | 3.970 | 0.4531 | Yes | ||
| 35 | ROM1 | 1757 | 3.940 | 0.4586 | Yes | ||
| 36 | GTPBP2 | 1786 | 3.900 | 0.4632 | Yes | ||
| 37 | MAMDC2 | 1796 | 3.865 | 0.4687 | Yes | ||
| 38 | ITGB4 | 1838 | 3.779 | 0.4724 | Yes | ||
| 39 | SLC35B1 | 1852 | 3.759 | 0.4776 | Yes | ||
| 40 | LTBP3 | 2008 | 3.448 | 0.4746 | Yes | ||
| 41 | BAZ2A | 2096 | 3.277 | 0.4750 | Yes | ||
| 42 | DHRS3 | 2137 | 3.202 | 0.4779 | Yes | ||
| 43 | EPHB2 | 2148 | 3.187 | 0.4823 | Yes | ||
| 44 | ABCD1 | 2167 | 3.159 | 0.4863 | Yes | ||
| 45 | GJA1 | 2333 | 2.878 | 0.4819 | Yes | ||
| 46 | USP2 | 2397 | 2.776 | 0.4828 | Yes | ||
| 47 | PPP2CA | 2450 | 2.707 | 0.4842 | Yes | ||
| 48 | AP2M1 | 2472 | 2.687 | 0.4873 | Yes | ||
| 49 | SSH2 | 2709 | 2.422 | 0.4783 | No | ||
| 50 | TNXB | 2718 | 2.414 | 0.4816 | No | ||
| 51 | VDR | 2762 | 2.369 | 0.4830 | No | ||
| 52 | TMEM24 | 2897 | 2.234 | 0.4792 | No | ||
| 53 | CSF1R | 3150 | 2.015 | 0.4687 | No | ||
| 54 | FABP4 | 3171 | 2.001 | 0.4708 | No | ||
| 55 | UBE2E3 | 3179 | 1.994 | 0.4735 | No | ||
| 56 | USP3 | 3442 | 1.789 | 0.4621 | No | ||
| 57 | BTK | 3816 | 1.546 | 0.4443 | No | ||
| 58 | IL11 | 3819 | 1.542 | 0.4466 | No | ||
| 59 | BLMH | 3986 | 1.445 | 0.4399 | No | ||
| 60 | LAMC2 | 4019 | 1.430 | 0.4404 | No | ||
| 61 | MAPRE3 | 4155 | 1.354 | 0.4352 | No | ||
| 62 | COL16A1 | 4193 | 1.337 | 0.4352 | No | ||
| 63 | MAP4K5 | 4210 | 1.327 | 0.4365 | No | ||
| 64 | PRX | 4548 | 1.162 | 0.4200 | No | ||
| 65 | HOXA11 | 4567 | 1.154 | 0.4208 | No | ||
| 66 | AQP5 | 4584 | 1.150 | 0.4218 | No | ||
| 67 | RELA | 4669 | 1.113 | 0.4189 | No | ||
| 68 | SYNGR1 | 4729 | 1.092 | 0.4175 | No | ||
| 69 | CD68 | 4781 | 1.068 | 0.4164 | No | ||
| 70 | SV2B | 4828 | 1.050 | 0.4155 | No | ||
| 71 | VIT | 4980 | 0.986 | 0.4089 | No | ||
| 72 | CHST4 | 5085 | 0.950 | 0.4047 | No | ||
| 73 | CALB2 | 5158 | 0.923 | 0.4022 | No | ||
| 74 | MMP9 | 5497 | 0.795 | 0.3851 | No | ||
| 75 | IGSF9 | 5792 | 0.685 | 0.3703 | No | ||
| 76 | FGF12 | 5820 | 0.676 | 0.3699 | No | ||
| 77 | HDAC3 | 5951 | 0.636 | 0.3638 | No | ||
| 78 | SLC38A3 | 6011 | 0.616 | 0.3616 | No | ||
| 79 | FBXW11 | 6082 | 0.599 | 0.3587 | No | ||
| 80 | GTF2B | 6097 | 0.596 | 0.3589 | No | ||
| 81 | SLC26A1 | 6130 | 0.584 | 0.3581 | No | ||
| 82 | SLC24A4 | 6312 | 0.529 | 0.3491 | No | ||
| 83 | IBRDC2 | 6417 | 0.495 | 0.3442 | No | ||
| 84 | PLCD1 | 6427 | 0.493 | 0.3445 | No | ||
| 85 | LOR | 6588 | 0.444 | 0.3365 | No | ||
| 86 | CRYBA2 | 6608 | 0.438 | 0.3361 | No | ||
| 87 | SRPK2 | 7114 | 0.291 | 0.3092 | No | ||
| 88 | CHST1 | 7172 | 0.276 | 0.3065 | No | ||
| 89 | FBXO44 | 7343 | 0.226 | 0.2977 | No | ||
| 90 | DYRK1A | 7838 | 0.077 | 0.2710 | No | ||
| 91 | SHC3 | 7883 | 0.066 | 0.2687 | No | ||
| 92 | SH3RF2 | 7966 | 0.041 | 0.2643 | No | ||
| 93 | LMOD3 | 8031 | 0.022 | 0.2609 | No | ||
| 94 | SNCB | 8035 | 0.021 | 0.2607 | No | ||
| 95 | SYT2 | 8190 | -0.021 | 0.2524 | No | ||
| 96 | FBS1 | 8217 | -0.028 | 0.2511 | No | ||
| 97 | DIRAS1 | 8303 | -0.056 | 0.2465 | No | ||
| 98 | MYH14 | 8343 | -0.067 | 0.2445 | No | ||
| 99 | LAMA3 | 8386 | -0.078 | 0.2424 | No | ||
| 100 | GNAI1 | 8493 | -0.104 | 0.2368 | No | ||
| 101 | AK5 | 8560 | -0.121 | 0.2334 | No | ||
| 102 | KCNA2 | 8584 | -0.128 | 0.2323 | No | ||
| 103 | UCN2 | 8740 | -0.166 | 0.2242 | No | ||
| 104 | ESRRG | 8830 | -0.188 | 0.2196 | No | ||
| 105 | COL7A1 | 9013 | -0.234 | 0.2101 | No | ||
| 106 | MARK1 | 9435 | -0.334 | 0.1878 | No | ||
| 107 | LRRTM3 | 9559 | -0.362 | 0.1817 | No | ||
| 108 | EDG1 | 9584 | -0.368 | 0.1810 | No | ||
| 109 | RIN1 | 9648 | -0.382 | 0.1782 | No | ||
| 110 | HCN3 | 10174 | -0.521 | 0.1505 | No | ||
| 111 | NR0B2 | 10212 | -0.531 | 0.1493 | No | ||
| 112 | MNT | 10284 | -0.553 | 0.1463 | No | ||
| 113 | ELAVL2 | 10390 | -0.579 | 0.1415 | No | ||
| 114 | ANXA7 | 10420 | -0.585 | 0.1409 | No | ||
| 115 | WNT6 | 10421 | -0.586 | 0.1418 | No | ||
| 116 | VASP | 10795 | -0.672 | 0.1226 | No | ||
| 117 | NDRG2 | 10858 | -0.686 | 0.1203 | No | ||
| 118 | COBL | 10893 | -0.694 | 0.1196 | No | ||
| 119 | SEC24D | 11016 | -0.720 | 0.1141 | No | ||
| 120 | PAPPA | 11156 | -0.755 | 0.1077 | No | ||
| 121 | RBP4 | 11382 | -0.809 | 0.0968 | No | ||
| 122 | GDNF | 11384 | -0.809 | 0.0980 | No | ||
| 123 | ADCY8 | 11499 | -0.842 | 0.0931 | No | ||
| 124 | BLR1 | 11536 | -0.850 | 0.0925 | No | ||
| 125 | ECM1 | 11793 | -0.914 | 0.0800 | No | ||
| 126 | FGF11 | 11914 | -0.947 | 0.0750 | No | ||
| 127 | CAMKK1 | 11954 | -0.957 | 0.0744 | No | ||
| 128 | XLKD1 | 12419 | -1.087 | 0.0509 | No | ||
| 129 | WNT7B | 12501 | -1.109 | 0.0482 | No | ||
| 130 | ADCK4 | 12551 | -1.123 | 0.0474 | No | ||
| 131 | OLR1 | 12718 | -1.176 | 0.0402 | No | ||
| 132 | SFN | 12828 | -1.208 | 0.0362 | No | ||
| 133 | NTN4 | 12881 | -1.224 | 0.0353 | No | ||
| 134 | MPV17 | 12944 | -1.243 | 0.0338 | No | ||
| 135 | CRYGS | 12990 | -1.260 | 0.0334 | No | ||
| 136 | RB1CC1 | 12996 | -1.261 | 0.0351 | No | ||
| 137 | FOSL1 | 13050 | -1.278 | 0.0342 | No | ||
| 138 | CDH23 | 13241 | -1.336 | 0.0260 | No | ||
| 139 | MYBPH | 13320 | -1.361 | 0.0239 | No | ||
| 140 | PHLDA2 | 13411 | -1.393 | 0.0212 | No | ||
| 141 | TAF15 | 13430 | -1.398 | 0.0224 | No | ||
| 142 | CMYA1 | 13590 | -1.456 | 0.0161 | No | ||
| 143 | DTNA | 13594 | -1.457 | 0.0182 | No | ||
| 144 | PTPRN | 13720 | -1.495 | 0.0137 | No | ||
| 145 | RIMS1 | 13727 | -1.497 | 0.0158 | No | ||
| 146 | OMG | 13841 | -1.543 | 0.0121 | No | ||
| 147 | BNIP3 | 13848 | -1.550 | 0.0142 | No | ||
| 148 | CSF3 | 13921 | -1.574 | 0.0127 | No | ||
| 149 | SPATS2 | 13933 | -1.579 | 0.0146 | No | ||
| 150 | WDFY3 | 13960 | -1.591 | 0.0157 | No | ||
| 151 | P2RXL1 | 14582 | -1.848 | -0.0151 | No | ||
| 152 | SLC26A9 | 14697 | -1.899 | -0.0183 | No | ||
| 153 | PRDM1 | 14786 | -1.938 | -0.0201 | No | ||
| 154 | TLL1 | 14812 | -1.953 | -0.0184 | No | ||
| 155 | MARK4 | 14820 | -1.955 | -0.0157 | No | ||
| 156 | GRIA1 | 14984 | -2.039 | -0.0213 | No | ||
| 157 | ABCF3 | 14997 | -2.045 | -0.0188 | No | ||
| 158 | PVALB | 15111 | -2.109 | -0.0216 | No | ||
| 159 | TUBB4 | 15192 | -2.151 | -0.0226 | No | ||
| 160 | NEFH | 15238 | -2.179 | -0.0216 | No | ||
| 161 | EN1 | 15285 | -2.206 | -0.0206 | No | ||
| 162 | TIAL1 | 15496 | -2.348 | -0.0284 | No | ||
| 163 | MYOZ2 | 15635 | -2.443 | -0.0320 | No | ||
| 164 | DUSP14 | 15708 | -2.495 | -0.0320 | No | ||
| 165 | CNTF | 15710 | -2.495 | -0.0282 | No | ||
| 166 | CSMD3 | 15819 | -2.609 | -0.0299 | No | ||
| 167 | VAPA | 15839 | -2.623 | -0.0268 | No | ||
| 168 | PSME4 | 15864 | -2.644 | -0.0240 | No | ||
| 169 | IL10 | 16046 | -2.825 | -0.0294 | No | ||
| 170 | PSMD4 | 16101 | -2.870 | -0.0278 | No | ||
| 171 | ALDOA | 16213 | -2.983 | -0.0292 | No | ||
| 172 | GKN1 | 16274 | -3.046 | -0.0277 | No | ||
| 173 | PSMD11 | 16285 | -3.054 | -0.0234 | No | ||
| 174 | TTC1 | 16364 | -3.129 | -0.0227 | No | ||
| 175 | FBXO2 | 16616 | -3.472 | -0.0309 | No | ||
| 176 | TRAF3 | 16727 | -3.648 | -0.0312 | No | ||
| 177 | NR1D1 | 16950 | -4.007 | -0.0369 | No | ||
| 178 | NCDN | 17016 | -4.130 | -0.0340 | No | ||
| 179 | NRAS | 17023 | -4.143 | -0.0278 | No | ||
| 180 | TYSND1 | 17193 | -4.529 | -0.0299 | No | ||
| 181 | ENO1 | 17258 | -4.676 | -0.0260 | No | ||
| 182 | TOB1 | 17459 | -5.211 | -0.0287 | No | ||
| 183 | RBBP7 | 17593 | -5.614 | -0.0271 | No | ||
| 184 | SFRS2 | 17611 | -5.660 | -0.0192 | No | ||
| 185 | RHBDF1 | 17818 | -6.522 | -0.0201 | No | ||
| 186 | BAG2 | 17848 | -6.620 | -0.0113 | No | ||
| 187 | SLC16A6 | 17919 | -6.979 | -0.0042 | No | ||
| 188 | TREX1 | 18210 | -8.616 | -0.0064 | No | ||
| 189 | XPOT | 18212 | -8.637 | 0.0070 | No | ||
| 190 | FSTL1 | 18298 | -9.456 | 0.0173 | No |