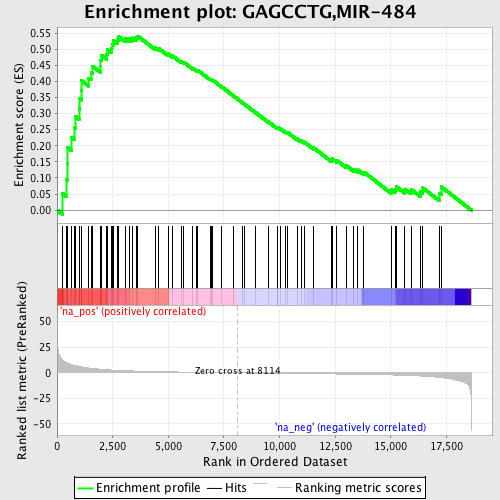
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | GAGCCTG,MIR-484 |
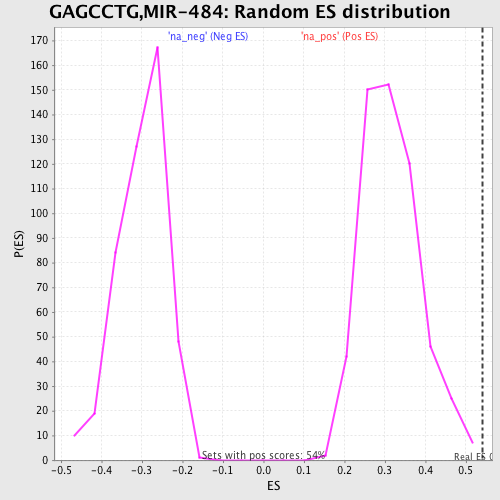
| Enrichment Score (ES) | 0.54072964 |
| Normalized Enrichment Score (NES) | 1.7131789 |
| Nominal p-value | 0.0018382353 |
| FDR q-value | 0.049117304 |
| FWER p-Value | 0.378 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MAPKAPK2 | 249 | 12.454 | 0.0516 | Yes | ||
| 2 | ZNRF1 | 421 | 10.123 | 0.0951 | Yes | ||
| 3 | TRIOBP | 453 | 9.789 | 0.1446 | Yes | ||
| 4 | TSGA14 | 457 | 9.754 | 0.1953 | Yes | ||
| 5 | RAB6IP1 | 658 | 7.944 | 0.2259 | Yes | ||
| 6 | MAP3K11 | 801 | 7.169 | 0.2557 | Yes | ||
| 7 | BCL11A | 818 | 7.061 | 0.2917 | Yes | ||
| 8 | ZFYVE1 | 988 | 6.306 | 0.3155 | Yes | ||
| 9 | HOXA5 | 1022 | 6.162 | 0.3458 | Yes | ||
| 10 | MAP4K2 | 1094 | 5.837 | 0.3725 | Yes | ||
| 11 | MTF2 | 1106 | 5.794 | 0.4021 | Yes | ||
| 12 | ACVR1B | 1414 | 4.804 | 0.4106 | Yes | ||
| 13 | DPYSL2 | 1527 | 4.522 | 0.4282 | Yes | ||
| 14 | CENPB | 1592 | 4.346 | 0.4474 | Yes | ||
| 15 | NFE2L1 | 1933 | 3.572 | 0.4477 | Yes | ||
| 16 | HIC2 | 1935 | 3.567 | 0.4662 | Yes | ||
| 17 | PHC2 | 1987 | 3.477 | 0.4816 | Yes | ||
| 18 | PRKCB1 | 2223 | 3.074 | 0.4850 | Yes | ||
| 19 | DAG1 | 2266 | 3.011 | 0.4985 | Yes | ||
| 20 | NFIA | 2432 | 2.728 | 0.5038 | Yes | ||
| 21 | YTHDF3 | 2468 | 2.691 | 0.5159 | Yes | ||
| 22 | PTPRF | 2528 | 2.629 | 0.5265 | Yes | ||
| 23 | NFKBIZ | 2713 | 2.418 | 0.5292 | Yes | ||
| 24 | HIPK1 | 2744 | 2.390 | 0.5400 | Yes | ||
| 25 | PCDH7 | 3057 | 2.089 | 0.5341 | Yes | ||
| 26 | LBH | 3249 | 1.940 | 0.5339 | Yes | ||
| 27 | FGF1 | 3407 | 1.812 | 0.5349 | Yes | ||
| 28 | NAV1 | 3549 | 1.713 | 0.5362 | Yes | ||
| 29 | TCF1 | 3628 | 1.668 | 0.5407 | Yes | ||
| 30 | IPO11 | 4403 | 1.227 | 0.5054 | No | ||
| 31 | EIF2C1 | 4573 | 1.153 | 0.5023 | No | ||
| 32 | PITPNA | 4984 | 0.984 | 0.4853 | No | ||
| 33 | PLEKHH2 | 5174 | 0.917 | 0.4799 | No | ||
| 34 | SERTAD1 | 5587 | 0.751 | 0.4616 | No | ||
| 35 | SCARA3 | 5684 | 0.718 | 0.4602 | No | ||
| 36 | PRPF4B | 6096 | 0.596 | 0.4411 | No | ||
| 37 | JMJD2A | 6265 | 0.541 | 0.4349 | No | ||
| 38 | TRPS1 | 6309 | 0.529 | 0.4353 | No | ||
| 39 | ZIC5 | 6873 | 0.363 | 0.4069 | No | ||
| 40 | BTG4 | 6946 | 0.338 | 0.4048 | No | ||
| 41 | EDA | 6988 | 0.326 | 0.4042 | No | ||
| 42 | MLLT7 | 7397 | 0.209 | 0.3833 | No | ||
| 43 | DLL4 | 7945 | 0.048 | 0.3541 | No | ||
| 44 | HIVEP2 | 8336 | -0.065 | 0.3334 | No | ||
| 45 | DND1 | 8424 | -0.086 | 0.3292 | No | ||
| 46 | LYPLA2 | 8934 | -0.212 | 0.3028 | No | ||
| 47 | SLC6A1 | 9489 | -0.346 | 0.2747 | No | ||
| 48 | EZH1 | 9888 | -0.450 | 0.2556 | No | ||
| 49 | DCBLD2 | 9927 | -0.459 | 0.2560 | No | ||
| 50 | PGBD5 | 10041 | -0.488 | 0.2524 | No | ||
| 51 | TEX261 | 10272 | -0.549 | 0.2429 | No | ||
| 52 | PTGER4 | 10337 | -0.567 | 0.2424 | No | ||
| 53 | RAF1 | 10793 | -0.671 | 0.2214 | No | ||
| 54 | NFATC4 | 10972 | -0.710 | 0.2155 | No | ||
| 55 | ZIC1 | 11113 | -0.741 | 0.2118 | No | ||
| 56 | SEMA4G | 11529 | -0.849 | 0.1938 | No | ||
| 57 | TRIM33 | 12338 | -1.061 | 0.1558 | No | ||
| 58 | COL12A1 | 12373 | -1.070 | 0.1596 | No | ||
| 59 | DACH1 | 12575 | -1.132 | 0.1546 | No | ||
| 60 | MINK1 | 13010 | -1.265 | 0.1378 | No | ||
| 61 | GRB10 | 13328 | -1.365 | 0.1278 | No | ||
| 62 | KLF12 | 13523 | -1.432 | 0.1249 | No | ||
| 63 | PALM2 | 13788 | -1.521 | 0.1186 | No | ||
| 64 | DGCR8 | 15014 | -2.053 | 0.0632 | No | ||
| 65 | SP6 | 15189 | -2.150 | 0.0650 | No | ||
| 66 | CBL | 15237 | -2.177 | 0.0739 | No | ||
| 67 | M6PR | 15634 | -2.442 | 0.0653 | No | ||
| 68 | DOLPP1 | 15911 | -2.695 | 0.0644 | No | ||
| 69 | PTPRE | 16345 | -3.114 | 0.0573 | No | ||
| 70 | HSPG2 | 16442 | -3.223 | 0.0690 | No | ||
| 71 | TYSND1 | 17193 | -4.529 | 0.0522 | No | ||
| 72 | EIF4G2 | 17271 | -4.699 | 0.0725 | No |