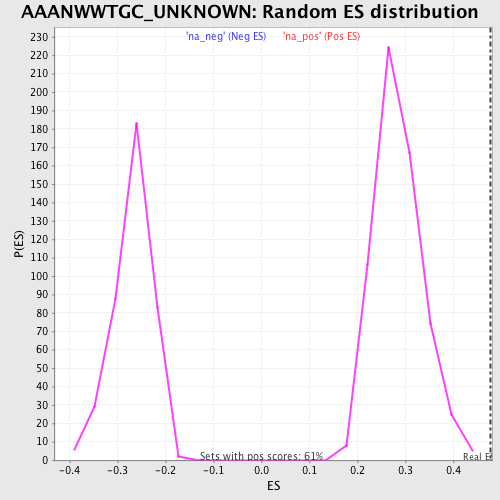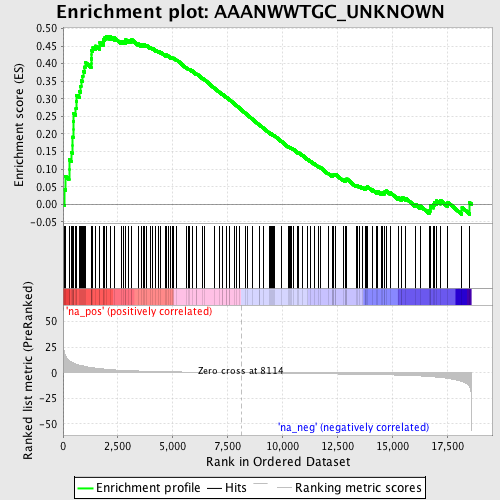
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | AAANWWTGC_UNKNOWN |
| Enrichment Score (ES) | 0.47827664 |
| Normalized Enrichment Score (NES) | 1.6821078 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.054734707 |
| FWER p-Value | 0.495 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MYLK | 70 | 19.287 | 0.0421 | Yes | ||
| 2 | ESR1 | 111 | 16.546 | 0.0792 | Yes | ||
| 3 | MEF2C | 271 | 12.059 | 0.0993 | Yes | ||
| 4 | BCL6 | 274 | 12.030 | 0.1278 | Yes | ||
| 5 | SORT1 | 375 | 10.673 | 0.1477 | Yes | ||
| 6 | SOX4 | 438 | 9.925 | 0.1680 | Yes | ||
| 7 | MEIS1 | 440 | 9.904 | 0.1915 | Yes | ||
| 8 | ITM2C | 460 | 9.734 | 0.2136 | Yes | ||
| 9 | HHEX | 462 | 9.718 | 0.2366 | Yes | ||
| 10 | TLE4 | 473 | 9.560 | 0.2588 | Yes | ||
| 11 | PPARGC1A | 584 | 8.495 | 0.2731 | Yes | ||
| 12 | MGLL | 605 | 8.330 | 0.2918 | Yes | ||
| 13 | DDIT3 | 618 | 8.219 | 0.3107 | Yes | ||
| 14 | EFNA5 | 750 | 7.452 | 0.3213 | Yes | ||
| 15 | DUSP1 | 797 | 7.185 | 0.3359 | Yes | ||
| 16 | BCL11A | 818 | 7.061 | 0.3516 | Yes | ||
| 17 | ATP1B1 | 875 | 6.834 | 0.3648 | Yes | ||
| 18 | DMD | 908 | 6.710 | 0.3790 | Yes | ||
| 19 | DSCAM | 976 | 6.359 | 0.3905 | Yes | ||
| 20 | SSBP3 | 1003 | 6.269 | 0.4040 | Yes | ||
| 21 | ZHX3 | 1278 | 5.193 | 0.4015 | Yes | ||
| 22 | PPP3CC | 1281 | 5.185 | 0.4137 | Yes | ||
| 23 | THBS2 | 1298 | 5.112 | 0.4250 | Yes | ||
| 24 | MBNL1 | 1300 | 5.110 | 0.4371 | Yes | ||
| 25 | RBMX | 1359 | 4.944 | 0.4457 | Yes | ||
| 26 | DLGAP4 | 1481 | 4.635 | 0.4502 | Yes | ||
| 27 | CDC42EP3 | 1671 | 4.140 | 0.4498 | Yes | ||
| 28 | SOX13 | 1675 | 4.134 | 0.4595 | Yes | ||
| 29 | MAPK3 | 1828 | 3.801 | 0.4603 | Yes | ||
| 30 | CHD2 | 1845 | 3.767 | 0.4684 | Yes | ||
| 31 | MAP3K4 | 1905 | 3.628 | 0.4738 | Yes | ||
| 32 | DNAJB5 | 1977 | 3.492 | 0.4783 | Yes | ||
| 33 | EPHB2 | 2148 | 3.187 | 0.4766 | No | ||
| 34 | APP | 2332 | 2.878 | 0.4736 | No | ||
| 35 | HOXB6 | 2638 | 2.506 | 0.4630 | No | ||
| 36 | CDK2AP1 | 2747 | 2.388 | 0.4629 | No | ||
| 37 | ZFPM1 | 2854 | 2.276 | 0.4625 | No | ||
| 38 | POU2F1 | 2859 | 2.270 | 0.4677 | No | ||
| 39 | OLFM1 | 2988 | 2.144 | 0.4659 | No | ||
| 40 | CHES1 | 3099 | 2.053 | 0.4648 | No | ||
| 41 | SNCAIP | 3131 | 2.031 | 0.4680 | No | ||
| 42 | FN1 | 3426 | 1.799 | 0.4563 | No | ||
| 43 | MRPL24 | 3553 | 1.710 | 0.4536 | No | ||
| 44 | RAB10 | 3664 | 1.643 | 0.4515 | No | ||
| 45 | PDGFRB | 3690 | 1.627 | 0.4540 | No | ||
| 46 | PHF15 | 3782 | 1.567 | 0.4528 | No | ||
| 47 | ELAVL4 | 3979 | 1.448 | 0.4456 | No | ||
| 48 | FZD7 | 4079 | 1.396 | 0.4436 | No | ||
| 49 | TRAM1 | 4232 | 1.317 | 0.4385 | No | ||
| 50 | CD14 | 4352 | 1.245 | 0.4350 | No | ||
| 51 | WDR23 | 4448 | 1.212 | 0.4328 | No | ||
| 52 | RORA | 4661 | 1.116 | 0.4239 | No | ||
| 53 | SOX5 | 4692 | 1.104 | 0.4249 | No | ||
| 54 | NFE2L2 | 4795 | 1.064 | 0.4219 | No | ||
| 55 | NEK6 | 4914 | 1.016 | 0.4180 | No | ||
| 56 | PAX1 | 4993 | 0.981 | 0.4161 | No | ||
| 57 | CITED2 | 5036 | 0.967 | 0.4161 | No | ||
| 58 | PHTF1 | 5182 | 0.913 | 0.4104 | No | ||
| 59 | LSAMP | 5635 | 0.736 | 0.3877 | No | ||
| 60 | EIF4EBP2 | 5736 | 0.702 | 0.3840 | No | ||
| 61 | GPC3 | 5772 | 0.692 | 0.3837 | No | ||
| 62 | HOXA3 | 5879 | 0.657 | 0.3795 | No | ||
| 63 | LRRN1 | 6077 | 0.600 | 0.3703 | No | ||
| 64 | PRPF4B | 6096 | 0.596 | 0.3707 | No | ||
| 65 | MMP3 | 6363 | 0.511 | 0.3575 | No | ||
| 66 | LOX | 6438 | 0.489 | 0.3547 | No | ||
| 67 | ADHFE1 | 6908 | 0.353 | 0.3301 | No | ||
| 68 | ATOH7 | 7126 | 0.288 | 0.3191 | No | ||
| 69 | NEUROG1 | 7251 | 0.253 | 0.3130 | No | ||
| 70 | DRD3 | 7258 | 0.251 | 0.3132 | No | ||
| 71 | HOXA2 | 7432 | 0.198 | 0.3043 | No | ||
| 72 | NTN1 | 7460 | 0.190 | 0.3033 | No | ||
| 73 | PCTP | 7563 | 0.156 | 0.2982 | No | ||
| 74 | GATA3 | 7584 | 0.149 | 0.2974 | No | ||
| 75 | ZNF462 | 7790 | 0.091 | 0.2866 | No | ||
| 76 | SHC3 | 7883 | 0.066 | 0.2817 | No | ||
| 77 | FGF7 | 8025 | 0.025 | 0.2742 | No | ||
| 78 | FTHL17 | 8304 | -0.058 | 0.2592 | No | ||
| 79 | ELF4 | 8385 | -0.077 | 0.2551 | No | ||
| 80 | CACNG3 | 8624 | -0.136 | 0.2425 | No | ||
| 81 | OLIG2 | 8957 | -0.218 | 0.2251 | No | ||
| 82 | MYH2 | 8972 | -0.222 | 0.2249 | No | ||
| 83 | ASPA | 9142 | -0.262 | 0.2163 | No | ||
| 84 | PAX6 | 9397 | -0.326 | 0.2033 | No | ||
| 85 | LIPG | 9472 | -0.342 | 0.2002 | No | ||
| 86 | LECT1 | 9480 | -0.344 | 0.2006 | No | ||
| 87 | CDC42EP5 | 9551 | -0.360 | 0.1977 | No | ||
| 88 | SPARCL1 | 9604 | -0.373 | 0.1957 | No | ||
| 89 | ANK2 | 9645 | -0.381 | 0.1945 | No | ||
| 90 | FBXW7 | 9957 | -0.465 | 0.1787 | No | ||
| 91 | PHOX2B | 10261 | -0.547 | 0.1636 | No | ||
| 92 | PRDM16 | 10323 | -0.564 | 0.1617 | No | ||
| 93 | CACNA1D | 10365 | -0.572 | 0.1608 | No | ||
| 94 | OTX2 | 10429 | -0.588 | 0.1588 | No | ||
| 95 | CEPT1 | 10483 | -0.598 | 0.1573 | No | ||
| 96 | GPC6 | 10701 | -0.648 | 0.1471 | No | ||
| 97 | SESN2 | 10705 | -0.649 | 0.1485 | No | ||
| 98 | FOXP1 | 10892 | -0.694 | 0.1401 | No | ||
| 99 | HOXB2 | 11151 | -0.753 | 0.1279 | No | ||
| 100 | PCSK1 | 11275 | -0.781 | 0.1231 | No | ||
| 101 | CDH13 | 11434 | -0.823 | 0.1165 | No | ||
| 102 | GPR21 | 11633 | -0.876 | 0.1079 | No | ||
| 103 | DSTN | 11724 | -0.897 | 0.1051 | No | ||
| 104 | SATB2 | 12075 | -0.992 | 0.0885 | No | ||
| 105 | LOXL4 | 12270 | -1.043 | 0.0805 | No | ||
| 106 | MLLT6 | 12275 | -1.044 | 0.0828 | No | ||
| 107 | CALM1 | 12307 | -1.052 | 0.0836 | No | ||
| 108 | PRIMA1 | 12344 | -1.062 | 0.0842 | No | ||
| 109 | DAB1 | 12401 | -1.079 | 0.0837 | No | ||
| 110 | CNTFR | 12433 | -1.091 | 0.0846 | No | ||
| 111 | SLMAP | 12771 | -1.190 | 0.0692 | No | ||
| 112 | GRIN2B | 12859 | -1.218 | 0.0674 | No | ||
| 113 | POU4F1 | 12866 | -1.221 | 0.0700 | No | ||
| 114 | NNAT | 12909 | -1.233 | 0.0706 | No | ||
| 115 | NR2F2 | 12936 | -1.241 | 0.0722 | No | ||
| 116 | BNC2 | 13350 | -1.373 | 0.0531 | No | ||
| 117 | SKP2 | 13406 | -1.391 | 0.0534 | No | ||
| 118 | KLF12 | 13523 | -1.432 | 0.0505 | No | ||
| 119 | MAN2A2 | 13651 | -1.475 | 0.0472 | No | ||
| 120 | TRPM3 | 13767 | -1.513 | 0.0445 | No | ||
| 121 | FOXP2 | 13796 | -1.525 | 0.0466 | No | ||
| 122 | OMG | 13841 | -1.543 | 0.0479 | No | ||
| 123 | SFRP2 | 13861 | -1.556 | 0.0506 | No | ||
| 124 | SGCD | 14103 | -1.644 | 0.0415 | No | ||
| 125 | SKIL | 14293 | -1.712 | 0.0353 | No | ||
| 126 | SPAG9 | 14336 | -1.735 | 0.0371 | No | ||
| 127 | GLRA2 | 14522 | -1.821 | 0.0315 | No | ||
| 128 | EPHA7 | 14572 | -1.843 | 0.0332 | No | ||
| 129 | ACTB | 14646 | -1.876 | 0.0337 | No | ||
| 130 | TLX3 | 14666 | -1.888 | 0.0372 | No | ||
| 131 | TLK1 | 14729 | -1.911 | 0.0383 | No | ||
| 132 | MID1 | 14907 | -2.003 | 0.0335 | No | ||
| 133 | DPYSL5 | 15275 | -2.201 | 0.0189 | No | ||
| 134 | GTF2E2 | 15441 | -2.314 | 0.0154 | No | ||
| 135 | SEMA6C | 15444 | -2.317 | 0.0208 | No | ||
| 136 | ZW10 | 15616 | -2.431 | 0.0174 | No | ||
| 137 | FGFR2 | 16060 | -2.833 | 0.0001 | No | ||
| 138 | NTRK3 | 16280 | -3.049 | -0.0045 | No | ||
| 139 | ANK3 | 16683 | -3.576 | -0.0178 | No | ||
| 140 | PPP2R2A | 16726 | -3.640 | -0.0114 | No | ||
| 141 | HOXC4 | 16736 | -3.657 | -0.0032 | No | ||
| 142 | GANAB | 16872 | -3.880 | -0.0013 | No | ||
| 143 | PPP1R10 | 16912 | -3.938 | 0.0060 | No | ||
| 144 | NRAS | 17023 | -4.143 | 0.0099 | No | ||
| 145 | RRS1 | 17190 | -4.513 | 0.0116 | No | ||
| 146 | PIK3R3 | 17535 | -5.453 | 0.0060 | No | ||
| 147 | PRKRIR | 18179 | -8.413 | -0.0088 | No | ||
| 148 | MRPS18B | 18534 | -13.647 | 0.0044 | No |