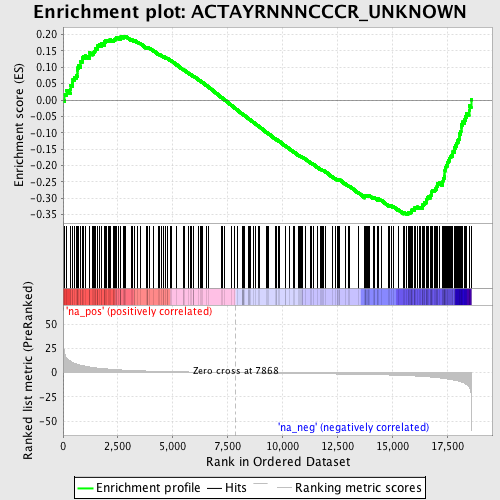
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | ACTAYRNNNCCCR_UNKNOWN |
| Enrichment Score (ES) | -0.3494351 |
| Normalized Enrichment Score (NES) | -1.360297 |
| Nominal p-value | 0.009302326 |
| FDR q-value | 0.3026845 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RAB8B | 68 | 20.364 | 0.0182 | No | ||
| 2 | ARID5A | 157 | 15.029 | 0.0296 | No | ||
| 3 | BACE1 | 324 | 12.035 | 0.0335 | No | ||
| 4 | TLE4 | 336 | 11.842 | 0.0456 | No | ||
| 5 | HBP1 | 422 | 10.739 | 0.0525 | No | ||
| 6 | INVS | 445 | 10.521 | 0.0626 | No | ||
| 7 | SESN1 | 522 | 9.731 | 0.0690 | No | ||
| 8 | RALBP1 | 597 | 9.033 | 0.0746 | No | ||
| 9 | PDE6D | 663 | 8.528 | 0.0803 | No | ||
| 10 | PPARGC1A | 665 | 8.507 | 0.0894 | No | ||
| 11 | MAF1 | 673 | 8.470 | 0.0981 | No | ||
| 12 | RBL2 | 689 | 8.374 | 0.1063 | No | ||
| 13 | ITCH | 800 | 7.737 | 0.1086 | No | ||
| 14 | ZFYVE1 | 801 | 7.729 | 0.1169 | No | ||
| 15 | RDH11 | 866 | 7.376 | 0.1214 | No | ||
| 16 | RRAS | 873 | 7.329 | 0.1289 | No | ||
| 17 | ZFP28 | 933 | 7.061 | 0.1333 | No | ||
| 18 | RABAC1 | 1031 | 6.597 | 0.1351 | No | ||
| 19 | ITPR1 | 1193 | 5.947 | 0.1327 | No | ||
| 20 | ZDHHC14 | 1203 | 5.904 | 0.1386 | No | ||
| 21 | KATNAL1 | 1205 | 5.898 | 0.1449 | No | ||
| 22 | CDCA3 | 1341 | 5.394 | 0.1433 | No | ||
| 23 | ZCCHC8 | 1398 | 5.228 | 0.1459 | No | ||
| 24 | RDH12 | 1435 | 5.128 | 0.1494 | No | ||
| 25 | FBXO9 | 1457 | 5.070 | 0.1538 | No | ||
| 26 | RAB5C | 1475 | 5.035 | 0.1582 | No | ||
| 27 | SNX17 | 1554 | 4.804 | 0.1592 | No | ||
| 28 | WRN | 1579 | 4.748 | 0.1630 | No | ||
| 29 | ORMDL3 | 1582 | 4.741 | 0.1680 | No | ||
| 30 | WBP2 | 1642 | 4.586 | 0.1697 | No | ||
| 31 | IRF2BP1 | 1739 | 4.355 | 0.1691 | No | ||
| 32 | POLK | 1753 | 4.326 | 0.1731 | No | ||
| 33 | TBX6 | 1876 | 4.078 | 0.1708 | No | ||
| 34 | PHF12 | 1877 | 4.077 | 0.1752 | No | ||
| 35 | HMG20A | 1887 | 4.052 | 0.1791 | No | ||
| 36 | PTDSS2 | 1930 | 3.962 | 0.1810 | No | ||
| 37 | DCTN2 | 1975 | 3.877 | 0.1828 | No | ||
| 38 | NUMA1 | 2062 | 3.697 | 0.1821 | No | ||
| 39 | CTF1 | 2109 | 3.624 | 0.1835 | No | ||
| 40 | COVA1 | 2148 | 3.551 | 0.1852 | No | ||
| 41 | OPHN1 | 2283 | 3.333 | 0.1815 | No | ||
| 42 | SETMAR | 2315 | 3.277 | 0.1834 | No | ||
| 43 | RPL15 | 2344 | 3.228 | 0.1853 | No | ||
| 44 | TRPC4AP | 2381 | 3.164 | 0.1867 | No | ||
| 45 | THAP11 | 2411 | 3.123 | 0.1885 | No | ||
| 46 | PPP2R5B | 2425 | 3.105 | 0.1912 | No | ||
| 47 | PLEKHB1 | 2512 | 2.994 | 0.1897 | No | ||
| 48 | USP48 | 2625 | 2.824 | 0.1866 | No | ||
| 49 | VTI1B | 2627 | 2.823 | 0.1896 | No | ||
| 50 | MFN2 | 2630 | 2.818 | 0.1925 | No | ||
| 51 | ZNF235 | 2636 | 2.811 | 0.1953 | No | ||
| 52 | SLC25A27 | 2748 | 2.685 | 0.1921 | No | ||
| 53 | SLC35B4 | 2784 | 2.645 | 0.1931 | No | ||
| 54 | CMAS | 2798 | 2.630 | 0.1952 | No | ||
| 55 | ETFDH | 2863 | 2.539 | 0.1944 | No | ||
| 56 | CBLL1 | 3111 | 2.290 | 0.1834 | No | ||
| 57 | CYB561D2 | 3150 | 2.246 | 0.1838 | No | ||
| 58 | SIRT2 | 3242 | 2.155 | 0.1811 | No | ||
| 59 | FGFR1 | 3267 | 2.130 | 0.1821 | No | ||
| 60 | SERTAD3 | 3406 | 2.009 | 0.1768 | No | ||
| 61 | MAX | 3505 | 1.931 | 0.1735 | No | ||
| 62 | FNTB | 3812 | 1.701 | 0.1587 | No | ||
| 63 | KDELC1 | 3832 | 1.689 | 0.1595 | No | ||
| 64 | NCOA3 | 3859 | 1.670 | 0.1598 | No | ||
| 65 | UBE2H | 3862 | 1.664 | 0.1615 | No | ||
| 66 | TP53 | 3947 | 1.611 | 0.1587 | No | ||
| 67 | AP2M1 | 4137 | 1.509 | 0.1500 | No | ||
| 68 | BRMS1L | 4351 | 1.379 | 0.1399 | No | ||
| 69 | DNAJB12 | 4399 | 1.349 | 0.1388 | No | ||
| 70 | ZCCHC7 | 4468 | 1.305 | 0.1365 | No | ||
| 71 | NUP153 | 4572 | 1.239 | 0.1322 | No | ||
| 72 | RPS11 | 4594 | 1.223 | 0.1324 | No | ||
| 73 | SFXN5 | 4681 | 1.185 | 0.1290 | No | ||
| 74 | TUSC4 | 4746 | 1.158 | 0.1267 | No | ||
| 75 | MITF | 4754 | 1.153 | 0.1276 | No | ||
| 76 | NFKBIB | 4891 | 1.090 | 0.1213 | No | ||
| 77 | DQX1 | 4948 | 1.062 | 0.1194 | No | ||
| 78 | SCAMP3 | 5186 | 0.954 | 0.1075 | No | ||
| 79 | RPL7 | 5478 | 0.827 | 0.0926 | No | ||
| 80 | NUDT13 | 5487 | 0.822 | 0.0930 | No | ||
| 81 | ABLIM2 | 5547 | 0.793 | 0.0907 | No | ||
| 82 | HCFC1R1 | 5729 | 0.720 | 0.0816 | No | ||
| 83 | ZNF8 | 5802 | 0.699 | 0.0784 | No | ||
| 84 | RPS27A | 5857 | 0.683 | 0.0762 | No | ||
| 85 | ACBD5 | 5929 | 0.656 | 0.0731 | No | ||
| 86 | TSC2 | 5954 | 0.645 | 0.0724 | No | ||
| 87 | PACRG | 5960 | 0.642 | 0.0729 | No | ||
| 88 | TBN | 6157 | 0.559 | 0.0628 | No | ||
| 89 | CPEB3 | 6244 | 0.526 | 0.0587 | No | ||
| 90 | PGM2L1 | 6320 | 0.497 | 0.0551 | No | ||
| 91 | BIVM | 6365 | 0.483 | 0.0532 | No | ||
| 92 | FBXW11 | 6527 | 0.426 | 0.0449 | No | ||
| 93 | DPM3 | 6645 | 0.383 | 0.0390 | No | ||
| 94 | SMARCAL1 | 7225 | 0.190 | 0.0076 | No | ||
| 95 | PPP1R7 | 7259 | 0.180 | 0.0060 | No | ||
| 96 | NUDT12 | 7340 | 0.156 | 0.0018 | No | ||
| 97 | SNX1 | 7662 | 0.059 | -0.0156 | No | ||
| 98 | RAB3C | 7806 | 0.016 | -0.0233 | No | ||
| 99 | NTAN1 | 7933 | -0.017 | -0.0302 | No | ||
| 100 | BET1 | 8167 | -0.093 | -0.0428 | No | ||
| 101 | GRK4 | 8172 | -0.096 | -0.0429 | No | ||
| 102 | ITSN1 | 8188 | -0.099 | -0.0436 | No | ||
| 103 | FANCG | 8202 | -0.102 | -0.0442 | No | ||
| 104 | PDCD10 | 8204 | -0.102 | -0.0441 | No | ||
| 105 | BUB1B | 8240 | -0.115 | -0.0459 | No | ||
| 106 | HSD17B4 | 8251 | -0.119 | -0.0463 | No | ||
| 107 | ADAM22 | 8432 | -0.170 | -0.0560 | No | ||
| 108 | BRCA1 | 8441 | -0.173 | -0.0562 | No | ||
| 109 | RDH14 | 8487 | -0.190 | -0.0585 | No | ||
| 110 | AAMP | 8516 | -0.198 | -0.0598 | No | ||
| 111 | PPP6C | 8531 | -0.201 | -0.0603 | No | ||
| 112 | RNF10 | 8690 | -0.253 | -0.0687 | No | ||
| 113 | WDFY2 | 8772 | -0.279 | -0.0728 | No | ||
| 114 | TMEM4 | 8892 | -0.311 | -0.0789 | No | ||
| 115 | MUS81 | 8939 | -0.327 | -0.0811 | No | ||
| 116 | ACTR1A | 9255 | -0.415 | -0.0978 | No | ||
| 117 | ASXL2 | 9300 | -0.428 | -0.0997 | No | ||
| 118 | FDPS | 9379 | -0.451 | -0.1035 | No | ||
| 119 | STCH | 9664 | -0.533 | -0.1184 | No | ||
| 120 | ZNF228 | 9706 | -0.546 | -0.1200 | No | ||
| 121 | MLLT7 | 9747 | -0.557 | -0.1216 | No | ||
| 122 | PANK3 | 9800 | -0.573 | -0.1238 | No | ||
| 123 | MAPBPIP | 9863 | -0.589 | -0.1266 | No | ||
| 124 | PURG | 10129 | -0.664 | -0.1403 | No | ||
| 125 | RPS6KB2 | 10135 | -0.665 | -0.1398 | No | ||
| 126 | DHX30 | 10336 | -0.720 | -0.1499 | No | ||
| 127 | JARID2 | 10491 | -0.764 | -0.1575 | No | ||
| 128 | BTRC | 10556 | -0.783 | -0.1602 | No | ||
| 129 | MAP3K10 | 10737 | -0.835 | -0.1691 | No | ||
| 130 | WASF1 | 10777 | -0.845 | -0.1703 | No | ||
| 131 | RPP21 | 10815 | -0.852 | -0.1714 | No | ||
| 132 | MAP2K6 | 10855 | -0.863 | -0.1726 | No | ||
| 133 | ADCK1 | 10900 | -0.877 | -0.1740 | No | ||
| 134 | COL4A3BP | 10926 | -0.884 | -0.1744 | No | ||
| 135 | NOS3 | 11056 | -0.918 | -0.1805 | No | ||
| 136 | CRYZL1 | 11281 | -0.982 | -0.1916 | No | ||
| 137 | COMMD9 | 11337 | -1.000 | -0.1935 | No | ||
| 138 | GDAP1 | 11391 | -1.016 | -0.1953 | No | ||
| 139 | STMN1 | 11601 | -1.074 | -0.2056 | No | ||
| 140 | RABL3 | 11731 | -1.117 | -0.2114 | No | ||
| 141 | KIAA0427 | 11770 | -1.128 | -0.2122 | No | ||
| 142 | PARK2 | 11825 | -1.144 | -0.2140 | No | ||
| 143 | IPO7 | 11849 | -1.155 | -0.2140 | No | ||
| 144 | TRIP10 | 11975 | -1.193 | -0.2195 | No | ||
| 145 | ZFYVE19 | 12275 | -1.280 | -0.2344 | No | ||
| 146 | ZFP1 | 12404 | -1.320 | -0.2399 | No | ||
| 147 | PHF13 | 12505 | -1.351 | -0.2439 | No | ||
| 148 | IPO13 | 12508 | -1.351 | -0.2426 | No | ||
| 149 | WDTC1 | 12562 | -1.364 | -0.2440 | No | ||
| 150 | BANP | 12603 | -1.376 | -0.2447 | No | ||
| 151 | PRCC | 12614 | -1.379 | -0.2438 | No | ||
| 152 | HACE1 | 12855 | -1.462 | -0.2553 | No | ||
| 153 | FEN1 | 13023 | -1.515 | -0.2627 | No | ||
| 154 | PSMD11 | 13068 | -1.528 | -0.2635 | No | ||
| 155 | PCM1 | 13467 | -1.679 | -0.2834 | No | ||
| 156 | UPF3B | 13757 | -1.792 | -0.2972 | No | ||
| 157 | UBL3 | 13759 | -1.793 | -0.2953 | No | ||
| 158 | SDCCAG8 | 13760 | -1.795 | -0.2934 | No | ||
| 159 | BUB3 | 13770 | -1.799 | -0.2919 | No | ||
| 160 | APBA3 | 13835 | -1.826 | -0.2935 | No | ||
| 161 | DNAJC12 | 13854 | -1.835 | -0.2925 | No | ||
| 162 | GSK3A | 13913 | -1.860 | -0.2936 | No | ||
| 163 | KCNE4 | 13925 | -1.865 | -0.2922 | No | ||
| 164 | SESN2 | 13944 | -1.875 | -0.2912 | No | ||
| 165 | SMUG1 | 14133 | -1.959 | -0.2993 | No | ||
| 166 | SMCR8 | 14176 | -1.978 | -0.2995 | No | ||
| 167 | LIG1 | 14187 | -1.982 | -0.2979 | No | ||
| 168 | ATP6V1F | 14341 | -2.061 | -0.3040 | No | ||
| 169 | RPIA | 14350 | -2.064 | -0.3022 | No | ||
| 170 | DTX2 | 14358 | -2.068 | -0.3004 | No | ||
| 171 | SNX16 | 14498 | -2.141 | -0.3056 | No | ||
| 172 | RPL6 | 14837 | -2.330 | -0.3215 | No | ||
| 173 | COX7C | 14883 | -2.355 | -0.3215 | No | ||
| 174 | TNPO2 | 14951 | -2.400 | -0.3225 | No | ||
| 175 | HDAC8 | 15056 | -2.461 | -0.3256 | No | ||
| 176 | PASK | 15264 | -2.613 | -0.3340 | No | ||
| 177 | PAFAH2 | 15521 | -2.805 | -0.3449 | No | ||
| 178 | PRIM1 | 15556 | -2.840 | -0.3437 | No | ||
| 179 | RAD51L1 | 15643 | -2.910 | -0.3453 | No | ||
| 180 | TIAL1 | 15720 | -2.984 | -0.3462 | Yes | ||
| 181 | NKIRAS1 | 15723 | -2.986 | -0.3431 | Yes | ||
| 182 | MDH1 | 15786 | -3.067 | -0.3432 | Yes | ||
| 183 | USP5 | 15824 | -3.116 | -0.3419 | Yes | ||
| 184 | RNF121 | 15876 | -3.163 | -0.3412 | Yes | ||
| 185 | BCKDK | 15885 | -3.171 | -0.3383 | Yes | ||
| 186 | UBE2N | 15900 | -3.182 | -0.3356 | Yes | ||
| 187 | SKP2 | 15923 | -3.216 | -0.3334 | Yes | ||
| 188 | UFD1L | 16010 | -3.308 | -0.3345 | Yes | ||
| 189 | TOP3A | 16030 | -3.331 | -0.3319 | Yes | ||
| 190 | RANGAP1 | 16032 | -3.337 | -0.3284 | Yes | ||
| 191 | RDH10 | 16067 | -3.376 | -0.3266 | Yes | ||
| 192 | M6PR | 16131 | -3.443 | -0.3263 | Yes | ||
| 193 | NMT1 | 16241 | -3.602 | -0.3284 | Yes | ||
| 194 | COX7B | 16303 | -3.686 | -0.3278 | Yes | ||
| 195 | FKBPL | 16374 | -3.792 | -0.3275 | Yes | ||
| 196 | SLC29A2 | 16376 | -3.795 | -0.3235 | Yes | ||
| 197 | DFFA | 16392 | -3.819 | -0.3202 | Yes | ||
| 198 | LCMT1 | 16426 | -3.876 | -0.3178 | Yes | ||
| 199 | TRIM23 | 16467 | -3.950 | -0.3157 | Yes | ||
| 200 | CDC25A | 16478 | -3.968 | -0.3120 | Yes | ||
| 201 | TMOD3 | 16540 | -4.066 | -0.3110 | Yes | ||
| 202 | CDK9 | 16560 | -4.094 | -0.3076 | Yes | ||
| 203 | RPP40 | 16575 | -4.118 | -0.3039 | Yes | ||
| 204 | SERPINI1 | 16587 | -4.143 | -0.3001 | Yes | ||
| 205 | SYT3 | 16616 | -4.191 | -0.2971 | Yes | ||
| 206 | DNM1L | 16666 | -4.286 | -0.2952 | Yes | ||
| 207 | VPS41 | 16742 | -4.438 | -0.2945 | Yes | ||
| 208 | MRPL54 | 16762 | -4.482 | -0.2907 | Yes | ||
| 209 | SUFU | 16799 | -4.561 | -0.2877 | Yes | ||
| 210 | VKORC1 | 16801 | -4.569 | -0.2829 | Yes | ||
| 211 | GCN5L2 | 16809 | -4.588 | -0.2783 | Yes | ||
| 212 | PSMC4 | 16841 | -4.655 | -0.2750 | Yes | ||
| 213 | CFL1 | 16942 | -4.834 | -0.2753 | Yes | ||
| 214 | YWHAG | 16965 | -4.874 | -0.2712 | Yes | ||
| 215 | ZW10 | 17016 | -4.975 | -0.2686 | Yes | ||
| 216 | ABCC5 | 17037 | -5.036 | -0.2643 | Yes | ||
| 217 | CYP39A1 | 17060 | -5.085 | -0.2600 | Yes | ||
| 218 | HSPC171 | 17068 | -5.102 | -0.2549 | Yes | ||
| 219 | EXO1 | 17135 | -5.314 | -0.2528 | Yes | ||
| 220 | DPYSL5 | 17279 | -5.667 | -0.2545 | Yes | ||
| 221 | PABPC1 | 17309 | -5.730 | -0.2499 | Yes | ||
| 222 | PA2G4 | 17316 | -5.750 | -0.2440 | Yes | ||
| 223 | HSPH1 | 17324 | -5.763 | -0.2382 | Yes | ||
| 224 | TTC13 | 17369 | -5.923 | -0.2342 | Yes | ||
| 225 | KCNN3 | 17396 | -6.000 | -0.2292 | Yes | ||
| 226 | QTRTD1 | 17398 | -6.003 | -0.2228 | Yes | ||
| 227 | LGMN | 17399 | -6.003 | -0.2164 | Yes | ||
| 228 | CALR3 | 17407 | -6.025 | -0.2103 | Yes | ||
| 229 | FKBP4 | 17422 | -6.059 | -0.2045 | Yes | ||
| 230 | CDC26 | 17452 | -6.169 | -0.1994 | Yes | ||
| 231 | IMMT | 17501 | -6.374 | -0.1952 | Yes | ||
| 232 | CREBL1 | 17525 | -6.454 | -0.1895 | Yes | ||
| 233 | DDX25 | 17564 | -6.582 | -0.1845 | Yes | ||
| 234 | CDC6 | 17598 | -6.709 | -0.1791 | Yes | ||
| 235 | RNASEH2A | 17633 | -6.837 | -0.1736 | Yes | ||
| 236 | MRPL50 | 17681 | -7.030 | -0.1686 | Yes | ||
| 237 | SFRS2 | 17755 | -7.315 | -0.1647 | Yes | ||
| 238 | CDC45L | 17767 | -7.357 | -0.1574 | Yes | ||
| 239 | PPRC1 | 17835 | -7.677 | -0.1528 | Yes | ||
| 240 | ING3 | 17844 | -7.748 | -0.1449 | Yes | ||
| 241 | ZNF24 | 17885 | -7.931 | -0.1385 | Yes | ||
| 242 | NR1D1 | 17942 | -8.198 | -0.1328 | Yes | ||
| 243 | PRPF4 | 17956 | -8.274 | -0.1246 | Yes | ||
| 244 | RUTBC3 | 18027 | -8.691 | -0.1191 | Yes | ||
| 245 | POP1 | 18075 | -9.018 | -0.1119 | Yes | ||
| 246 | TTLL4 | 18081 | -9.064 | -0.1024 | Yes | ||
| 247 | HRSP12 | 18124 | -9.360 | -0.0947 | Yes | ||
| 248 | MRPL19 | 18138 | -9.542 | -0.0851 | Yes | ||
| 249 | SH3BGRL2 | 18143 | -9.581 | -0.0750 | Yes | ||
| 250 | MRPL34 | 18189 | -9.934 | -0.0668 | Yes | ||
| 251 | EIF2B4 | 18287 | -10.786 | -0.0605 | Yes | ||
| 252 | PET112L | 18339 | -11.373 | -0.0510 | Yes | ||
| 253 | SHMT2 | 18386 | -12.073 | -0.0406 | Yes | ||
| 254 | GRWD1 | 18501 | -13.937 | -0.0318 | Yes | ||
| 255 | NTHL1 | 18509 | -14.161 | -0.0169 | Yes | ||
| 256 | POLR1B | 18596 | -21.115 | 0.0011 | Yes |
