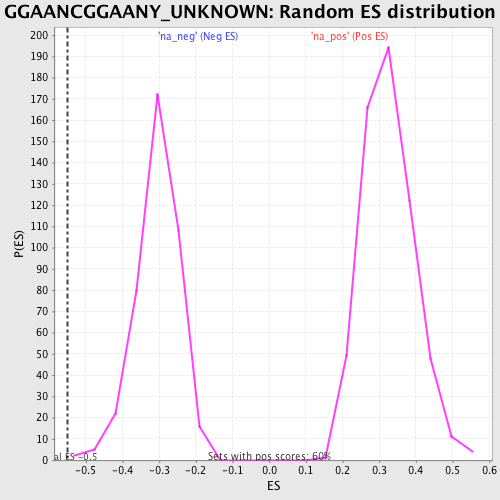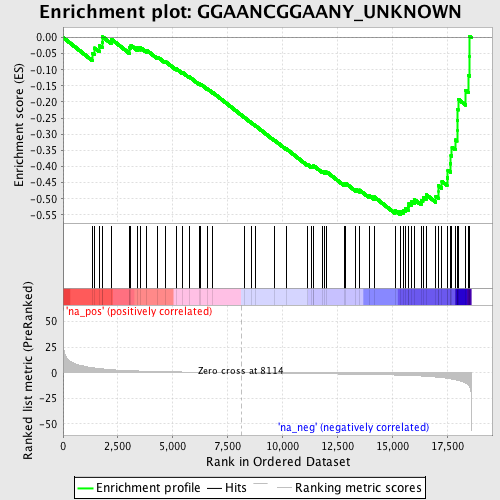
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | GGAANCGGAANY_UNKNOWN |
| Enrichment Score (ES) | -0.54887897 |
| Normalized Enrichment Score (NES) | -1.8005297 |
| Nominal p-value | 0.004938272 |
| FDR q-value | 0.023266975 |
| FWER p-Value | 0.112 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MTA2 | 1349 | 4.965 | -0.0500 | No | ||
| 2 | RRAS | 1428 | 4.766 | -0.0325 | No | ||
| 3 | SDF2 | 1657 | 4.174 | -0.0257 | No | ||
| 4 | PTPRCAP | 1784 | 3.903 | -0.0146 | No | ||
| 5 | SEC24C | 1802 | 3.855 | 0.0021 | No | ||
| 6 | RPL28 | 2204 | 3.102 | -0.0054 | No | ||
| 7 | GGA1 | 3022 | 2.119 | -0.0397 | No | ||
| 8 | BIRC4 | 3023 | 2.117 | -0.0301 | No | ||
| 9 | PEX6 | 3091 | 2.063 | -0.0243 | No | ||
| 10 | DPM1 | 3389 | 1.823 | -0.0319 | No | ||
| 11 | RNF25 | 3520 | 1.729 | -0.0310 | No | ||
| 12 | UBL5 | 3805 | 1.553 | -0.0393 | No | ||
| 13 | PRKAB1 | 4310 | 1.271 | -0.0606 | No | ||
| 14 | UBA52 | 4650 | 1.122 | -0.0738 | No | ||
| 15 | ATP6V1E1 | 5155 | 0.924 | -0.0967 | No | ||
| 16 | FEV | 5446 | 0.814 | -0.1086 | No | ||
| 17 | ATP6V1D | 5737 | 0.702 | -0.1211 | No | ||
| 18 | PSMB4 | 6195 | 0.564 | -0.1431 | No | ||
| 19 | DLX4 | 6277 | 0.537 | -0.1450 | No | ||
| 20 | ZDHHC5 | 6581 | 0.446 | -0.1593 | No | ||
| 21 | BAT3 | 6827 | 0.373 | -0.1708 | No | ||
| 22 | TRIM39 | 8246 | -0.040 | -0.2471 | No | ||
| 23 | SRP54 | 8592 | -0.129 | -0.2651 | No | ||
| 24 | SEC61G | 8772 | -0.173 | -0.2740 | No | ||
| 25 | SNRPE | 9624 | -0.376 | -0.3181 | No | ||
| 26 | UGCGL1 | 10163 | -0.518 | -0.3448 | No | ||
| 27 | FBXO38 | 11154 | -0.754 | -0.3947 | No | ||
| 28 | MRPL21 | 11300 | -0.787 | -0.3989 | No | ||
| 29 | MBD1 | 11391 | -0.811 | -0.4001 | No | ||
| 30 | MTF1 | 11417 | -0.817 | -0.3977 | No | ||
| 31 | TTC17 | 11833 | -0.926 | -0.4158 | No | ||
| 32 | PARVA | 11928 | -0.951 | -0.4165 | No | ||
| 33 | SFRS16 | 12021 | -0.975 | -0.4170 | No | ||
| 34 | SUPT6H | 12808 | -1.200 | -0.4539 | No | ||
| 35 | MRPS21 | 12877 | -1.223 | -0.4520 | No | ||
| 36 | EIF3S3 | 13336 | -1.368 | -0.4704 | No | ||
| 37 | CSNK2B | 13503 | -1.425 | -0.4729 | No | ||
| 38 | IGHMBP2 | 13944 | -1.586 | -0.4894 | No | ||
| 39 | GYS1 | 14177 | -1.670 | -0.4942 | No | ||
| 40 | U2AF2 | 15127 | -2.117 | -0.5357 | No | ||
| 41 | COMMD6 | 15372 | -2.265 | -0.5385 | Yes | ||
| 42 | SMUG1 | 15526 | -2.370 | -0.5359 | Yes | ||
| 43 | BAT4 | 15612 | -2.427 | -0.5294 | Yes | ||
| 44 | SNIP1 | 15733 | -2.521 | -0.5244 | Yes | ||
| 45 | EPN1 | 15759 | -2.550 | -0.5141 | Yes | ||
| 46 | TAF10 | 15882 | -2.663 | -0.5085 | Yes | ||
| 47 | SEC61A1 | 16022 | -2.804 | -0.5032 | Yes | ||
| 48 | VPS16 | 16322 | -3.092 | -0.5052 | Yes | ||
| 49 | EIF2S1 | 16403 | -3.164 | -0.4950 | Yes | ||
| 50 | MRPL43 | 16546 | -3.366 | -0.4873 | Yes | ||
| 51 | GNB2L1 | 16989 | -4.080 | -0.4925 | Yes | ||
| 52 | TFB2M | 17101 | -4.311 | -0.4788 | Yes | ||
| 53 | BANF1 | 17104 | -4.315 | -0.4591 | Yes | ||
| 54 | COX7A2 | 17265 | -4.690 | -0.4463 | Yes | ||
| 55 | NCBP2 | 17500 | -5.321 | -0.4346 | Yes | ||
| 56 | DDX55 | 17538 | -5.467 | -0.4117 | Yes | ||
| 57 | MRPS18A | 17654 | -5.807 | -0.3913 | Yes | ||
| 58 | MRPS23 | 17676 | -5.901 | -0.3655 | Yes | ||
| 59 | RPL38 | 17723 | -6.106 | -0.3401 | Yes | ||
| 60 | PDAP1 | 17887 | -6.820 | -0.3177 | Yes | ||
| 61 | PRPF3 | 17953 | -7.160 | -0.2885 | Yes | ||
| 62 | HSPH1 | 17959 | -7.187 | -0.2559 | Yes | ||
| 63 | PEO1 | 17985 | -7.286 | -0.2239 | Yes | ||
| 64 | RUVBL2 | 18004 | -7.399 | -0.1911 | Yes | ||
| 65 | MAP4K1 | 18350 | -9.884 | -0.1645 | Yes | ||
| 66 | EBNA1BP2 | 18478 | -11.855 | -0.1172 | Yes | ||
| 67 | TIMM8A | 18529 | -13.528 | -0.0581 | Yes | ||
| 68 | RARS | 18536 | -13.724 | 0.0043 | Yes |