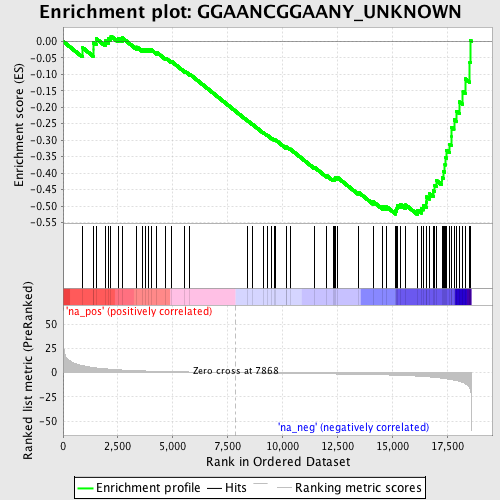
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | GGAANCGGAANY_UNKNOWN |
| Enrichment Score (ES) | -0.5256619 |
| Normalized Enrichment Score (NES) | -1.7453395 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.069161996 |
| FWER p-Value | 0.184 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RRAS | 873 | 7.329 | -0.0184 | No | ||
| 2 | MTA2 | 1370 | 5.306 | -0.0243 | No | ||
| 3 | RPL28 | 1371 | 5.305 | -0.0035 | No | ||
| 4 | SEC24C | 1509 | 4.917 | 0.0083 | No | ||
| 5 | PEX6 | 1923 | 3.975 | 0.0016 | No | ||
| 6 | PTPRCAP | 2075 | 3.678 | 0.0079 | No | ||
| 7 | SDF2 | 2180 | 3.507 | 0.0160 | No | ||
| 8 | BIRC4 | 2510 | 2.995 | 0.0100 | No | ||
| 9 | TRIM39 | 2692 | 2.744 | 0.0110 | No | ||
| 10 | RNF25 | 3362 | 2.053 | -0.0170 | No | ||
| 11 | SFRS16 | 3619 | 1.842 | -0.0236 | No | ||
| 12 | GGA1 | 3776 | 1.728 | -0.0252 | No | ||
| 13 | ATP6V1D | 3882 | 1.650 | -0.0244 | No | ||
| 14 | UBL5 | 4009 | 1.573 | -0.0251 | No | ||
| 15 | TTC17 | 4276 | 1.425 | -0.0338 | No | ||
| 16 | SUPT6H | 4678 | 1.187 | -0.0508 | No | ||
| 17 | FBXO38 | 4951 | 1.061 | -0.0613 | No | ||
| 18 | MTF1 | 5550 | 0.793 | -0.0904 | No | ||
| 19 | PRKAB1 | 5779 | 0.705 | -0.1000 | No | ||
| 20 | ATP6V1E1 | 8400 | -0.162 | -0.2406 | No | ||
| 21 | UBA52 | 8613 | -0.226 | -0.2511 | No | ||
| 22 | MRPL21 | 9139 | -0.381 | -0.2779 | No | ||
| 23 | PARVA | 9311 | -0.432 | -0.2854 | No | ||
| 24 | GYS1 | 9517 | -0.491 | -0.2946 | No | ||
| 25 | BAT4 | 9653 | -0.530 | -0.2998 | No | ||
| 26 | DLX4 | 9682 | -0.537 | -0.2992 | No | ||
| 27 | SRP54 | 10169 | -0.674 | -0.3227 | No | ||
| 28 | BAT3 | 10175 | -0.675 | -0.3204 | No | ||
| 29 | EPN1 | 10346 | -0.723 | -0.3267 | No | ||
| 30 | SNRPE | 11463 | -1.035 | -0.3828 | No | ||
| 31 | FEV | 12009 | -1.199 | -0.4075 | No | ||
| 32 | UGCGL1 | 12310 | -1.289 | -0.4186 | No | ||
| 33 | CSNK2B | 12389 | -1.315 | -0.4177 | No | ||
| 34 | ZDHHC5 | 12419 | -1.324 | -0.4140 | No | ||
| 35 | VPS16 | 12511 | -1.352 | -0.4137 | No | ||
| 36 | MBD1 | 13460 | -1.677 | -0.4582 | No | ||
| 37 | SMUG1 | 14133 | -1.959 | -0.4868 | No | ||
| 38 | PSMB4 | 14569 | -2.181 | -0.5017 | No | ||
| 39 | IGHMBP2 | 14750 | -2.278 | -0.5024 | No | ||
| 40 | EIF2S1 | 15160 | -2.538 | -0.5145 | Yes | ||
| 41 | U2AF2 | 15211 | -2.574 | -0.5072 | Yes | ||
| 42 | SEC61G | 15239 | -2.594 | -0.4985 | Yes | ||
| 43 | TAF10 | 15368 | -2.693 | -0.4948 | Yes | ||
| 44 | MRPS21 | 15611 | -2.886 | -0.4965 | Yes | ||
| 45 | DDX55 | 16152 | -3.474 | -0.5121 | Yes | ||
| 46 | EIF3S3 | 16327 | -3.726 | -0.5068 | Yes | ||
| 47 | SEC61A1 | 16425 | -3.874 | -0.4969 | Yes | ||
| 48 | SNIP1 | 16551 | -4.081 | -0.4876 | Yes | ||
| 49 | TFB2M | 16564 | -4.101 | -0.4722 | Yes | ||
| 50 | MRPL43 | 16690 | -4.340 | -0.4620 | Yes | ||
| 51 | COMMD6 | 16870 | -4.708 | -0.4532 | Yes | ||
| 52 | DPM1 | 16937 | -4.825 | -0.4378 | Yes | ||
| 53 | GNB2L1 | 17014 | -4.974 | -0.4224 | Yes | ||
| 54 | PDAP1 | 17268 | -5.641 | -0.4140 | Yes | ||
| 55 | HSPH1 | 17324 | -5.763 | -0.3944 | Yes | ||
| 56 | NCBP2 | 17401 | -6.011 | -0.3749 | Yes | ||
| 57 | COX7A2 | 17413 | -6.040 | -0.3519 | Yes | ||
| 58 | BANF1 | 17469 | -6.226 | -0.3304 | Yes | ||
| 59 | PEO1 | 17620 | -6.769 | -0.3120 | Yes | ||
| 60 | RPL38 | 17691 | -7.055 | -0.2882 | Yes | ||
| 61 | PRPF3 | 17696 | -7.071 | -0.2607 | Yes | ||
| 62 | MRPS18A | 17816 | -7.611 | -0.2373 | Yes | ||
| 63 | MRPS23 | 17929 | -8.115 | -0.2115 | Yes | ||
| 64 | RUVBL2 | 18063 | -8.937 | -0.1837 | Yes | ||
| 65 | MAP4K1 | 18224 | -10.205 | -0.1523 | Yes | ||
| 66 | EBNA1BP2 | 18326 | -11.235 | -0.1138 | Yes | ||
| 67 | TIMM8A | 18538 | -15.604 | -0.0640 | Yes | ||
| 68 | RARS | 18565 | -17.404 | 0.0027 | Yes |
