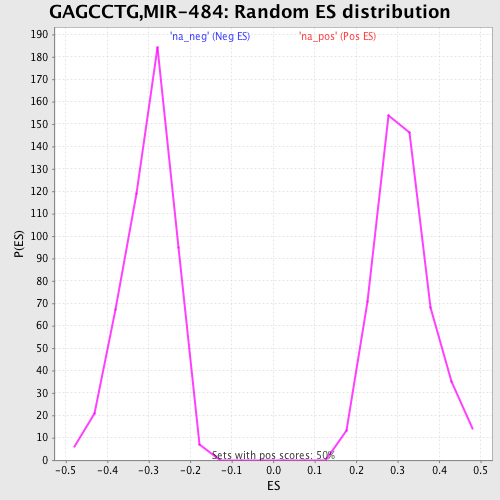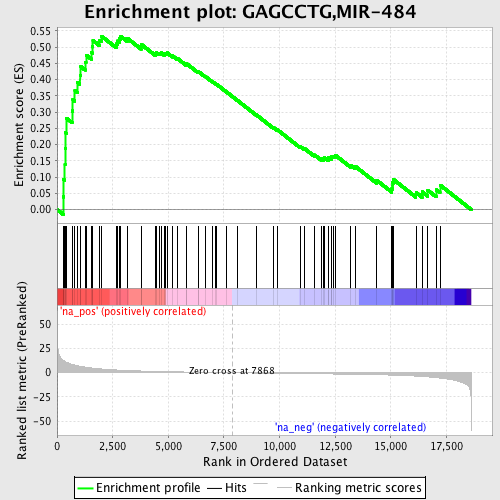
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | GAGCCTG,MIR-484 |
| Enrichment Score (ES) | 0.5338681 |
| Normalized Enrichment Score (NES) | 1.7088125 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.04248097 |
| FWER p-Value | 0.406 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MAPKAPK2 | 276 | 12.560 | 0.0394 | Yes | ||
| 2 | TRIOBP | 290 | 12.449 | 0.0925 | Yes | ||
| 3 | TSGA14 | 353 | 11.617 | 0.1393 | Yes | ||
| 4 | BCL11A | 362 | 11.533 | 0.1887 | Yes | ||
| 5 | MAP3K11 | 373 | 11.326 | 0.2371 | Yes | ||
| 6 | ZNRF1 | 419 | 10.759 | 0.2812 | Yes | ||
| 7 | RAB6IP1 | 677 | 8.457 | 0.3039 | Yes | ||
| 8 | MAP4K2 | 708 | 8.299 | 0.3381 | Yes | ||
| 9 | ZFYVE1 | 801 | 7.729 | 0.3666 | Yes | ||
| 10 | NFE2L1 | 914 | 7.145 | 0.3914 | Yes | ||
| 11 | PRKCB1 | 1028 | 6.604 | 0.4138 | Yes | ||
| 12 | HOXA5 | 1065 | 6.487 | 0.4399 | Yes | ||
| 13 | MTF2 | 1263 | 5.659 | 0.4538 | Yes | ||
| 14 | HIC2 | 1301 | 5.527 | 0.4757 | Yes | ||
| 15 | ACVR1B | 1553 | 4.804 | 0.4829 | Yes | ||
| 16 | DPYSL2 | 1581 | 4.741 | 0.5019 | Yes | ||
| 17 | CENPB | 1611 | 4.668 | 0.5205 | Yes | ||
| 18 | DAG1 | 1918 | 3.983 | 0.5212 | Yes | ||
| 19 | NFKBIZ | 1992 | 3.836 | 0.5339 | Yes | ||
| 20 | DND1 | 2660 | 2.785 | 0.5099 | No | ||
| 21 | PHC2 | 2712 | 2.718 | 0.5189 | No | ||
| 22 | NFIA | 2793 | 2.635 | 0.5260 | No | ||
| 23 | LBH | 2861 | 2.542 | 0.5334 | No | ||
| 24 | HIPK1 | 3149 | 2.247 | 0.5276 | No | ||
| 25 | TRIM33 | 3775 | 1.728 | 0.5014 | No | ||
| 26 | TCF1 | 3777 | 1.726 | 0.5088 | No | ||
| 27 | PTPRF | 4425 | 1.335 | 0.4797 | No | ||
| 28 | YTHDF3 | 4453 | 1.310 | 0.4839 | No | ||
| 29 | PCDH7 | 4595 | 1.223 | 0.4815 | No | ||
| 30 | SCARA3 | 4669 | 1.191 | 0.4827 | No | ||
| 31 | JMJD2A | 4816 | 1.122 | 0.4797 | No | ||
| 32 | PITPNA | 4880 | 1.094 | 0.4811 | No | ||
| 33 | EIF2C1 | 4939 | 1.066 | 0.4825 | No | ||
| 34 | EZH1 | 5185 | 0.954 | 0.4734 | No | ||
| 35 | FGF1 | 5403 | 0.858 | 0.4654 | No | ||
| 36 | RAF1 | 5794 | 0.702 | 0.4475 | No | ||
| 37 | EDA | 5809 | 0.696 | 0.4497 | No | ||
| 38 | PGBD5 | 6344 | 0.489 | 0.4230 | No | ||
| 39 | DLL4 | 6355 | 0.486 | 0.4246 | No | ||
| 40 | MINK1 | 6663 | 0.378 | 0.4097 | No | ||
| 41 | PLEKHH2 | 6967 | 0.278 | 0.3945 | No | ||
| 42 | DCBLD2 | 7109 | 0.230 | 0.3879 | No | ||
| 43 | PTGER4 | 7157 | 0.217 | 0.3863 | No | ||
| 44 | SEMA4G | 7594 | 0.088 | 0.3632 | No | ||
| 45 | LYPLA2 | 8093 | -0.073 | 0.3366 | No | ||
| 46 | ZIC5 | 8945 | -0.329 | 0.2922 | No | ||
| 47 | MLLT7 | 9747 | -0.557 | 0.2514 | No | ||
| 48 | PRPF4B | 9901 | -0.602 | 0.2457 | No | ||
| 49 | SLC6A1 | 10934 | -0.886 | 0.1939 | No | ||
| 50 | ZIC1 | 11114 | -0.935 | 0.1883 | No | ||
| 51 | COL12A1 | 11571 | -1.065 | 0.1683 | No | ||
| 52 | GRB10 | 11883 | -1.163 | 0.1566 | No | ||
| 53 | HSPG2 | 11966 | -1.190 | 0.1573 | No | ||
| 54 | DACH1 | 12019 | -1.204 | 0.1597 | No | ||
| 55 | NFATC4 | 12203 | -1.261 | 0.1553 | No | ||
| 56 | TEX261 | 12208 | -1.262 | 0.1605 | No | ||
| 57 | CBL | 12327 | -1.294 | 0.1597 | No | ||
| 58 | PALM2 | 12340 | -1.299 | 0.1647 | No | ||
| 59 | IPO11 | 12443 | -1.332 | 0.1650 | No | ||
| 60 | KLF12 | 12519 | -1.353 | 0.1668 | No | ||
| 61 | SERTAD1 | 13195 | -1.574 | 0.1372 | No | ||
| 62 | SP6 | 13411 | -1.656 | 0.1327 | No | ||
| 63 | TRPS1 | 14365 | -2.071 | 0.0903 | No | ||
| 64 | BTG4 | 15048 | -2.459 | 0.0641 | No | ||
| 65 | NAV1 | 15070 | -2.469 | 0.0737 | No | ||
| 66 | HIVEP2 | 15075 | -2.473 | 0.0841 | No | ||
| 67 | DGCR8 | 15122 | -2.510 | 0.0925 | No | ||
| 68 | M6PR | 16131 | -3.443 | 0.0530 | No | ||
| 69 | PTPRE | 16407 | -3.847 | 0.0548 | No | ||
| 70 | DOLPP1 | 16661 | -4.280 | 0.0597 | No | ||
| 71 | EIF4G2 | 17044 | -5.050 | 0.0609 | No | ||
| 72 | TYSND1 | 17213 | -5.511 | 0.0757 | No |