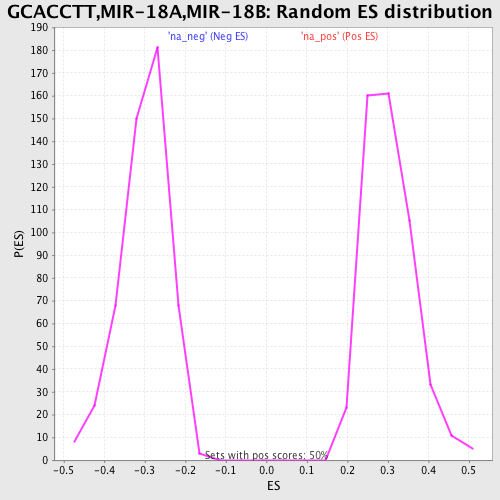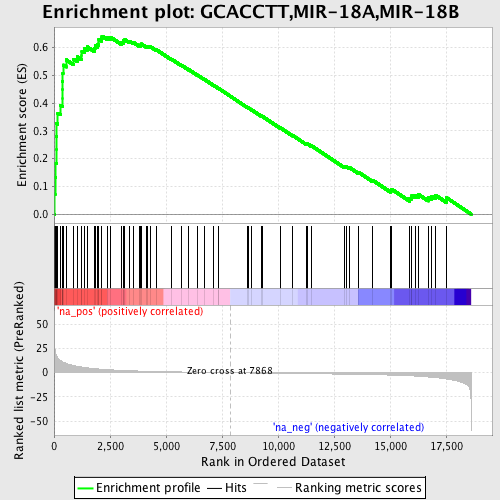
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | GCACCTT,MIR-18A,MIR-18B |
| Enrichment Score (ES) | 0.6417697 |
| Normalized Enrichment Score (NES) | 2.1116664 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0016616541 |
| FWER p-Value | 0.0010 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GAB1 | 24 | 27.320 | 0.0728 | Yes | ||
| 2 | KIT | 44 | 22.388 | 0.1324 | Yes | ||
| 3 | JARID1B | 76 | 19.731 | 0.1843 | Yes | ||
| 4 | ARHGEF5 | 97 | 18.082 | 0.2322 | Yes | ||
| 5 | FRMD4A | 101 | 17.980 | 0.2808 | Yes | ||
| 6 | ESR1 | 115 | 17.309 | 0.3270 | Yes | ||
| 7 | NEDD9 | 165 | 14.716 | 0.3643 | Yes | ||
| 8 | TRIOBP | 290 | 12.449 | 0.3913 | Yes | ||
| 9 | RUNX1 | 369 | 11.391 | 0.4180 | Yes | ||
| 10 | CTDSPL | 371 | 11.384 | 0.4488 | Yes | ||
| 11 | ETV6 | 375 | 11.290 | 0.4793 | Yes | ||
| 12 | PIAS3 | 389 | 11.066 | 0.5086 | Yes | ||
| 13 | PRKACB | 400 | 10.923 | 0.5377 | Yes | ||
| 14 | ZFP36L1 | 536 | 9.560 | 0.5563 | Yes | ||
| 15 | NFAT5 | 871 | 7.334 | 0.5582 | Yes | ||
| 16 | TEX2 | 1036 | 6.575 | 0.5671 | Yes | ||
| 17 | IGF1 | 1238 | 5.779 | 0.5720 | Yes | ||
| 18 | EHMT1 | 1241 | 5.761 | 0.5875 | Yes | ||
| 19 | ATXN1 | 1347 | 5.374 | 0.5964 | Yes | ||
| 20 | RAB5C | 1475 | 5.035 | 0.6032 | Yes | ||
| 21 | FCHSD2 | 1797 | 4.240 | 0.5974 | Yes | ||
| 22 | IRF2 | 1831 | 4.176 | 0.6069 | Yes | ||
| 23 | ALCAM | 1949 | 3.924 | 0.6112 | Yes | ||
| 24 | PHF2 | 1995 | 3.830 | 0.6192 | Yes | ||
| 25 | STK4 | 1996 | 3.829 | 0.6296 | Yes | ||
| 26 | PURB | 2106 | 3.628 | 0.6335 | Yes | ||
| 27 | PACSIN1 | 2134 | 3.572 | 0.6418 | Yes | ||
| 28 | ANKRD50 | 2382 | 3.162 | 0.6370 | No | ||
| 29 | RABGAP1 | 2518 | 2.986 | 0.6378 | No | ||
| 30 | PARP6 | 3006 | 2.389 | 0.6180 | No | ||
| 31 | NR3C1 | 3088 | 2.317 | 0.6200 | No | ||
| 32 | NCOA1 | 3097 | 2.308 | 0.6258 | No | ||
| 33 | CDC42 | 3158 | 2.237 | 0.6286 | No | ||
| 34 | WSB1 | 3372 | 2.039 | 0.6227 | No | ||
| 35 | SOCS5 | 3531 | 1.913 | 0.6193 | No | ||
| 36 | AEBP2 | 3818 | 1.699 | 0.6085 | No | ||
| 37 | CLK2 | 3874 | 1.657 | 0.6100 | No | ||
| 38 | PERQ1 | 3890 | 1.643 | 0.6137 | No | ||
| 39 | CTGF | 4138 | 1.509 | 0.6044 | No | ||
| 40 | SH3BP4 | 4190 | 1.476 | 0.6057 | No | ||
| 41 | NR1H2 | 4308 | 1.403 | 0.6032 | No | ||
| 42 | SAR1A | 4560 | 1.245 | 0.5930 | No | ||
| 43 | SON | 5246 | 0.921 | 0.5586 | No | ||
| 44 | GCLC | 5696 | 0.734 | 0.5363 | No | ||
| 45 | LIN28 | 6013 | 0.622 | 0.5210 | No | ||
| 46 | SMAD2 | 6395 | 0.473 | 0.5017 | No | ||
| 47 | HIF1A | 6695 | 0.367 | 0.4866 | No | ||
| 48 | MESP1 | 7111 | 0.230 | 0.4648 | No | ||
| 49 | PTGFRN | 7343 | 0.156 | 0.4528 | No | ||
| 50 | SIM2 | 7346 | 0.155 | 0.4531 | No | ||
| 51 | ARL15 | 8641 | -0.237 | 0.3840 | No | ||
| 52 | PHF19 | 8657 | -0.243 | 0.3838 | No | ||
| 53 | SNURF | 8831 | -0.296 | 0.3753 | No | ||
| 54 | BHLHB5 | 9241 | -0.411 | 0.3543 | No | ||
| 55 | ASXL2 | 9300 | -0.428 | 0.3524 | No | ||
| 56 | PRICKLE2 | 10091 | -0.657 | 0.3115 | No | ||
| 57 | CLASP2 | 10648 | -0.809 | 0.2838 | No | ||
| 58 | MAN1A2 | 11283 | -0.983 | 0.2522 | No | ||
| 59 | MDGA1 | 11291 | -0.985 | 0.2545 | No | ||
| 60 | RAB11FIP2 | 11487 | -1.040 | 0.2468 | No | ||
| 61 | GLRB | 12972 | -1.501 | 0.1708 | No | ||
| 62 | DPP10 | 13049 | -1.522 | 0.1709 | No | ||
| 63 | AKR1D1 | 13200 | -1.576 | 0.1671 | No | ||
| 64 | MEF2D | 13598 | -1.728 | 0.1503 | No | ||
| 65 | RHOT1 | 14206 | -1.993 | 0.1230 | No | ||
| 66 | ZBTB4 | 15024 | -2.444 | 0.0856 | No | ||
| 67 | NAV1 | 15070 | -2.469 | 0.0898 | No | ||
| 68 | XYLT2 | 15871 | -3.160 | 0.0552 | No | ||
| 69 | TRIB2 | 15938 | -3.234 | 0.0605 | No | ||
| 70 | HSF2 | 15971 | -3.266 | 0.0676 | No | ||
| 71 | CRIM1 | 16133 | -3.445 | 0.0682 | No | ||
| 72 | BTG3 | 16276 | -3.651 | 0.0705 | No | ||
| 73 | ADD3 | 16707 | -4.375 | 0.0591 | No | ||
| 74 | TRIM2 | 16845 | -4.660 | 0.0644 | No | ||
| 75 | FNBP1 | 17020 | -4.986 | 0.0685 | No | ||
| 76 | TARDBP | 17527 | -6.456 | 0.0587 | No |