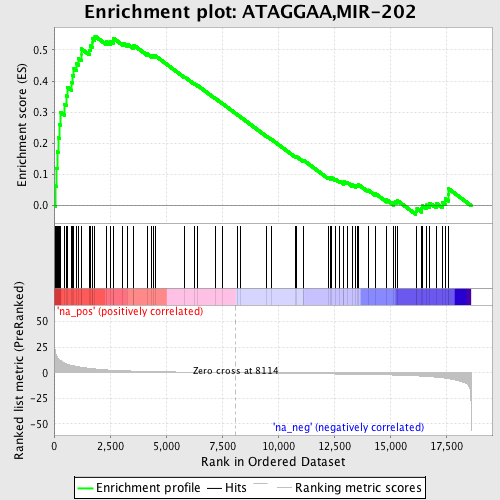
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | ATAGGAA,MIR-202 |
| Enrichment Score (ES) | 0.54439116 |
| Normalized Enrichment Score (NES) | 1.7114398 |
| Nominal p-value | 0.0033783785 |
| FDR q-value | 0.043767996 |
| FWER p-Value | 0.386 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SSBP2 | 78 | 18.923 | 0.0632 | Yes | ||
| 2 | ESR1 | 111 | 16.546 | 0.1205 | Yes | ||
| 3 | MARCKS | 141 | 15.219 | 0.1732 | Yes | ||
| 4 | NDRG1 | 200 | 13.450 | 0.2180 | Yes | ||
| 5 | VANGL2 | 246 | 12.473 | 0.2600 | Yes | ||
| 6 | ZFAND3 | 297 | 11.761 | 0.2993 | Yes | ||
| 7 | ELF1 | 458 | 9.738 | 0.3253 | Yes | ||
| 8 | ACVR1 | 542 | 8.880 | 0.3525 | Yes | ||
| 9 | CNN3 | 587 | 8.469 | 0.3803 | Yes | ||
| 10 | STAT3 | 790 | 7.215 | 0.3952 | Yes | ||
| 11 | BCL11A | 818 | 7.061 | 0.4189 | Yes | ||
| 12 | RNF11 | 883 | 6.809 | 0.4397 | Yes | ||
| 13 | LBR | 989 | 6.305 | 0.4565 | Yes | ||
| 14 | THRAP2 | 1096 | 5.831 | 0.4716 | Yes | ||
| 15 | RAB22A | 1214 | 5.380 | 0.4844 | Yes | ||
| 16 | ATXN1 | 1217 | 5.372 | 0.5035 | Yes | ||
| 17 | KRAS | 1579 | 4.377 | 0.4996 | Yes | ||
| 18 | DDEF1 | 1615 | 4.273 | 0.5130 | Yes | ||
| 19 | RASA1 | 1707 | 4.056 | 0.5225 | Yes | ||
| 20 | PTEN | 1710 | 4.050 | 0.5368 | Yes | ||
| 21 | LUZP1 | 1823 | 3.814 | 0.5444 | Yes | ||
| 22 | USP15 | 2343 | 2.855 | 0.5266 | No | ||
| 23 | GRIA3 | 2509 | 2.650 | 0.5271 | No | ||
| 24 | MIB1 | 2643 | 2.498 | 0.5289 | No | ||
| 25 | RAD52 | 2647 | 2.492 | 0.5376 | No | ||
| 26 | CREBBP | 3069 | 2.079 | 0.5223 | No | ||
| 27 | ANP32E | 3288 | 1.909 | 0.5173 | No | ||
| 28 | ERBB2IP | 3546 | 1.714 | 0.5096 | No | ||
| 29 | TCF12 | 3555 | 1.709 | 0.5152 | No | ||
| 30 | USP8 | 4149 | 1.355 | 0.4881 | No | ||
| 31 | PCGF2 | 4365 | 1.241 | 0.4809 | No | ||
| 32 | MAPK6 | 4447 | 1.212 | 0.4809 | No | ||
| 33 | CTBP2 | 4514 | 1.176 | 0.4815 | No | ||
| 34 | CD28 | 5835 | 0.671 | 0.4127 | No | ||
| 35 | APPBP2 | 6263 | 0.542 | 0.3916 | No | ||
| 36 | HLF | 6412 | 0.496 | 0.3854 | No | ||
| 37 | ACSL3 | 7185 | 0.272 | 0.3447 | No | ||
| 38 | EI24 | 7510 | 0.173 | 0.3279 | No | ||
| 39 | SNAP91 | 8204 | -0.026 | 0.2906 | No | ||
| 40 | RPS6KA3 | 8319 | -0.060 | 0.2847 | No | ||
| 41 | KIAA1715 | 9491 | -0.346 | 0.2227 | No | ||
| 42 | DNAJC10 | 9704 | -0.396 | 0.2127 | No | ||
| 43 | RKHD3 | 10790 | -0.670 | 0.1566 | No | ||
| 44 | SFPQ | 10840 | -0.683 | 0.1564 | No | ||
| 45 | YAF2 | 11127 | -0.746 | 0.1436 | No | ||
| 46 | HOXB2 | 11151 | -0.753 | 0.1451 | No | ||
| 47 | IQGAP1 | 12255 | -1.039 | 0.0893 | No | ||
| 48 | TRIM33 | 12338 | -1.061 | 0.0886 | No | ||
| 49 | SORCS1 | 12362 | -1.067 | 0.0912 | No | ||
| 50 | CPEB3 | 12560 | -1.126 | 0.0846 | No | ||
| 51 | RNF121 | 12744 | -1.184 | 0.0789 | No | ||
| 52 | SPRED1 | 12934 | -1.240 | 0.0732 | No | ||
| 53 | NR2F2 | 12936 | -1.241 | 0.0775 | No | ||
| 54 | SNX16 | 13076 | -1.283 | 0.0746 | No | ||
| 55 | FAM60A | 13309 | -1.358 | 0.0669 | No | ||
| 56 | ARSB | 13458 | -1.406 | 0.0640 | No | ||
| 57 | KLF12 | 13523 | -1.432 | 0.0656 | No | ||
| 58 | BTG1 | 13585 | -1.454 | 0.0675 | No | ||
| 59 | ENAH | 14013 | -1.609 | 0.0502 | No | ||
| 60 | KHDRBS2 | 14340 | -1.736 | 0.0388 | No | ||
| 61 | BICD2 | 14848 | -1.969 | 0.0185 | No | ||
| 62 | HIP2 | 15161 | -2.141 | 0.0093 | No | ||
| 63 | CBL | 15237 | -2.177 | 0.0130 | No | ||
| 64 | TGFBR2 | 15324 | -2.233 | 0.0163 | No | ||
| 65 | EAF1 | 16155 | -2.929 | -0.0180 | No | ||
| 66 | DDX3X | 16197 | -2.966 | -0.0096 | No | ||
| 67 | ROBO2 | 16405 | -3.169 | -0.0095 | No | ||
| 68 | PPP5C | 16423 | -3.196 | 0.0010 | No | ||
| 69 | SENP1 | 16634 | -3.505 | 0.0022 | No | ||
| 70 | BCL2 | 16768 | -3.720 | 0.0083 | No | ||
| 71 | TARDBP | 17058 | -4.225 | 0.0077 | No | ||
| 72 | TTC13 | 17340 | -4.872 | 0.0099 | No | ||
| 73 | PPARBP | 17456 | -5.199 | 0.0223 | No | ||
| 74 | NARG1 | 17598 | -5.625 | 0.0347 | No | ||
| 75 | BAG4 | 17607 | -5.647 | 0.0544 | No |
