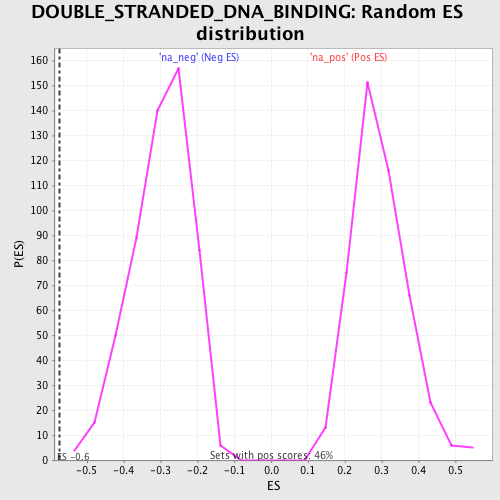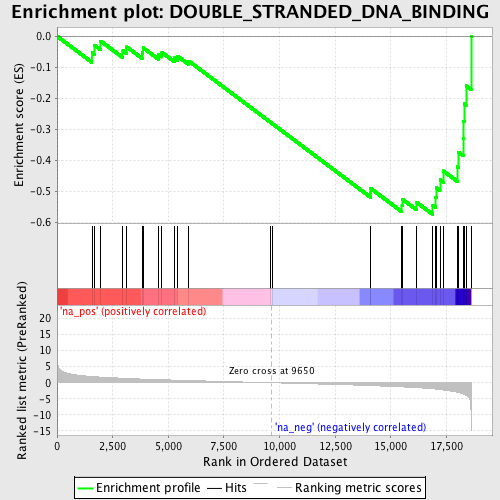
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | DOUBLE_STRANDED_DNA_BINDING |
| Enrichment Score (ES) | -0.5746134 |
| Normalized Enrichment Score (NES) | -1.9221298 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.053043224 |
| FWER p-Value | 0.389 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SMAD2 | 1568 | 1.934 | -0.0531 | No | ||
| 2 | MSH6 | 1681 | 1.874 | -0.0289 | No | ||
| 3 | CGGBP1 | 1959 | 1.744 | -0.0156 | No | ||
| 4 | PNKP | 2959 | 1.394 | -0.0469 | No | ||
| 5 | HMGB2 | 3135 | 1.340 | -0.0346 | No | ||
| 6 | MSH3 | 3824 | 1.135 | -0.0533 | No | ||
| 7 | TERF1 | 3860 | 1.125 | -0.0370 | No | ||
| 8 | XRCC5 | 4569 | 0.965 | -0.0595 | No | ||
| 9 | HLF | 4709 | 0.933 | -0.0519 | No | ||
| 10 | XRCC6 | 5288 | 0.814 | -0.0699 | No | ||
| 11 | APTX | 5422 | 0.784 | -0.0644 | No | ||
| 12 | FEN1 | 5919 | 0.680 | -0.0801 | No | ||
| 13 | PMS2 | 9581 | 0.014 | -0.2769 | No | ||
| 14 | SATB1 | 9672 | -0.005 | -0.2816 | No | ||
| 15 | MSH2 | 14083 | -0.890 | -0.5045 | No | ||
| 16 | RBMS1 | 14088 | -0.891 | -0.4904 | No | ||
| 17 | IFI16 | 15501 | -1.285 | -0.5456 | Yes | ||
| 18 | PURA | 15510 | -1.289 | -0.5252 | Yes | ||
| 19 | NTHL1 | 16154 | -1.533 | -0.5350 | Yes | ||
| 20 | RAD51 | 16891 | -1.889 | -0.5441 | Yes | ||
| 21 | KIN | 17024 | -1.968 | -0.5194 | Yes | ||
| 22 | MTERF | 17049 | -1.995 | -0.4885 | Yes | ||
| 23 | MRE11A | 17211 | -2.133 | -0.4627 | Yes | ||
| 24 | CSDA | 17360 | -2.253 | -0.4343 | Yes | ||
| 25 | MLH1 | 17987 | -2.959 | -0.4202 | Yes | ||
| 26 | ZNF638 | 18052 | -3.037 | -0.3746 | Yes | ||
| 27 | HNRPDL | 18252 | -3.448 | -0.3296 | Yes | ||
| 28 | RAD51AP1 | 18264 | -3.474 | -0.2741 | Yes | ||
| 29 | ERCC5 | 18295 | -3.572 | -0.2180 | Yes | ||
| 30 | YBX1 | 18384 | -3.897 | -0.1598 | Yes | ||
| 31 | TP53 | 18610 | -10.661 | 0.0003 | Yes |