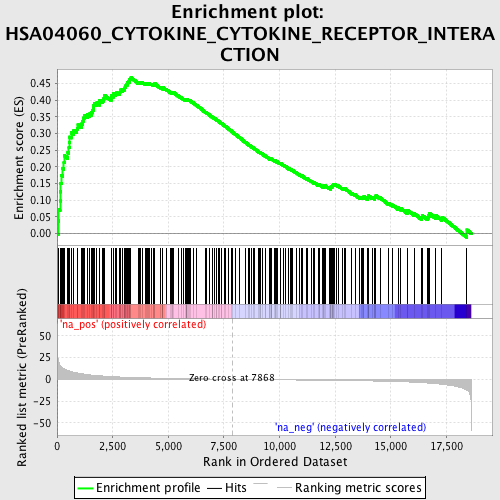
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | HSA04060_CYTOKINE_CYTOKINE_RECEPTOR_INTERACTION |
| Enrichment Score (ES) | 0.46831685 |
| Normalized Enrichment Score (NES) | 1.7783175 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.045043647 |
| FWER p-Value | 0.527 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | KIT | 44 | 22.388 | 0.0379 | Yes | ||
| 2 | IL15 | 74 | 19.919 | 0.0721 | Yes | ||
| 3 | IL28RA | 132 | 15.988 | 0.0978 | Yes | ||
| 4 | CD27 | 142 | 15.671 | 0.1255 | Yes | ||
| 5 | IL3RA | 162 | 14.808 | 0.1511 | Yes | ||
| 6 | IL17RA | 197 | 14.044 | 0.1745 | Yes | ||
| 7 | IFNAR2 | 256 | 12.952 | 0.1947 | Yes | ||
| 8 | PRLR | 307 | 12.224 | 0.2139 | Yes | ||
| 9 | TNFRSF18 | 339 | 11.745 | 0.2334 | Yes | ||
| 10 | TNFRSF1A | 483 | 10.110 | 0.2438 | Yes | ||
| 11 | TNFSF10 | 532 | 9.625 | 0.2585 | Yes | ||
| 12 | IL5 | 548 | 9.406 | 0.2746 | Yes | ||
| 13 | VEGFA | 573 | 9.227 | 0.2899 | Yes | ||
| 14 | IL7R | 631 | 8.739 | 0.3025 | Yes | ||
| 15 | IL12A | 753 | 8.069 | 0.3104 | Yes | ||
| 16 | TGFBR1 | 893 | 7.218 | 0.3159 | Yes | ||
| 17 | LTB | 937 | 7.029 | 0.3262 | Yes | ||
| 18 | ACVR1 | 1096 | 6.352 | 0.3290 | Yes | ||
| 19 | TNFRSF19 | 1149 | 6.146 | 0.3372 | Yes | ||
| 20 | TNFSF11 | 1170 | 6.036 | 0.3470 | Yes | ||
| 21 | TGFB1 | 1215 | 5.864 | 0.3552 | Yes | ||
| 22 | CCL4 | 1364 | 5.312 | 0.3567 | Yes | ||
| 23 | IL7 | 1474 | 5.042 | 0.3598 | Yes | ||
| 24 | ACVR1B | 1553 | 4.804 | 0.3642 | Yes | ||
| 25 | TNFSF9 | 1597 | 4.704 | 0.3703 | Yes | ||
| 26 | IL21R | 1628 | 4.617 | 0.3770 | Yes | ||
| 27 | CNTF | 1647 | 4.564 | 0.3843 | Yes | ||
| 28 | CCL27 | 1670 | 4.517 | 0.3912 | Yes | ||
| 29 | CSF2RA | 1789 | 4.251 | 0.3924 | Yes | ||
| 30 | EGF | 1914 | 3.994 | 0.3929 | Yes | ||
| 31 | IFNGR2 | 1916 | 3.987 | 0.4000 | Yes | ||
| 32 | CXCR3 | 2051 | 3.723 | 0.3994 | Yes | ||
| 33 | TNFRSF21 | 2083 | 3.667 | 0.4043 | Yes | ||
| 34 | CTF1 | 2109 | 3.624 | 0.4095 | Yes | ||
| 35 | PDGFRB | 2130 | 3.579 | 0.4148 | Yes | ||
| 36 | IL18RAP | 2427 | 3.105 | 0.4043 | Yes | ||
| 37 | IFNGR1 | 2431 | 3.101 | 0.4097 | Yes | ||
| 38 | CLCF1 | 2437 | 3.094 | 0.4150 | Yes | ||
| 39 | CX3CR1 | 2535 | 2.958 | 0.4151 | Yes | ||
| 40 | TNFRSF1B | 2536 | 2.957 | 0.4204 | Yes | ||
| 41 | CXCL12 | 2637 | 2.810 | 0.4200 | Yes | ||
| 42 | CCL25 | 2682 | 2.751 | 0.4226 | Yes | ||
| 43 | TNFSF13 | 2800 | 2.629 | 0.4209 | Yes | ||
| 44 | IL10RB | 2822 | 2.604 | 0.4245 | Yes | ||
| 45 | CXCR4 | 2830 | 2.592 | 0.4288 | Yes | ||
| 46 | CRLF2 | 2869 | 2.533 | 0.4313 | Yes | ||
| 47 | IL6ST | 2945 | 2.449 | 0.4316 | Yes | ||
| 48 | CCL5 | 3009 | 2.386 | 0.4324 | Yes | ||
| 49 | CSF2 | 3037 | 2.356 | 0.4352 | Yes | ||
| 50 | CXCL5 | 3049 | 2.346 | 0.4388 | Yes | ||
| 51 | XCL1 | 3074 | 2.325 | 0.4417 | Yes | ||
| 52 | LEPR | 3092 | 2.312 | 0.4450 | Yes | ||
| 53 | IL13RA1 | 3110 | 2.292 | 0.4482 | Yes | ||
| 54 | IL8RB | 3154 | 2.241 | 0.4498 | Yes | ||
| 55 | IL11 | 3166 | 2.229 | 0.4533 | Yes | ||
| 56 | CSF3R | 3189 | 2.206 | 0.4560 | Yes | ||
| 57 | EPOR | 3243 | 2.155 | 0.4570 | Yes | ||
| 58 | CCL3 | 3251 | 2.146 | 0.4605 | Yes | ||
| 59 | ACVR2A | 3262 | 2.133 | 0.4638 | Yes | ||
| 60 | TNFSF14 | 3289 | 2.113 | 0.4662 | Yes | ||
| 61 | CSF1R | 3320 | 2.091 | 0.4683 | Yes | ||
| 62 | TNFRSF14 | 3647 | 1.816 | 0.4539 | No | ||
| 63 | IFNAR1 | 3707 | 1.777 | 0.4539 | No | ||
| 64 | EPO | 3761 | 1.738 | 0.4541 | No | ||
| 65 | TNF | 3842 | 1.682 | 0.4528 | No | ||
| 66 | PPBP | 3950 | 1.609 | 0.4499 | No | ||
| 67 | FAS | 4002 | 1.576 | 0.4499 | No | ||
| 68 | IL1B | 4040 | 1.558 | 0.4507 | No | ||
| 69 | BMP7 | 4112 | 1.521 | 0.4496 | No | ||
| 70 | CCL2 | 4155 | 1.500 | 0.4500 | No | ||
| 71 | IL9R | 4226 | 1.449 | 0.4488 | No | ||
| 72 | KITLG | 4320 | 1.398 | 0.4463 | No | ||
| 73 | IL20 | 4327 | 1.396 | 0.4485 | No | ||
| 74 | HGF | 4370 | 1.364 | 0.4486 | No | ||
| 75 | CXCL2 | 4383 | 1.357 | 0.4504 | No | ||
| 76 | IL17RB | 4668 | 1.191 | 0.4371 | No | ||
| 77 | IL23R | 4731 | 1.167 | 0.4359 | No | ||
| 78 | INHBE | 4734 | 1.164 | 0.4378 | No | ||
| 79 | IL1R1 | 4749 | 1.155 | 0.4392 | No | ||
| 80 | CCL8 | 4913 | 1.079 | 0.4322 | No | ||
| 81 | CCL28 | 5103 | 0.992 | 0.4238 | No | ||
| 82 | IL22RA1 | 5162 | 0.965 | 0.4223 | No | ||
| 83 | TNFSF8 | 5175 | 0.959 | 0.4234 | No | ||
| 84 | LTBR | 5191 | 0.951 | 0.4243 | No | ||
| 85 | LTA | 5223 | 0.932 | 0.4243 | No | ||
| 86 | CCR5 | 5434 | 0.843 | 0.4144 | No | ||
| 87 | IFNA13 | 5587 | 0.779 | 0.4075 | No | ||
| 88 | BMPR1B | 5701 | 0.730 | 0.4027 | No | ||
| 89 | IL18 | 5767 | 0.708 | 0.4005 | No | ||
| 90 | EDA | 5809 | 0.696 | 0.3995 | No | ||
| 91 | CXCR6 | 5812 | 0.695 | 0.4006 | No | ||
| 92 | IL22 | 5819 | 0.693 | 0.4015 | No | ||
| 93 | IL23A | 5833 | 0.690 | 0.4021 | No | ||
| 94 | TSLP | 5847 | 0.686 | 0.4026 | No | ||
| 95 | GHR | 5906 | 0.664 | 0.4006 | No | ||
| 96 | PF4 | 5939 | 0.652 | 0.4001 | No | ||
| 97 | IFNB1 | 5987 | 0.632 | 0.3987 | No | ||
| 98 | TNFRSF13B | 6114 | 0.576 | 0.3928 | No | ||
| 99 | CCL19 | 6277 | 0.515 | 0.3850 | No | ||
| 100 | IL4R | 6677 | 0.373 | 0.3640 | No | ||
| 101 | CCR6 | 6705 | 0.364 | 0.3631 | No | ||
| 102 | IL1R2 | 6862 | 0.310 | 0.3552 | No | ||
| 103 | LEP | 6966 | 0.279 | 0.3501 | No | ||
| 104 | TNFSF15 | 6989 | 0.270 | 0.3494 | No | ||
| 105 | IL25 | 7005 | 0.264 | 0.3491 | No | ||
| 106 | IL12B | 7068 | 0.243 | 0.3461 | No | ||
| 107 | ACVR2B | 7178 | 0.210 | 0.3406 | No | ||
| 108 | TPO | 7181 | 0.209 | 0.3409 | No | ||
| 109 | IL21 | 7272 | 0.176 | 0.3363 | No | ||
| 110 | CCR4 | 7303 | 0.169 | 0.3350 | No | ||
| 111 | EDA2R | 7380 | 0.148 | 0.3311 | No | ||
| 112 | TNFRSF13C | 7404 | 0.141 | 0.3301 | No | ||
| 113 | BMP2 | 7510 | 0.111 | 0.3246 | No | ||
| 114 | EDAR | 7581 | 0.092 | 0.3210 | No | ||
| 115 | FASLG | 7706 | 0.045 | 0.3143 | No | ||
| 116 | TNFRSF11B | 7831 | 0.009 | 0.3076 | No | ||
| 117 | IL8RA | 7903 | -0.010 | 0.3037 | No | ||
| 118 | TNFRSF4 | 8004 | -0.043 | 0.2984 | No | ||
| 119 | CCL1 | 8183 | -0.098 | 0.2889 | No | ||
| 120 | IL4 | 8484 | -0.187 | 0.2729 | No | ||
| 121 | IL17A | 8622 | -0.230 | 0.2659 | No | ||
| 122 | IFNK | 8638 | -0.236 | 0.2655 | No | ||
| 123 | CXCL10 | 8733 | -0.265 | 0.2609 | No | ||
| 124 | PRL | 8814 | -0.294 | 0.2570 | No | ||
| 125 | IL5RA | 8867 | -0.304 | 0.2548 | No | ||
| 126 | CD40LG | 9073 | -0.363 | 0.2443 | No | ||
| 127 | CX3CL1 | 9096 | -0.369 | 0.2437 | No | ||
| 128 | GDF5 | 9153 | -0.385 | 0.2414 | No | ||
| 129 | IL19 | 9162 | -0.388 | 0.2416 | No | ||
| 130 | CCR2 | 9217 | -0.402 | 0.2394 | No | ||
| 131 | NGFR | 9360 | -0.448 | 0.2325 | No | ||
| 132 | CD70 | 9365 | -0.448 | 0.2331 | No | ||
| 133 | IFNA4 | 9550 | -0.501 | 0.2240 | No | ||
| 134 | INHBB | 9553 | -0.501 | 0.2248 | No | ||
| 135 | PDGFB | 9591 | -0.513 | 0.2237 | No | ||
| 136 | CCL24 | 9623 | -0.522 | 0.2230 | No | ||
| 137 | MPL | 9624 | -0.522 | 0.2239 | No | ||
| 138 | TGFB3 | 9645 | -0.526 | 0.2238 | No | ||
| 139 | CCR3 | 9766 | -0.562 | 0.2183 | No | ||
| 140 | IL2RG | 9795 | -0.571 | 0.2178 | No | ||
| 141 | CXCL1 | 9802 | -0.573 | 0.2185 | No | ||
| 142 | IFNA14 | 9880 | -0.596 | 0.2154 | No | ||
| 143 | CXCL16 | 9884 | -0.597 | 0.2163 | No | ||
| 144 | KDR | 10022 | -0.636 | 0.2100 | No | ||
| 145 | IFNA5 | 10028 | -0.638 | 0.2108 | No | ||
| 146 | IFNA7 | 10043 | -0.642 | 0.2112 | No | ||
| 147 | INHBC | 10184 | -0.679 | 0.2048 | No | ||
| 148 | BMPR2 | 10269 | -0.699 | 0.2015 | No | ||
| 149 | CCR9 | 10422 | -0.744 | 0.1946 | No | ||
| 150 | CCL11 | 10495 | -0.766 | 0.1921 | No | ||
| 151 | IL18R1 | 10522 | -0.772 | 0.1921 | No | ||
| 152 | CXCL14 | 10601 | -0.793 | 0.1892 | No | ||
| 153 | CCR8 | 10762 | -0.841 | 0.1821 | No | ||
| 154 | CCL17 | 10902 | -0.878 | 0.1761 | No | ||
| 155 | MET | 10966 | -0.894 | 0.1743 | No | ||
| 156 | IL6 | 11007 | -0.906 | 0.1737 | No | ||
| 157 | AMH | 11199 | -0.958 | 0.1651 | No | ||
| 158 | CCL7 | 11241 | -0.969 | 0.1646 | No | ||
| 159 | IL17B | 11424 | -1.026 | 0.1565 | No | ||
| 160 | IFNG | 11523 | -1.051 | 0.1531 | No | ||
| 161 | EGFR | 11555 | -1.062 | 0.1533 | No | ||
| 162 | IL2 | 11727 | -1.116 | 0.1460 | No | ||
| 163 | PDGFC | 11771 | -1.129 | 0.1457 | No | ||
| 164 | IL9 | 11790 | -1.135 | 0.1468 | No | ||
| 165 | TNFSF18 | 11949 | -1.182 | 0.1403 | No | ||
| 166 | TNFSF12 | 11990 | -1.196 | 0.1403 | No | ||
| 167 | IL24 | 11995 | -1.197 | 0.1422 | No | ||
| 168 | LIFR | 12005 | -1.199 | 0.1439 | No | ||
| 169 | FLT4 | 12043 | -1.210 | 0.1441 | No | ||
| 170 | TNFSF4 | 12224 | -1.265 | 0.1366 | No | ||
| 171 | CCL22 | 12292 | -1.284 | 0.1352 | No | ||
| 172 | PDGFRA | 12296 | -1.285 | 0.1374 | No | ||
| 173 | CNTFR | 12298 | -1.286 | 0.1397 | No | ||
| 174 | OSMR | 12314 | -1.290 | 0.1412 | No | ||
| 175 | IFNA6 | 12359 | -1.306 | 0.1411 | No | ||
| 176 | TNFRSF11A | 12379 | -1.312 | 0.1424 | No | ||
| 177 | CXCL13 | 12382 | -1.313 | 0.1447 | No | ||
| 178 | TNFRSF25 | 12413 | -1.322 | 0.1454 | No | ||
| 179 | IL1A | 12427 | -1.327 | 0.1471 | No | ||
| 180 | CXCL11 | 12471 | -1.340 | 0.1472 | No | ||
| 181 | IL3 | 12552 | -1.361 | 0.1453 | No | ||
| 182 | TGFBR2 | 12638 | -1.390 | 0.1432 | No | ||
| 183 | IFNE1 | 12826 | -1.452 | 0.1356 | No | ||
| 184 | CXCL9 | 12907 | -1.481 | 0.1339 | No | ||
| 185 | TNFRSF8 | 12940 | -1.490 | 0.1349 | No | ||
| 186 | CSF3 | 13246 | -1.592 | 0.1212 | No | ||
| 187 | FLT3 | 13392 | -1.646 | 0.1163 | No | ||
| 188 | TNFRSF9 | 13599 | -1.728 | 0.1082 | No | ||
| 189 | BMPR1A | 13669 | -1.753 | 0.1076 | No | ||
| 190 | FLT1 | 13723 | -1.780 | 0.1079 | No | ||
| 191 | IL6R | 13769 | -1.799 | 0.1087 | No | ||
| 192 | CSF1 | 13790 | -1.808 | 0.1108 | No | ||
| 193 | IFNA2 | 13947 | -1.878 | 0.1057 | No | ||
| 194 | IL12RB2 | 13975 | -1.890 | 0.1077 | No | ||
| 195 | IFNA1 | 13979 | -1.891 | 0.1109 | No | ||
| 196 | TNFRSF17 | 14003 | -1.902 | 0.1131 | No | ||
| 197 | IL15RA | 14179 | -1.979 | 0.1071 | No | ||
| 198 | IL10RA | 14265 | -2.018 | 0.1061 | No | ||
| 199 | TNFSF13B | 14286 | -2.029 | 0.1087 | No | ||
| 200 | TNFRSF12A | 14325 | -2.054 | 0.1103 | No | ||
| 201 | IL22RA2 | 14331 | -2.055 | 0.1138 | No | ||
| 202 | TNFRSF10B | 14515 | -2.155 | 0.1077 | No | ||
| 203 | LIF | 14916 | -2.375 | 0.0902 | No | ||
| 204 | AMHR2 | 15083 | -2.477 | 0.0857 | No | ||
| 205 | IL13 | 15328 | -2.665 | 0.0772 | No | ||
| 206 | XCR1 | 15456 | -2.759 | 0.0753 | No | ||
| 207 | CD40 | 15735 | -3.002 | 0.0655 | No | ||
| 208 | TGFB2 | 15757 | -3.017 | 0.0698 | No | ||
| 209 | CCR1 | 16054 | -3.361 | 0.0598 | No | ||
| 210 | IL2RB | 16377 | -3.795 | 0.0491 | No | ||
| 211 | IL1RAP | 16410 | -3.854 | 0.0543 | No | ||
| 212 | IL12RB1 | 16657 | -4.264 | 0.0486 | No | ||
| 213 | IL2RA | 16704 | -4.369 | 0.0540 | No | ||
| 214 | VEGFC | 16745 | -4.447 | 0.0598 | No | ||
| 215 | VEGFB | 17022 | -4.989 | 0.0538 | No | ||
| 216 | IL10 | 17298 | -5.709 | 0.0491 | No | ||
| 217 | CCR7 | 18423 | -12.502 | 0.0105 | No |
