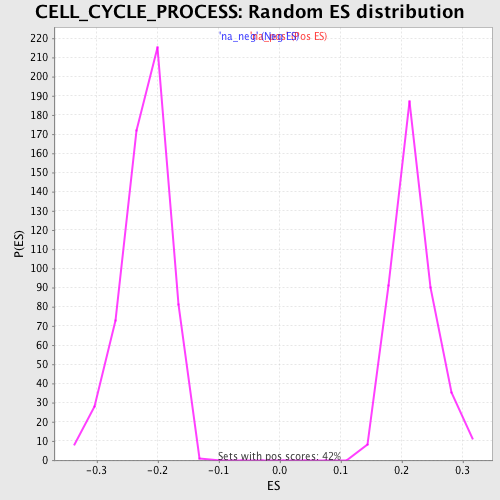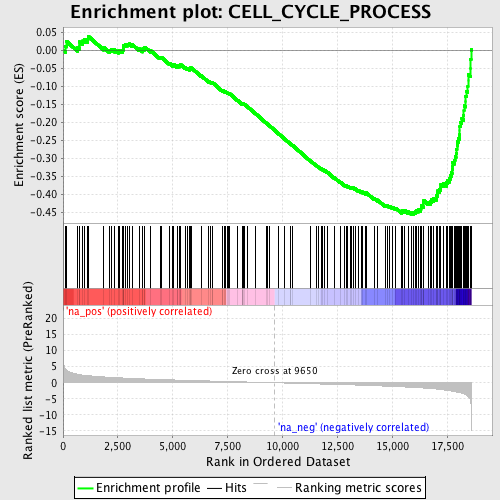
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | CELL_CYCLE_PROCESS |
| Enrichment Score (ES) | -0.4567098 |
| Normalized Enrichment Score (NES) | -2.061225 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.030417418 |
| FWER p-Value | 0.063 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CDKN1C | 103 | 4.297 | 0.0116 | No | ||
| 2 | MSH5 | 154 | 3.949 | 0.0247 | No | ||
| 3 | CDKN2A | 661 | 2.646 | 0.0079 | No | ||
| 4 | PAM | 751 | 2.532 | 0.0132 | No | ||
| 5 | CHMP1A | 754 | 2.528 | 0.0232 | No | ||
| 6 | CDKN2D | 886 | 2.387 | 0.0256 | No | ||
| 7 | PAFAH1B1 | 972 | 2.320 | 0.0303 | No | ||
| 8 | MAD2L1 | 1128 | 2.202 | 0.0307 | No | ||
| 9 | KPNA2 | 1136 | 2.200 | 0.0392 | No | ||
| 10 | CCNA2 | 1853 | 1.799 | 0.0076 | No | ||
| 11 | TIPIN | 2131 | 1.671 | -0.0008 | No | ||
| 12 | CDKN1A | 2200 | 1.649 | 0.0022 | No | ||
| 13 | NPM2 | 2336 | 1.598 | 0.0012 | No | ||
| 14 | NUSAP1 | 2509 | 1.543 | -0.0019 | No | ||
| 15 | PIN1 | 2583 | 1.520 | 0.0002 | No | ||
| 16 | PPP6C | 2701 | 1.485 | -0.0002 | No | ||
| 17 | RAD21 | 2736 | 1.470 | 0.0039 | No | ||
| 18 | TTN | 2754 | 1.461 | 0.0088 | No | ||
| 19 | RAD51L1 | 2761 | 1.459 | 0.0143 | No | ||
| 20 | CUL4A | 2827 | 1.441 | 0.0165 | No | ||
| 21 | LATS2 | 2920 | 1.410 | 0.0172 | No | ||
| 22 | RAD17 | 3013 | 1.378 | 0.0177 | No | ||
| 23 | SUGT1 | 3148 | 1.333 | 0.0158 | No | ||
| 24 | DUSP13 | 3462 | 1.238 | 0.0038 | No | ||
| 25 | CDKN3 | 3626 | 1.190 | -0.0003 | No | ||
| 26 | ATM | 3638 | 1.186 | 0.0039 | No | ||
| 27 | FBXO5 | 3706 | 1.166 | 0.0049 | No | ||
| 28 | CUL5 | 3722 | 1.163 | 0.0088 | No | ||
| 29 | CETN3 | 3986 | 1.097 | -0.0011 | No | ||
| 30 | ZWINT | 4415 | 1.000 | -0.0203 | No | ||
| 31 | ANAPC10 | 4467 | 0.987 | -0.0191 | No | ||
| 32 | TGFA | 4867 | 0.900 | -0.0371 | No | ||
| 33 | KATNA1 | 5004 | 0.872 | -0.0410 | No | ||
| 34 | STMN1 | 5029 | 0.866 | -0.0388 | No | ||
| 35 | TPD52L1 | 5206 | 0.832 | -0.0450 | No | ||
| 36 | NDC80 | 5232 | 0.826 | -0.0431 | No | ||
| 37 | CHFR | 5301 | 0.811 | -0.0435 | No | ||
| 38 | CDC23 | 5353 | 0.800 | -0.0431 | No | ||
| 39 | NEK2 | 5362 | 0.796 | -0.0403 | No | ||
| 40 | PIM2 | 5562 | 0.756 | -0.0481 | No | ||
| 41 | ANAPC11 | 5680 | 0.730 | -0.0515 | No | ||
| 42 | CENTD2 | 5755 | 0.712 | -0.0527 | No | ||
| 43 | APBB2 | 5787 | 0.706 | -0.0515 | No | ||
| 44 | RCC1 | 5794 | 0.704 | -0.0490 | No | ||
| 45 | HSPA2 | 5849 | 0.695 | -0.0492 | No | ||
| 46 | ANLN | 6293 | 0.612 | -0.0707 | No | ||
| 47 | PTPRC | 6637 | 0.548 | -0.0871 | No | ||
| 48 | USH1C | 6713 | 0.535 | -0.0890 | No | ||
| 49 | CHEK1 | 6785 | 0.522 | -0.0908 | No | ||
| 50 | CDKN2B | 6825 | 0.514 | -0.0909 | No | ||
| 51 | RAN | 7264 | 0.422 | -0.1129 | No | ||
| 52 | UBE2C | 7270 | 0.421 | -0.1115 | No | ||
| 53 | MAD2L2 | 7364 | 0.408 | -0.1149 | No | ||
| 54 | BOLL | 7388 | 0.403 | -0.1145 | No | ||
| 55 | CDKN1B | 7498 | 0.383 | -0.1189 | No | ||
| 56 | XRCC2 | 7529 | 0.380 | -0.1190 | No | ||
| 57 | POLA1 | 7588 | 0.368 | -0.1207 | No | ||
| 58 | RB1 | 7963 | 0.297 | -0.1398 | No | ||
| 59 | PRMT5 | 8157 | 0.265 | -0.1491 | No | ||
| 60 | RAD1 | 8160 | 0.265 | -0.1482 | No | ||
| 61 | ANAPC4 | 8162 | 0.265 | -0.1472 | No | ||
| 62 | BUB1 | 8225 | 0.254 | -0.1495 | No | ||
| 63 | CCNA1 | 8247 | 0.249 | -0.1497 | No | ||
| 64 | CDCA5 | 8391 | 0.226 | -0.1565 | No | ||
| 65 | TIMELESS | 8756 | 0.160 | -0.1756 | No | ||
| 66 | POLS | 9273 | 0.071 | -0.2033 | No | ||
| 67 | AURKA | 9293 | 0.067 | -0.2040 | No | ||
| 68 | NPM1 | 9417 | 0.047 | -0.2105 | No | ||
| 69 | CUL2 | 9838 | -0.039 | -0.2331 | No | ||
| 70 | STAG3 | 10080 | -0.083 | -0.2458 | No | ||
| 71 | NEK6 | 10344 | -0.127 | -0.2596 | No | ||
| 72 | CENPE | 10373 | -0.134 | -0.2606 | No | ||
| 73 | CDC16 | 10470 | -0.152 | -0.2652 | No | ||
| 74 | CD28 | 11286 | -0.304 | -0.3081 | No | ||
| 75 | CETN1 | 11531 | -0.350 | -0.3199 | No | ||
| 76 | KIF11 | 11659 | -0.378 | -0.3253 | No | ||
| 77 | BUB1B | 11770 | -0.400 | -0.3296 | No | ||
| 78 | CUL3 | 11837 | -0.412 | -0.3316 | No | ||
| 79 | SYCP1 | 11925 | -0.429 | -0.3346 | No | ||
| 80 | TOP3A | 12047 | -0.450 | -0.3393 | No | ||
| 81 | BIRC5 | 12364 | -0.512 | -0.3544 | No | ||
| 82 | RACGAP1 | 12635 | -0.565 | -0.3668 | No | ||
| 83 | MDM4 | 12836 | -0.607 | -0.3752 | No | ||
| 84 | EPGN | 12897 | -0.619 | -0.3759 | No | ||
| 85 | CDC6 | 12983 | -0.636 | -0.3780 | No | ||
| 86 | TTK | 13086 | -0.658 | -0.3809 | No | ||
| 87 | EGF | 13134 | -0.666 | -0.3808 | No | ||
| 88 | CDK6 | 13219 | -0.686 | -0.3826 | No | ||
| 89 | TGFB1 | 13308 | -0.704 | -0.3845 | No | ||
| 90 | SSSCA1 | 13452 | -0.737 | -0.3893 | No | ||
| 91 | AKAP8 | 13583 | -0.766 | -0.3933 | No | ||
| 92 | DCTN3 | 13654 | -0.783 | -0.3940 | No | ||
| 93 | KIF15 | 13789 | -0.817 | -0.3980 | No | ||
| 94 | SMC4 | 13804 | -0.821 | -0.3954 | No | ||
| 95 | GFI1B | 14183 | -0.915 | -0.4123 | No | ||
| 96 | CUL1 | 14329 | -0.955 | -0.4163 | No | ||
| 97 | MSH4 | 14673 | -1.045 | -0.4307 | No | ||
| 98 | DMC1 | 14768 | -1.072 | -0.4315 | No | ||
| 99 | CDC7 | 14883 | -1.103 | -0.4333 | No | ||
| 100 | ACVR1 | 15033 | -1.143 | -0.4368 | No | ||
| 101 | RAD51L3 | 15159 | -1.181 | -0.4388 | No | ||
| 102 | KNTC1 | 15435 | -1.269 | -0.4486 | No | ||
| 103 | KHDRBS1 | 15450 | -1.270 | -0.4443 | No | ||
| 104 | BCAT1 | 15569 | -1.312 | -0.4455 | No | ||
| 105 | SMC3 | 15722 | -1.364 | -0.4482 | No | ||
| 106 | ZW10 | 15866 | -1.415 | -0.4503 | No | ||
| 107 | RAD52 | 15985 | -1.464 | -0.4509 | Yes | ||
| 108 | TUBE1 | 16068 | -1.496 | -0.4493 | Yes | ||
| 109 | E2F1 | 16090 | -1.502 | -0.4444 | Yes | ||
| 110 | CDK4 | 16191 | -1.548 | -0.4437 | Yes | ||
| 111 | TARDBP | 16295 | -1.582 | -0.4429 | Yes | ||
| 112 | CDK2 | 16312 | -1.588 | -0.4374 | Yes | ||
| 113 | MAP3K11 | 16319 | -1.593 | -0.4314 | Yes | ||
| 114 | CENPF | 16425 | -1.644 | -0.4305 | Yes | ||
| 115 | BRCA2 | 16426 | -1.645 | -0.4239 | Yes | ||
| 116 | CDC25B | 16441 | -1.655 | -0.4180 | Yes | ||
| 117 | NCAPH | 16656 | -1.758 | -0.4226 | Yes | ||
| 118 | PCBP4 | 16759 | -1.815 | -0.4209 | Yes | ||
| 119 | GSPT1 | 16804 | -1.840 | -0.4159 | Yes | ||
| 120 | RAD51 | 16891 | -1.889 | -0.4130 | Yes | ||
| 121 | TBRG4 | 17003 | -1.957 | -0.4112 | Yes | ||
| 122 | CDC25C | 17009 | -1.959 | -0.4036 | Yes | ||
| 123 | POLE | 17055 | -2.001 | -0.3980 | Yes | ||
| 124 | SKP2 | 17079 | -2.016 | -0.3912 | Yes | ||
| 125 | PPP5C | 17161 | -2.096 | -0.3872 | Yes | ||
| 126 | MRE11A | 17211 | -2.133 | -0.3813 | Yes | ||
| 127 | EREG | 17216 | -2.137 | -0.3730 | Yes | ||
| 128 | CIT | 17333 | -2.227 | -0.3704 | Yes | ||
| 129 | TPX2 | 17473 | -2.353 | -0.3685 | Yes | ||
| 130 | GFI1 | 17529 | -2.405 | -0.3619 | Yes | ||
| 131 | ANAPC5 | 17619 | -2.499 | -0.3567 | Yes | ||
| 132 | LIG3 | 17653 | -2.540 | -0.3483 | Yes | ||
| 133 | KIF22 | 17690 | -2.585 | -0.3399 | Yes | ||
| 134 | CDK2AP1 | 17730 | -2.620 | -0.3315 | Yes | ||
| 135 | NOLC1 | 17748 | -2.639 | -0.3219 | Yes | ||
| 136 | KIF23 | 17755 | -2.653 | -0.3116 | Yes | ||
| 137 | PLK1 | 17847 | -2.764 | -0.3055 | Yes | ||
| 138 | NUMA1 | 17864 | -2.791 | -0.2952 | Yes | ||
| 139 | DDX11 | 17909 | -2.844 | -0.2862 | Yes | ||
| 140 | RAD50 | 17930 | -2.875 | -0.2758 | Yes | ||
| 141 | CLIP1 | 17953 | -2.913 | -0.2653 | Yes | ||
| 142 | NBN | 17973 | -2.942 | -0.2546 | Yes | ||
| 143 | CDK10 | 18008 | -2.992 | -0.2444 | Yes | ||
| 144 | SPO11 | 18067 | -3.063 | -0.2353 | Yes | ||
| 145 | PML | 18083 | -3.084 | -0.2238 | Yes | ||
| 146 | ABL1 | 18084 | -3.088 | -0.2114 | Yes | ||
| 147 | MPHOSPH6 | 18116 | -3.151 | -0.2005 | Yes | ||
| 148 | ACVR1B | 18173 | -3.255 | -0.1905 | Yes | ||
| 149 | DCTN2 | 18256 | -3.458 | -0.1811 | Yes | ||
| 150 | DLG7 | 18270 | -3.498 | -0.1678 | Yes | ||
| 151 | CDKN2C | 18274 | -3.503 | -0.1540 | Yes | ||
| 152 | KIF2C | 18349 | -3.764 | -0.1429 | Yes | ||
| 153 | SMC1A | 18351 | -3.769 | -0.1279 | Yes | ||
| 154 | APBB1 | 18379 | -3.882 | -0.1138 | Yes | ||
| 155 | RAD54L | 18448 | -4.193 | -0.1007 | Yes | ||
| 156 | PRC1 | 18454 | -4.207 | -0.0842 | Yes | ||
| 157 | NDE1 | 18455 | -4.231 | -0.0673 | Yes | ||
| 158 | POLD1 | 18575 | -5.904 | -0.0501 | Yes | ||
| 159 | TUBG1 | 18578 | -5.997 | -0.0262 | Yes | ||
| 160 | MYH10 | 18593 | -7.046 | 0.0012 | Yes |