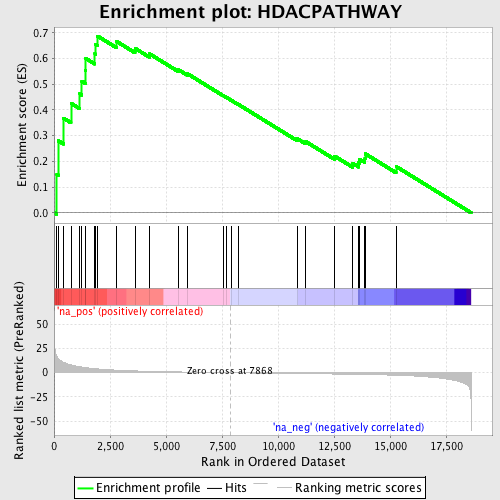
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | HDACPATHWAY |
| Enrichment Score (ES) | 0.6872712 |
| Normalized Enrichment Score (NES) | 1.8986318 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.02611738 |
| FWER p-Value | 0.119 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NFATC1 | 123 | 16.658 | 0.1498 | Yes | ||
| 2 | MEF2C | 185 | 14.237 | 0.2801 | Yes | ||
| 3 | CAMK1G | 424 | 10.715 | 0.3679 | Yes | ||
| 4 | MAPK14 | 754 | 8.054 | 0.4258 | Yes | ||
| 5 | MEF2A | 1147 | 6.154 | 0.4625 | Yes | ||
| 6 | IGF1 | 1238 | 5.779 | 0.5119 | Yes | ||
| 7 | CALM3 | 1400 | 5.225 | 0.5523 | Yes | ||
| 8 | CAMK1 | 1413 | 5.202 | 0.6005 | Yes | ||
| 9 | IGF1R | 1820 | 4.193 | 0.6180 | Yes | ||
| 10 | PPP3CC | 1849 | 4.130 | 0.6553 | Yes | ||
| 11 | PPP3CB | 1943 | 3.938 | 0.6873 | Yes | ||
| 12 | PPP3CA | 2779 | 2.647 | 0.6672 | No | ||
| 13 | CALM2 | 3612 | 1.847 | 0.6398 | No | ||
| 14 | PIK3R1 | 4260 | 1.432 | 0.6184 | No | ||
| 15 | HDAC5 | 5554 | 0.791 | 0.5563 | No | ||
| 16 | MEF2B | 5948 | 0.647 | 0.5412 | No | ||
| 17 | PIK3CA | 7547 | 0.100 | 0.4562 | No | ||
| 18 | SYT1 | 7677 | 0.056 | 0.4498 | No | ||
| 19 | MYOD1 | 7897 | -0.007 | 0.4381 | No | ||
| 20 | CABIN1 | 8241 | -0.116 | 0.4207 | No | ||
| 21 | MAP2K6 | 10855 | -0.863 | 0.2882 | No | ||
| 22 | NFATC2 | 11235 | -0.968 | 0.2769 | No | ||
| 23 | CALM1 | 12515 | -1.352 | 0.2208 | No | ||
| 24 | MAPK7 | 13332 | -1.628 | 0.1922 | No | ||
| 25 | MEF2D | 13598 | -1.728 | 0.1942 | No | ||
| 26 | AKT1 | 13635 | -1.741 | 0.2086 | No | ||
| 27 | INSR | 13875 | -1.845 | 0.2130 | No | ||
| 28 | YWHAH | 13894 | -1.851 | 0.2294 | No | ||
| 29 | AVP | 15266 | -2.614 | 0.1802 | No |
