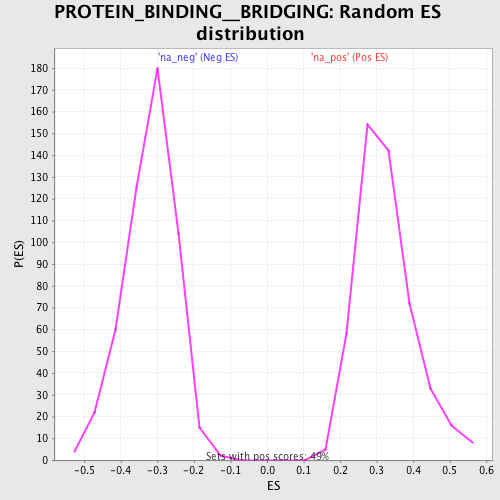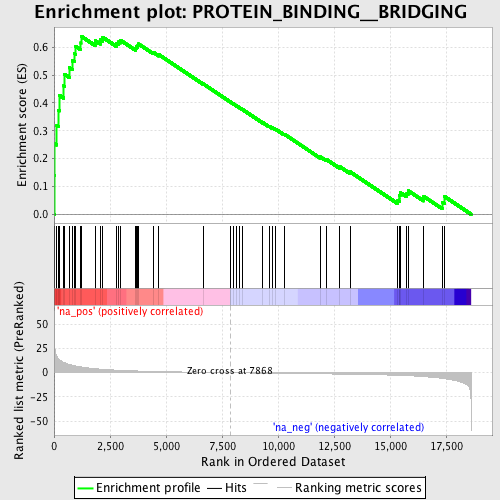
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | PROTEIN_BINDING__BRIDGING |
| Enrichment Score (ES) | 0.64029956 |
| Normalized Enrichment Score (NES) | 1.9735638 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.02422246 |
| FWER p-Value | 0.067 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | VAV3 | 11 | 33.496 | 0.1400 | Yes | ||
| 2 | GAB1 | 24 | 27.320 | 0.2540 | Yes | ||
| 3 | GRB7 | 120 | 16.803 | 0.3194 | Yes | ||
| 4 | LASP1 | 206 | 13.841 | 0.3729 | Yes | ||
| 5 | BLNK | 239 | 13.319 | 0.4271 | Yes | ||
| 6 | LAT | 432 | 10.644 | 0.4615 | Yes | ||
| 7 | SOCS2 | 459 | 10.367 | 0.5036 | Yes | ||
| 8 | SH3BP2 | 676 | 8.459 | 0.5274 | Yes | ||
| 9 | SH2D2A | 813 | 7.647 | 0.5522 | Yes | ||
| 10 | ANXA1 | 903 | 7.188 | 0.5776 | Yes | ||
| 11 | ARHGAP4 | 967 | 6.888 | 0.6031 | Yes | ||
| 12 | SH3BGRL | 1197 | 5.931 | 0.6157 | Yes | ||
| 13 | STAM | 1201 | 5.908 | 0.6403 | Yes | ||
| 14 | SHB | 1838 | 4.155 | 0.6235 | No | ||
| 15 | SH2D3C | 2058 | 3.709 | 0.6273 | No | ||
| 16 | GRB2 | 2166 | 3.530 | 0.6363 | No | ||
| 17 | ARHGAP6 | 2790 | 2.639 | 0.6138 | No | ||
| 18 | CRKL | 2883 | 2.519 | 0.6195 | No | ||
| 19 | CHN2 | 2957 | 2.437 | 0.6257 | No | ||
| 20 | ITSN2 | 3654 | 1.812 | 0.5959 | No | ||
| 21 | GRAP2 | 3668 | 1.800 | 0.6027 | No | ||
| 22 | ABI2 | 3739 | 1.755 | 0.6063 | No | ||
| 23 | IRS4 | 3765 | 1.736 | 0.6123 | No | ||
| 24 | GRB14 | 4423 | 1.337 | 0.5825 | No | ||
| 25 | EVPL | 4679 | 1.186 | 0.5737 | No | ||
| 26 | CHN1 | 6662 | 0.378 | 0.4686 | No | ||
| 27 | SNX9 | 7873 | -0.001 | 0.4034 | No | ||
| 28 | ARPC4 | 7885 | -0.005 | 0.4028 | No | ||
| 29 | SKAP2 | 8006 | -0.044 | 0.3965 | No | ||
| 30 | LOR | 8124 | -0.080 | 0.3906 | No | ||
| 31 | SLA | 8271 | -0.125 | 0.3832 | No | ||
| 32 | SH3BGR | 8406 | -0.164 | 0.3767 | No | ||
| 33 | RUSC1 | 9318 | -0.434 | 0.3295 | No | ||
| 34 | RAD50 | 9604 | -0.516 | 0.3163 | No | ||
| 35 | IVL | 9737 | -0.555 | 0.3115 | No | ||
| 36 | CNKSR1 | 9888 | -0.599 | 0.3059 | No | ||
| 37 | GRAP | 10268 | -0.699 | 0.2885 | No | ||
| 38 | GRB10 | 11883 | -1.163 | 0.2064 | No | ||
| 39 | SRC | 12150 | -1.245 | 0.1973 | No | ||
| 40 | SPRR1A | 12759 | -1.429 | 0.1706 | No | ||
| 41 | STUB1 | 13210 | -1.579 | 0.1529 | No | ||
| 42 | SPRR1B | 15342 | -2.675 | 0.0494 | No | ||
| 43 | ARHGAP1 | 15417 | -2.732 | 0.0569 | No | ||
| 44 | KHDRBS1 | 15419 | -2.733 | 0.0683 | No | ||
| 45 | CNTNAP1 | 15448 | -2.754 | 0.0784 | No | ||
| 46 | BCL3 | 15747 | -3.008 | 0.0749 | No | ||
| 47 | TOB1 | 15802 | -3.090 | 0.0850 | No | ||
| 48 | EPS8 | 16493 | -3.994 | 0.0646 | No | ||
| 49 | ST13 | 17326 | -5.768 | 0.0440 | No | ||
| 50 | FKBP4 | 17422 | -6.059 | 0.0643 | No |