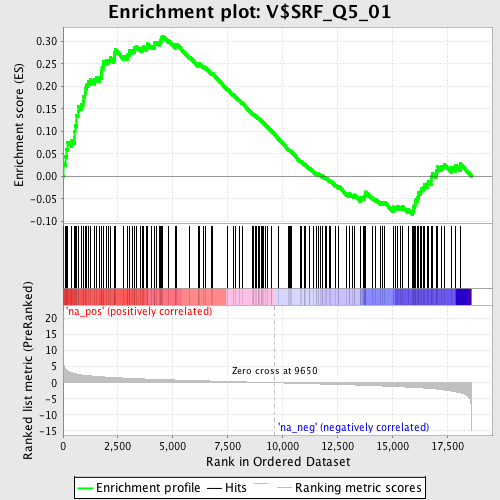
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | V$SRF_Q5_01 |
| Enrichment Score (ES) | 0.31153575 |
| Normalized Enrichment Score (NES) | 1.450136 |
| Nominal p-value | 0.0043290043 |
| FDR q-value | 1.0 |
| FWER p-Value | 0.998 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MYLK | 21 | 6.191 | 0.0286 | Yes | ||
| 2 | ASB2 | 102 | 4.314 | 0.0449 | Yes | ||
| 3 | CNN2 | 166 | 3.876 | 0.0601 | Yes | ||
| 4 | EML4 | 206 | 3.707 | 0.0758 | Yes | ||
| 5 | H3F3A | 401 | 3.070 | 0.0800 | Yes | ||
| 6 | EFHD1 | 528 | 2.853 | 0.0869 | Yes | ||
| 7 | SGK | 535 | 2.839 | 0.1002 | Yes | ||
| 8 | PCDH7 | 542 | 2.824 | 0.1134 | Yes | ||
| 9 | TANK | 617 | 2.704 | 0.1224 | Yes | ||
| 10 | LCP1 | 619 | 2.703 | 0.1353 | Yes | ||
| 11 | MYL9 | 679 | 2.624 | 0.1447 | Yes | ||
| 12 | PICALM | 712 | 2.578 | 0.1553 | Yes | ||
| 13 | MYL6 | 843 | 2.441 | 0.1600 | Yes | ||
| 14 | PELI2 | 908 | 2.375 | 0.1679 | Yes | ||
| 15 | ABR | 935 | 2.346 | 0.1778 | Yes | ||
| 16 | PPARG | 997 | 2.298 | 0.1855 | Yes | ||
| 17 | RBBP7 | 1031 | 2.270 | 0.1946 | Yes | ||
| 18 | CALD1 | 1074 | 2.239 | 0.2031 | Yes | ||
| 19 | TAF5L | 1138 | 2.198 | 0.2102 | Yes | ||
| 20 | RASSF2 | 1243 | 2.131 | 0.2148 | Yes | ||
| 21 | CXCL5 | 1426 | 2.015 | 0.2146 | Yes | ||
| 22 | BLOC1S1 | 1506 | 1.971 | 0.2198 | Yes | ||
| 23 | MITF | 1677 | 1.875 | 0.2195 | Yes | ||
| 24 | SERTAD4 | 1734 | 1.850 | 0.2254 | Yes | ||
| 25 | TAGLN | 1743 | 1.847 | 0.2338 | Yes | ||
| 26 | ERBB2IP | 1771 | 1.835 | 0.2412 | Yes | ||
| 27 | EEF1B2 | 1823 | 1.812 | 0.2471 | Yes | ||
| 28 | MAPK14 | 1847 | 1.802 | 0.2545 | Yes | ||
| 29 | MYH11 | 1956 | 1.745 | 0.2570 | Yes | ||
| 30 | FLNA | 2087 | 1.690 | 0.2581 | Yes | ||
| 31 | STAT5B | 2139 | 1.667 | 0.2633 | Yes | ||
| 32 | MAP1A | 2321 | 1.606 | 0.2612 | Yes | ||
| 33 | H2AFX | 2344 | 1.595 | 0.2677 | Yes | ||
| 34 | ARF6 | 2352 | 1.593 | 0.2749 | Yes | ||
| 35 | CFL1 | 2382 | 1.583 | 0.2809 | Yes | ||
| 36 | TLE3 | 2771 | 1.457 | 0.2669 | Yes | ||
| 37 | CSRP1 | 2918 | 1.410 | 0.2658 | Yes | ||
| 38 | SRF | 2954 | 1.397 | 0.2706 | Yes | ||
| 39 | RSU1 | 3011 | 1.380 | 0.2741 | Yes | ||
| 40 | ITGB6 | 3022 | 1.374 | 0.2802 | Yes | ||
| 41 | NR2F1 | 3152 | 1.332 | 0.2796 | Yes | ||
| 42 | MRPS21 | 3236 | 1.307 | 0.2814 | Yes | ||
| 43 | SCOC | 3261 | 1.301 | 0.2863 | Yes | ||
| 44 | VASP | 3351 | 1.275 | 0.2876 | Yes | ||
| 45 | WNT3 | 3525 | 1.217 | 0.2841 | Yes | ||
| 46 | EMILIN2 | 3634 | 1.188 | 0.2839 | Yes | ||
| 47 | GADD45G | 3647 | 1.183 | 0.2889 | Yes | ||
| 48 | DGKG | 3810 | 1.138 | 0.2856 | Yes | ||
| 49 | THBS1 | 3840 | 1.132 | 0.2895 | Yes | ||
| 50 | NKX2-2 | 3854 | 1.127 | 0.2942 | Yes | ||
| 51 | OTX1 | 4050 | 1.083 | 0.2888 | Yes | ||
| 52 | MYL7 | 4160 | 1.056 | 0.2880 | Yes | ||
| 53 | ELAVL4 | 4175 | 1.052 | 0.2923 | Yes | ||
| 54 | BNC2 | 4186 | 1.050 | 0.2968 | Yes | ||
| 55 | MUS81 | 4271 | 1.033 | 0.2972 | Yes | ||
| 56 | SLC25A4 | 4375 | 1.011 | 0.2964 | Yes | ||
| 57 | VCL | 4419 | 1.000 | 0.2989 | Yes | ||
| 58 | MYADM | 4446 | 0.993 | 0.3022 | Yes | ||
| 59 | THRAP1 | 4483 | 0.983 | 0.3050 | Yes | ||
| 60 | ACTR3 | 4502 | 0.978 | 0.3087 | Yes | ||
| 61 | HOXC5 | 4537 | 0.970 | 0.3115 | Yes | ||
| 62 | SLC4A3 | 4786 | 0.917 | 0.3025 | No | ||
| 63 | GPR20 | 5113 | 0.850 | 0.2889 | No | ||
| 64 | DIXDC1 | 5124 | 0.848 | 0.2924 | No | ||
| 65 | IL17B | 5187 | 0.837 | 0.2931 | No | ||
| 66 | TRDN | 5775 | 0.709 | 0.2647 | No | ||
| 67 | RIMS1 | 6162 | 0.635 | 0.2468 | No | ||
| 68 | HOXD10 | 6167 | 0.635 | 0.2496 | No | ||
| 69 | NRXN3 | 6205 | 0.627 | 0.2506 | No | ||
| 70 | UBE2H | 6378 | 0.595 | 0.2441 | No | ||
| 71 | CORO1C | 6485 | 0.577 | 0.2412 | No | ||
| 72 | FLNC | 6776 | 0.523 | 0.2279 | No | ||
| 73 | MAP2K6 | 6819 | 0.514 | 0.2281 | No | ||
| 74 | CDKN1B | 7498 | 0.383 | 0.1932 | No | ||
| 75 | MID1 | 7751 | 0.340 | 0.1812 | No | ||
| 76 | MAPKAPK2 | 7864 | 0.318 | 0.1766 | No | ||
| 77 | NPPA | 8034 | 0.284 | 0.1688 | No | ||
| 78 | ASPA | 8166 | 0.263 | 0.1630 | No | ||
| 79 | KRTCAP2 | 8640 | 0.183 | 0.1382 | No | ||
| 80 | LRRTM4 | 8661 | 0.178 | 0.1380 | No | ||
| 81 | COL23A1 | 8698 | 0.170 | 0.1368 | No | ||
| 82 | ARPC4 | 8759 | 0.158 | 0.1344 | No | ||
| 83 | SRD5A2 | 8800 | 0.152 | 0.1329 | No | ||
| 84 | STARD13 | 8896 | 0.135 | 0.1284 | No | ||
| 85 | FOXE3 | 8906 | 0.134 | 0.1286 | No | ||
| 86 | CASQ1 | 8960 | 0.124 | 0.1263 | No | ||
| 87 | PDLIM7 | 9042 | 0.110 | 0.1224 | No | ||
| 88 | RERE | 9060 | 0.107 | 0.1220 | No | ||
| 89 | RUNDC1 | 9082 | 0.103 | 0.1214 | No | ||
| 90 | SLC2A4 | 9144 | 0.094 | 0.1185 | No | ||
| 91 | CHST8 | 9240 | 0.077 | 0.1137 | No | ||
| 92 | TNMD | 9328 | 0.062 | 0.1093 | No | ||
| 93 | DLL1 | 9482 | 0.035 | 0.1012 | No | ||
| 94 | PLN | 9490 | 0.034 | 0.1010 | No | ||
| 95 | NRAS | 9797 | -0.029 | 0.0845 | No | ||
| 96 | MYO1E | 10281 | -0.116 | 0.0589 | No | ||
| 97 | KCNA1 | 10319 | -0.122 | 0.0575 | No | ||
| 98 | JUNB | 10366 | -0.133 | 0.0556 | No | ||
| 99 | EGR1 | 10428 | -0.145 | 0.0530 | No | ||
| 100 | ANXA6 | 10814 | -0.213 | 0.0331 | No | ||
| 101 | PADI2 | 10845 | -0.219 | 0.0326 | No | ||
| 102 | MRGPRF | 10992 | -0.245 | 0.0258 | No | ||
| 103 | LPP | 11016 | -0.248 | 0.0258 | No | ||
| 104 | CRH | 11027 | -0.250 | 0.0264 | No | ||
| 105 | PODN | 11221 | -0.293 | 0.0174 | No | ||
| 106 | DUSP10 | 11239 | -0.295 | 0.0179 | No | ||
| 107 | EGR3 | 11427 | -0.329 | 0.0093 | No | ||
| 108 | TGFB1I1 | 11538 | -0.352 | 0.0050 | No | ||
| 109 | DSTN | 11555 | -0.355 | 0.0059 | No | ||
| 110 | ADAMTSL1 | 11625 | -0.368 | 0.0039 | No | ||
| 111 | EGR2 | 11632 | -0.370 | 0.0053 | No | ||
| 112 | KCNH2 | 11712 | -0.389 | 0.0029 | No | ||
| 113 | PHF12 | 11812 | -0.408 | -0.0005 | No | ||
| 114 | NPAS2 | 11819 | -0.410 | 0.0011 | No | ||
| 115 | ACTG2 | 11946 | -0.431 | -0.0036 | No | ||
| 116 | CNN1 | 12004 | -0.441 | -0.0046 | No | ||
| 117 | NR2F2 | 12123 | -0.465 | -0.0088 | No | ||
| 118 | IER2 | 12201 | -0.481 | -0.0106 | No | ||
| 119 | FOXP2 | 12436 | -0.526 | -0.0208 | No | ||
| 120 | NPR3 | 12542 | -0.547 | -0.0239 | No | ||
| 121 | NFATC4 | 12565 | -0.551 | -0.0224 | No | ||
| 122 | CITED2 | 12937 | -0.627 | -0.0395 | No | ||
| 123 | FOSB | 13036 | -0.648 | -0.0417 | No | ||
| 124 | COL1A1 | 13048 | -0.650 | -0.0392 | No | ||
| 125 | TAZ | 13168 | -0.676 | -0.0424 | No | ||
| 126 | CKM | 13270 | -0.696 | -0.0445 | No | ||
| 127 | CACNA1B | 13280 | -0.698 | -0.0417 | No | ||
| 128 | MLLT7 | 13540 | -0.756 | -0.0521 | No | ||
| 129 | ITGB1BP2 | 13548 | -0.757 | -0.0488 | No | ||
| 130 | ACTB | 13575 | -0.764 | -0.0466 | No | ||
| 131 | RHOJ | 13677 | -0.789 | -0.0483 | No | ||
| 132 | TRIM3 | 13724 | -0.800 | -0.0469 | No | ||
| 133 | FGF17 | 13739 | -0.805 | -0.0438 | No | ||
| 134 | RARB | 13766 | -0.811 | -0.0413 | No | ||
| 135 | CHD2 | 13783 | -0.816 | -0.0383 | No | ||
| 136 | SDPR | 13796 | -0.819 | -0.0350 | No | ||
| 137 | LRRFIP1 | 14119 | -0.897 | -0.0482 | No | ||
| 138 | SLC16A6 | 14251 | -0.932 | -0.0508 | No | ||
| 139 | PPAP2B | 14476 | -0.995 | -0.0582 | No | ||
| 140 | ABCA1 | 14551 | -1.018 | -0.0573 | No | ||
| 141 | ASXL2 | 14666 | -1.044 | -0.0585 | No | ||
| 142 | RAI2 | 15041 | -1.146 | -0.0732 | No | ||
| 143 | FGF7 | 15053 | -1.149 | -0.0683 | No | ||
| 144 | TRIM46 | 15163 | -1.182 | -0.0686 | No | ||
| 145 | MBNL1 | 15243 | -1.202 | -0.0671 | No | ||
| 146 | ATF3 | 15382 | -1.250 | -0.0686 | No | ||
| 147 | TPM3 | 15476 | -1.279 | -0.0675 | No | ||
| 148 | HOXB5 | 15729 | -1.366 | -0.0746 | No | ||
| 149 | TIMP3 | 15923 | -1.439 | -0.0781 | No | ||
| 150 | HOXC4 | 15953 | -1.452 | -0.0727 | No | ||
| 151 | FGF8 | 15975 | -1.460 | -0.0669 | No | ||
| 152 | PFN1 | 16037 | -1.483 | -0.0631 | No | ||
| 153 | PPP1R12A | 16058 | -1.494 | -0.0570 | No | ||
| 154 | CALN1 | 16081 | -1.500 | -0.0510 | No | ||
| 155 | TADA3L | 16163 | -1.537 | -0.0480 | No | ||
| 156 | HOXB4 | 16194 | -1.549 | -0.0422 | No | ||
| 157 | GGN | 16209 | -1.553 | -0.0355 | No | ||
| 158 | FGFRL1 | 16305 | -1.585 | -0.0330 | No | ||
| 159 | PRKAB2 | 16353 | -1.606 | -0.0279 | No | ||
| 160 | CDC25B | 16441 | -1.655 | -0.0246 | No | ||
| 161 | CFL2 | 16478 | -1.669 | -0.0186 | No | ||
| 162 | ATF4 | 16604 | -1.731 | -0.0171 | No | ||
| 163 | CA3 | 16646 | -1.754 | -0.0109 | No | ||
| 164 | DVL3 | 16786 | -1.831 | -0.0096 | No | ||
| 165 | GRK6 | 16788 | -1.832 | -0.0009 | No | ||
| 166 | SVIL | 16814 | -1.843 | 0.0066 | No | ||
| 167 | LDB1 | 16997 | -1.951 | 0.0061 | No | ||
| 168 | TPM2 | 17032 | -1.981 | 0.0138 | No | ||
| 169 | FOS | 17081 | -2.016 | 0.0209 | No | ||
| 170 | FOXP1 | 17248 | -2.162 | 0.0222 | No | ||
| 171 | CSDA | 17360 | -2.253 | 0.0270 | No | ||
| 172 | KCNK3 | 17711 | -2.601 | 0.0205 | No | ||
| 173 | ACTA1 | 17879 | -2.807 | 0.0249 | No | ||
| 174 | PLCB3 | 18096 | -3.117 | 0.0282 | No |
