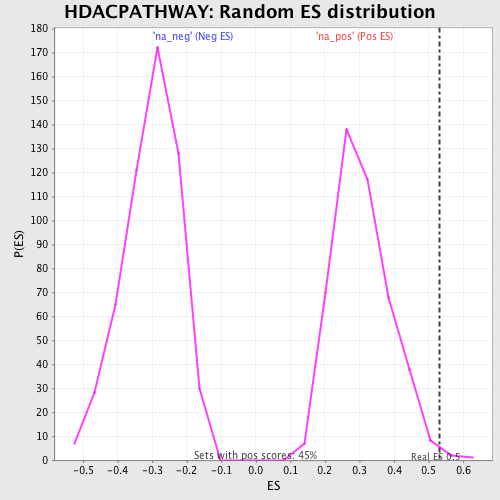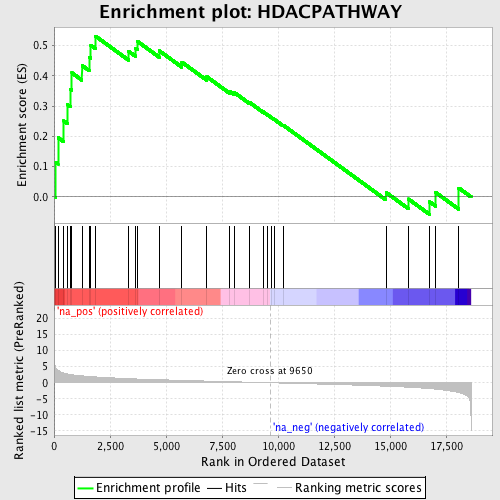
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | HDACPATHWAY |
| Enrichment Score (ES) | 0.5309354 |
| Normalized Enrichment Score (NES) | 1.7299056 |
| Nominal p-value | 0.0066815144 |
| FDR q-value | 0.16391183 |
| FWER p-Value | 0.813 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CAMK1G | 53 | 5.162 | 0.1153 | Yes | ||
| 2 | MEF2A | 178 | 3.816 | 0.1960 | Yes | ||
| 3 | CALM2 | 433 | 3.005 | 0.2511 | Yes | ||
| 4 | PIK3R1 | 600 | 2.724 | 0.3046 | Yes | ||
| 5 | NFATC1 | 739 | 2.547 | 0.3555 | Yes | ||
| 6 | CAMK1 | 793 | 2.494 | 0.4097 | Yes | ||
| 7 | PPP3CA | 1244 | 2.131 | 0.4343 | Yes | ||
| 8 | IGF1 | 1567 | 1.934 | 0.4612 | Yes | ||
| 9 | YWHAH | 1639 | 1.895 | 0.5008 | Yes | ||
| 10 | MAPK14 | 1847 | 1.802 | 0.5309 | Yes | ||
| 11 | MEF2B | 3324 | 1.285 | 0.4810 | No | ||
| 12 | IGF1R | 3633 | 1.189 | 0.4916 | No | ||
| 13 | MEF2C | 3730 | 1.161 | 0.5130 | No | ||
| 14 | NFATC2 | 4695 | 0.936 | 0.4826 | No | ||
| 15 | SYT1 | 5701 | 0.725 | 0.4451 | No | ||
| 16 | MAP2K6 | 6819 | 0.514 | 0.3968 | No | ||
| 17 | MAPK7 | 7846 | 0.321 | 0.3489 | No | ||
| 18 | HDAC5 | 8057 | 0.281 | 0.3441 | No | ||
| 19 | CABIN1 | 8729 | 0.165 | 0.3118 | No | ||
| 20 | AVP | 9326 | 0.062 | 0.2811 | No | ||
| 21 | PIK3CA | 9521 | 0.028 | 0.2713 | No | ||
| 22 | PPP3CB | 9709 | -0.011 | 0.2615 | No | ||
| 23 | MYOD1 | 9821 | -0.035 | 0.2563 | No | ||
| 24 | AKT1 | 10228 | -0.108 | 0.2370 | No | ||
| 25 | CALM3 | 14815 | -1.086 | 0.0151 | No | ||
| 26 | MEF2D | 15813 | -1.397 | -0.0066 | No | ||
| 27 | INSR | 16765 | -1.820 | -0.0161 | No | ||
| 28 | CALM1 | 17037 | -1.987 | 0.0148 | No | ||
| 29 | PPP3CC | 18065 | -3.060 | 0.0296 | No |