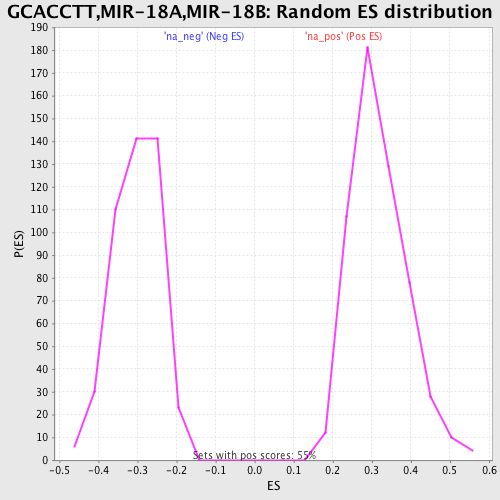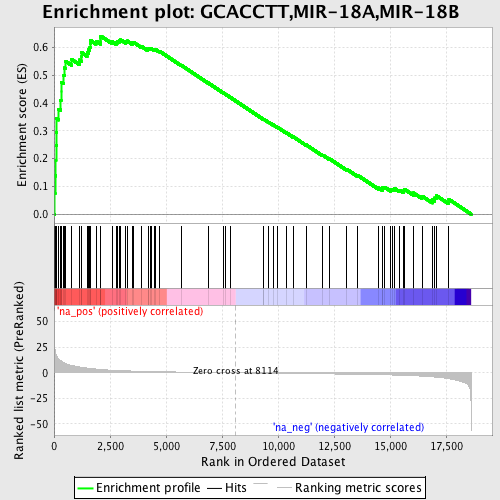
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | GCACCTT,MIR-18A,MIR-18B |
| Enrichment Score (ES) | 0.64081883 |
| Normalized Enrichment Score (NES) | 2.020346 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0042221225 |
| FWER p-Value | 0.0030 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GAB1 | 22 | 26.495 | 0.0770 | Yes | ||
| 2 | KIT | 44 | 21.341 | 0.1388 | Yes | ||
| 3 | JARID1B | 77 | 18.935 | 0.1930 | Yes | ||
| 4 | FRMD4A | 85 | 18.192 | 0.2463 | Yes | ||
| 5 | ARHGEF5 | 101 | 16.999 | 0.2956 | Yes | ||
| 6 | ESR1 | 111 | 16.546 | 0.3439 | Yes | ||
| 7 | NEDD9 | 207 | 13.363 | 0.3782 | Yes | ||
| 8 | RUNX1 | 286 | 11.889 | 0.4091 | Yes | ||
| 9 | ETV6 | 313 | 11.475 | 0.4416 | Yes | ||
| 10 | CTDSPL | 314 | 11.456 | 0.4753 | Yes | ||
| 11 | PIAS3 | 433 | 9.998 | 0.4985 | Yes | ||
| 12 | TRIOBP | 453 | 9.789 | 0.5263 | Yes | ||
| 13 | PRKACB | 519 | 9.130 | 0.5498 | Yes | ||
| 14 | TEX2 | 787 | 7.234 | 0.5567 | Yes | ||
| 15 | ZFP36L1 | 1130 | 5.692 | 0.5551 | Yes | ||
| 16 | ATXN1 | 1217 | 5.372 | 0.5663 | Yes | ||
| 17 | IGF1 | 1226 | 5.346 | 0.5816 | Yes | ||
| 18 | ANKRD50 | 1473 | 4.649 | 0.5821 | Yes | ||
| 19 | EHMT1 | 1546 | 4.470 | 0.5914 | Yes | ||
| 20 | RABGAP1 | 1596 | 4.330 | 0.6015 | Yes | ||
| 21 | RAB5C | 1612 | 4.278 | 0.6133 | Yes | ||
| 22 | NFAT5 | 1621 | 4.255 | 0.6254 | Yes | ||
| 23 | SAR1A | 1870 | 3.709 | 0.6230 | Yes | ||
| 24 | IRF2 | 2076 | 3.300 | 0.6217 | Yes | ||
| 25 | ALCAM | 2080 | 3.291 | 0.6312 | Yes | ||
| 26 | PACSIN1 | 2083 | 3.287 | 0.6408 | Yes | ||
| 27 | ARL15 | 2598 | 2.553 | 0.6206 | No | ||
| 28 | PARP6 | 2771 | 2.360 | 0.6183 | No | ||
| 29 | SOCS5 | 2819 | 2.312 | 0.6226 | No | ||
| 30 | STK4 | 2902 | 2.227 | 0.6248 | No | ||
| 31 | WSB1 | 2973 | 2.156 | 0.6273 | No | ||
| 32 | CTGF | 3202 | 1.978 | 0.6209 | No | ||
| 33 | NR1H2 | 3259 | 1.931 | 0.6236 | No | ||
| 34 | PHF2 | 3485 | 1.761 | 0.6166 | No | ||
| 35 | NAV1 | 3549 | 1.713 | 0.6183 | No | ||
| 36 | NR3C1 | 3907 | 1.496 | 0.6034 | No | ||
| 37 | SNURF | 4191 | 1.339 | 0.5921 | No | ||
| 38 | NCOA1 | 4214 | 1.326 | 0.5948 | No | ||
| 39 | MESP1 | 4284 | 1.286 | 0.5949 | No | ||
| 40 | PURB | 4327 | 1.262 | 0.5964 | No | ||
| 41 | FCHSD2 | 4493 | 1.188 | 0.5910 | No | ||
| 42 | PHF19 | 4534 | 1.167 | 0.5923 | No | ||
| 43 | SMAD2 | 4698 | 1.102 | 0.5867 | No | ||
| 44 | LIN28 | 5677 | 0.720 | 0.5361 | No | ||
| 45 | SH3BP4 | 6875 | 0.362 | 0.4726 | No | ||
| 46 | PTGFRN | 7572 | 0.153 | 0.4355 | No | ||
| 47 | SIM2 | 7649 | 0.132 | 0.4318 | No | ||
| 48 | HIF1A | 7886 | 0.065 | 0.4193 | No | ||
| 49 | BHLHB5 | 9343 | -0.312 | 0.3417 | No | ||
| 50 | AEBP2 | 9355 | -0.314 | 0.3420 | No | ||
| 51 | DPP10 | 9589 | -0.370 | 0.3305 | No | ||
| 52 | PERQ1 | 9782 | -0.419 | 0.3214 | No | ||
| 53 | GLRB | 9968 | -0.468 | 0.3128 | No | ||
| 54 | RHOT1 | 10355 | -0.570 | 0.2937 | No | ||
| 55 | GCLC | 10676 | -0.644 | 0.2783 | No | ||
| 56 | HSF2 | 11244 | -0.772 | 0.2500 | No | ||
| 57 | CRIM1 | 11989 | -0.967 | 0.2127 | No | ||
| 58 | RAB11FIP2 | 12304 | -1.051 | 0.1989 | No | ||
| 59 | CLK2 | 13069 | -1.282 | 0.1615 | No | ||
| 60 | ASXL2 | 13545 | -1.438 | 0.1401 | No | ||
| 61 | MEF2D | 14489 | -1.807 | 0.0946 | No | ||
| 62 | PRICKLE2 | 14657 | -1.882 | 0.0911 | No | ||
| 63 | AKR1D1 | 14668 | -1.888 | 0.0961 | No | ||
| 64 | MAN1A2 | 14755 | -1.924 | 0.0972 | No | ||
| 65 | ZBTB4 | 15027 | -2.061 | 0.0886 | No | ||
| 66 | SON | 15122 | -2.116 | 0.0898 | No | ||
| 67 | CDC42 | 15186 | -2.149 | 0.0928 | No | ||
| 68 | XYLT2 | 15424 | -2.298 | 0.0868 | No | ||
| 69 | FNBP1 | 15617 | -2.432 | 0.0836 | No | ||
| 70 | CLASP2 | 15623 | -2.436 | 0.0905 | No | ||
| 71 | MDGA1 | 16038 | -2.817 | 0.0765 | No | ||
| 72 | TRIB2 | 16422 | -3.191 | 0.0652 | No | ||
| 73 | TRIM2 | 16889 | -3.905 | 0.0516 | No | ||
| 74 | BTG3 | 16991 | -4.084 | 0.0582 | No | ||
| 75 | TARDBP | 17058 | -4.225 | 0.0671 | No | ||
| 76 | ADD3 | 17620 | -5.709 | 0.0537 | No |