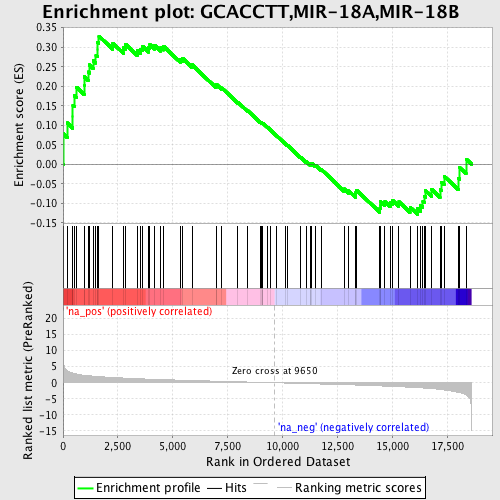
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | GCACCTT,MIR-18A,MIR-18B |
| Enrichment Score (ES) | 0.32842225 |
| Normalized Enrichment Score (NES) | 1.3135933 |
| Nominal p-value | 0.07942974 |
| FDR q-value | 0.95438397 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | KIT | 9 | 8.006 | 0.0785 | Yes | ||
| 2 | JARID1B | 188 | 3.774 | 0.1062 | Yes | ||
| 3 | WSB1 | 446 | 2.988 | 0.1218 | Yes | ||
| 4 | NEDD9 | 447 | 2.983 | 0.1512 | Yes | ||
| 5 | CDC42 | 507 | 2.885 | 0.1765 | Yes | ||
| 6 | ESR1 | 614 | 2.705 | 0.1975 | Yes | ||
| 7 | ZFP36L1 | 960 | 2.328 | 0.2018 | Yes | ||
| 8 | CTGF | 968 | 2.322 | 0.2244 | Yes | ||
| 9 | SOCS5 | 1143 | 2.192 | 0.2366 | Yes | ||
| 10 | ADD3 | 1200 | 2.154 | 0.2548 | Yes | ||
| 11 | NR3C1 | 1370 | 2.054 | 0.2660 | Yes | ||
| 12 | RAB5C | 1496 | 1.974 | 0.2787 | Yes | ||
| 13 | IGF1 | 1567 | 1.934 | 0.2940 | Yes | ||
| 14 | SMAD2 | 1568 | 1.934 | 0.3131 | Yes | ||
| 15 | CRIM1 | 1633 | 1.899 | 0.3284 | Yes | ||
| 16 | ALCAM | 2264 | 1.626 | 0.3105 | No | ||
| 17 | GAB1 | 2755 | 1.461 | 0.2985 | No | ||
| 18 | FCHSD2 | 2848 | 1.433 | 0.3077 | No | ||
| 19 | PACSIN1 | 3376 | 1.265 | 0.2917 | No | ||
| 20 | STK4 | 3535 | 1.212 | 0.2952 | No | ||
| 21 | MESP1 | 3610 | 1.193 | 0.3029 | No | ||
| 22 | GLRB | 3899 | 1.117 | 0.2984 | No | ||
| 23 | SIM2 | 3954 | 1.104 | 0.3064 | No | ||
| 24 | SNURF | 4165 | 1.054 | 0.3055 | No | ||
| 25 | TRIM2 | 4455 | 0.990 | 0.2997 | No | ||
| 26 | SAR1A | 4584 | 0.962 | 0.3023 | No | ||
| 27 | ARL15 | 5358 | 0.798 | 0.2684 | No | ||
| 28 | PIAS3 | 5445 | 0.779 | 0.2715 | No | ||
| 29 | TRIB2 | 5877 | 0.689 | 0.2550 | No | ||
| 30 | TEX2 | 6974 | 0.482 | 0.2007 | No | ||
| 31 | BHLHB5 | 6997 | 0.477 | 0.2042 | No | ||
| 32 | BTG3 | 7233 | 0.428 | 0.1957 | No | ||
| 33 | AEBP2 | 7960 | 0.297 | 0.1595 | No | ||
| 34 | CTDSPL | 8398 | 0.225 | 0.1382 | No | ||
| 35 | MDGA1 | 8986 | 0.119 | 0.1077 | No | ||
| 36 | GCLC | 9027 | 0.112 | 0.1066 | No | ||
| 37 | SH3BP4 | 9081 | 0.103 | 0.1048 | No | ||
| 38 | DPP10 | 9088 | 0.102 | 0.1055 | No | ||
| 39 | HSF2 | 9104 | 0.099 | 0.1056 | No | ||
| 40 | PRKACB | 9310 | 0.065 | 0.0952 | No | ||
| 41 | ZBTB4 | 9435 | 0.045 | 0.0890 | No | ||
| 42 | FNBP1 | 9743 | -0.019 | 0.0726 | No | ||
| 43 | RAB11FIP2 | 10146 | -0.093 | 0.0518 | No | ||
| 44 | IRF2 | 10226 | -0.107 | 0.0486 | No | ||
| 45 | ETV6 | 10825 | -0.216 | 0.0185 | No | ||
| 46 | PTGFRN | 11077 | -0.260 | 0.0075 | No | ||
| 47 | PHF19 | 11274 | -0.300 | -0.0001 | No | ||
| 48 | ATXN1 | 11290 | -0.304 | 0.0021 | No | ||
| 49 | RABGAP1 | 11337 | -0.315 | 0.0027 | No | ||
| 50 | PURB | 11499 | -0.343 | -0.0026 | No | ||
| 51 | RHOT1 | 11756 | -0.396 | -0.0125 | No | ||
| 52 | NCOA1 | 12805 | -0.599 | -0.0631 | No | ||
| 53 | PERQ1 | 13015 | -0.642 | -0.0680 | No | ||
| 54 | PRICKLE2 | 13334 | -0.710 | -0.0782 | No | ||
| 55 | HIF1A | 13342 | -0.712 | -0.0715 | No | ||
| 56 | MAN1A2 | 13377 | -0.719 | -0.0662 | No | ||
| 57 | ANKRD50 | 14438 | -0.982 | -0.1137 | No | ||
| 58 | NAV1 | 14453 | -0.988 | -0.1047 | No | ||
| 59 | CLASP2 | 14460 | -0.990 | -0.0953 | No | ||
| 60 | ASXL2 | 14666 | -1.044 | -0.0960 | No | ||
| 61 | NFAT5 | 14909 | -1.109 | -0.0981 | No | ||
| 62 | XYLT2 | 14998 | -1.134 | -0.0917 | No | ||
| 63 | AKR1D1 | 15307 | -1.222 | -0.0963 | No | ||
| 64 | MEF2D | 15813 | -1.397 | -0.1097 | No | ||
| 65 | LIN28 | 16164 | -1.537 | -0.1134 | No | ||
| 66 | TARDBP | 16295 | -1.582 | -0.1048 | No | ||
| 67 | PHF2 | 16399 | -1.633 | -0.0943 | No | ||
| 68 | SON | 16462 | -1.660 | -0.0812 | No | ||
| 69 | NR1H2 | 16524 | -1.693 | -0.0678 | No | ||
| 70 | FRMD4A | 16802 | -1.839 | -0.0646 | No | ||
| 71 | EHMT1 | 17181 | -2.106 | -0.0642 | No | ||
| 72 | TRIOBP | 17239 | -2.156 | -0.0460 | No | ||
| 73 | RUNX1 | 17368 | -2.260 | -0.0306 | No | ||
| 74 | CLK2 | 18015 | -2.997 | -0.0359 | No | ||
| 75 | ARHGEF5 | 18080 | -3.082 | -0.0089 | No | ||
| 76 | PARP6 | 18367 | -3.828 | 0.0134 | No |
