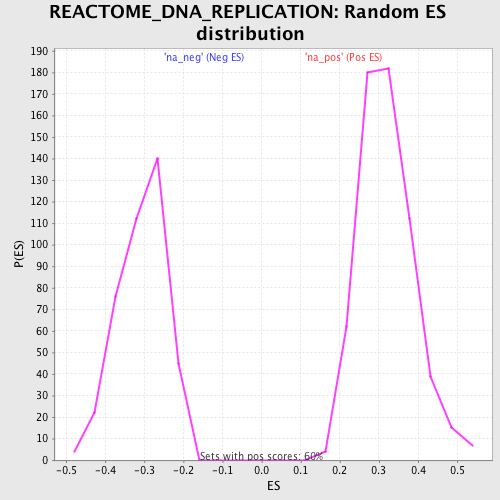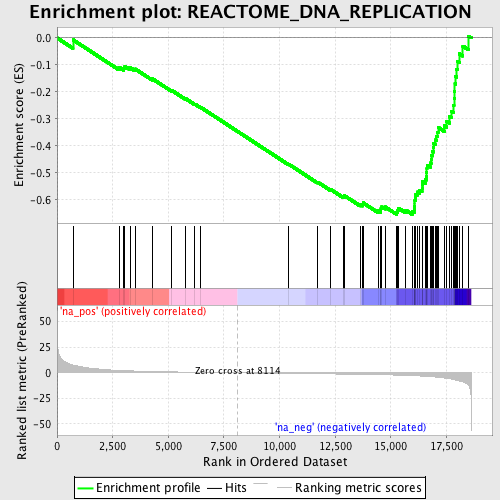
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_transDMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | REACTOME_DNA_REPLICATION |
| Enrichment Score (ES) | -0.65442866 |
| Normalized Enrichment Score (NES) | -2.1353328 |
| Nominal p-value | 0.0 |
| FDR q-value | 6.284341E-5 |
| FWER p-Value | 0.0010 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PSMB9 | 715 | 7.628 | -0.0070 | No | ||
| 2 | PSMA6 | 2787 | 2.344 | -0.1090 | No | ||
| 3 | POLD1 | 2998 | 2.135 | -0.1115 | No | ||
| 4 | PSMA3 | 3045 | 2.102 | -0.1053 | No | ||
| 5 | PSMB6 | 3276 | 1.921 | -0.1097 | No | ||
| 6 | PSMA2 | 3509 | 1.736 | -0.1150 | No | ||
| 7 | PSMB10 | 4286 | 1.286 | -0.1516 | No | ||
| 8 | PSMC3 | 5139 | 0.928 | -0.1937 | No | ||
| 9 | POLA1 | 5762 | 0.693 | -0.2243 | No | ||
| 10 | PSMB4 | 6195 | 0.564 | -0.2453 | No | ||
| 11 | RB1 | 6458 | 0.483 | -0.2574 | No | ||
| 12 | PCNA | 10397 | -0.580 | -0.4673 | No | ||
| 13 | POLD3 | 11705 | -0.892 | -0.5341 | No | ||
| 14 | POLE2 | 12276 | -1.045 | -0.5605 | No | ||
| 15 | PSMD10 | 12884 | -1.226 | -0.5882 | No | ||
| 16 | RPA1 | 12898 | -1.231 | -0.5838 | No | ||
| 17 | PSMC6 | 13626 | -1.466 | -0.6169 | No | ||
| 18 | PSMB1 | 13741 | -1.503 | -0.6169 | No | ||
| 19 | PSMC1 | 13752 | -1.507 | -0.6112 | No | ||
| 20 | PSMD8 | 14434 | -1.779 | -0.6405 | No | ||
| 21 | FEN1 | 14546 | -1.834 | -0.6389 | No | ||
| 22 | RFC1 | 14556 | -1.839 | -0.6318 | No | ||
| 23 | MCM6 | 14599 | -1.854 | -0.6264 | No | ||
| 24 | PSMA5 | 14746 | -1.921 | -0.6263 | No | ||
| 25 | PSME1 | 15268 | -2.197 | -0.6453 | Yes | ||
| 26 | MCM5 | 15314 | -2.227 | -0.6386 | Yes | ||
| 27 | PSMB5 | 15337 | -2.243 | -0.6305 | Yes | ||
| 28 | PSMD5 | 15669 | -2.467 | -0.6381 | Yes | ||
| 29 | PSMD3 | 15959 | -2.739 | -0.6424 | Yes | ||
| 30 | LIG1 | 16050 | -2.827 | -0.6355 | Yes | ||
| 31 | PRIM1 | 16066 | -2.838 | -0.6246 | Yes | ||
| 32 | PSMA4 | 16079 | -2.851 | -0.6135 | Yes | ||
| 33 | CDC7 | 16081 | -2.853 | -0.6017 | Yes | ||
| 34 | PSMD4 | 16101 | -2.870 | -0.5909 | Yes | ||
| 35 | PSMC2 | 16121 | -2.892 | -0.5800 | Yes | ||
| 36 | PSMD9 | 16182 | -2.953 | -0.5710 | Yes | ||
| 37 | PSMD11 | 16285 | -3.054 | -0.5638 | Yes | ||
| 38 | MCM2 | 16404 | -3.166 | -0.5571 | Yes | ||
| 39 | ORC4L | 16414 | -3.183 | -0.5444 | Yes | ||
| 40 | RFC3 | 16431 | -3.206 | -0.5321 | Yes | ||
| 41 | MCM7 | 16549 | -3.369 | -0.5244 | Yes | ||
| 42 | PSMD14 | 16593 | -3.440 | -0.5125 | Yes | ||
| 43 | RPA3 | 16606 | -3.458 | -0.4989 | Yes | ||
| 44 | MCM4 | 16623 | -3.480 | -0.4854 | Yes | ||
| 45 | PSMD12 | 16629 | -3.498 | -0.4712 | Yes | ||
| 46 | PSMB3 | 16772 | -3.724 | -0.4634 | Yes | ||
| 47 | E2F2 | 16811 | -3.778 | -0.4498 | Yes | ||
| 48 | PSMC4 | 16828 | -3.809 | -0.4350 | Yes | ||
| 49 | CDK2 | 16879 | -3.891 | -0.4216 | Yes | ||
| 50 | MCM3 | 16907 | -3.932 | -0.4068 | Yes | ||
| 51 | PSMB2 | 16925 | -3.969 | -0.3913 | Yes | ||
| 52 | PSMA1 | 16995 | -4.092 | -0.3781 | Yes | ||
| 53 | PSMA7 | 17051 | -4.211 | -0.3636 | Yes | ||
| 54 | GMNN | 17110 | -4.344 | -0.3488 | Yes | ||
| 55 | PSMD2 | 17128 | -4.378 | -0.3316 | Yes | ||
| 56 | PSME3 | 17404 | -5.037 | -0.3256 | Yes | ||
| 57 | RPA2 | 17511 | -5.355 | -0.3092 | Yes | ||
| 58 | CDC45L | 17635 | -5.759 | -0.2920 | Yes | ||
| 59 | POLE | 17722 | -6.104 | -0.2714 | Yes | ||
| 60 | ORC2L | 17822 | -6.535 | -0.2497 | Yes | ||
| 61 | MCM10 | 17841 | -6.601 | -0.2234 | Yes | ||
| 62 | ORC1L | 17868 | -6.700 | -0.1971 | Yes | ||
| 63 | PSMC5 | 17883 | -6.771 | -0.1698 | Yes | ||
| 64 | RFC5 | 17910 | -6.942 | -0.1425 | Yes | ||
| 65 | POLA2 | 17962 | -7.207 | -0.1155 | Yes | ||
| 66 | RFC4 | 18000 | -7.383 | -0.0870 | Yes | ||
| 67 | PSMD7 | 18082 | -7.872 | -0.0588 | Yes | ||
| 68 | POLD2 | 18238 | -8.926 | -0.0302 | Yes | ||
| 69 | ORC5L | 18491 | -12.227 | 0.0067 | Yes |