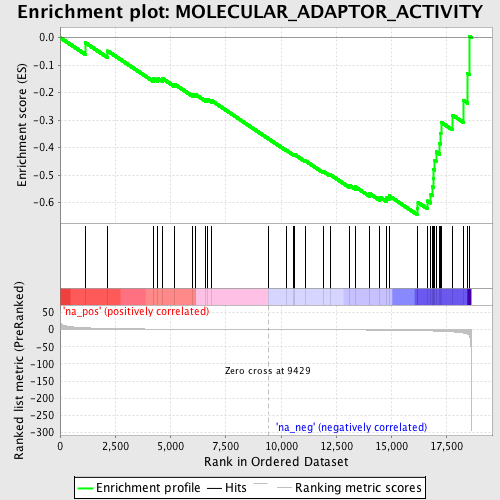
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_WTpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | MOLECULAR_ADAPTOR_ACTIVITY |
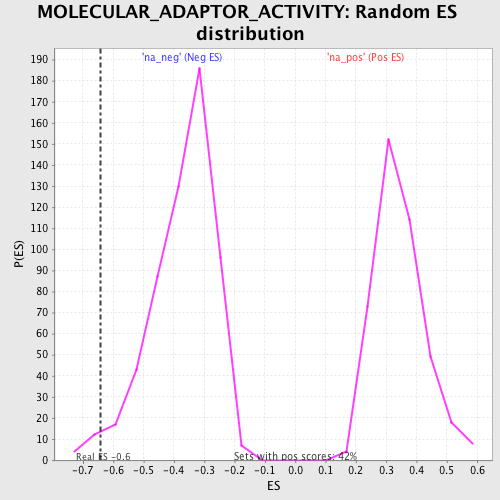
| Enrichment Score (ES) | -0.6431609 |
| Normalized Enrichment Score (NES) | -1.7339631 |
| Nominal p-value | 0.024054984 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SOCS2 | 1134 | 5.338 | -0.0184 | No | ||
| 2 | GRB7 | 2157 | 3.067 | -0.0489 | No | ||
| 3 | KHDRBS1 | 4221 | 1.463 | -0.1483 | No | ||
| 4 | ITSN2 | 4429 | 1.382 | -0.1484 | No | ||
| 5 | ANK1 | 4622 | 1.307 | -0.1483 | No | ||
| 6 | SRC | 5173 | 1.116 | -0.1690 | No | ||
| 7 | GRAP2 | 5998 | 0.867 | -0.2064 | No | ||
| 8 | SHB | 6132 | 0.830 | -0.2069 | No | ||
| 9 | SKAP2 | 6589 | 0.711 | -0.2258 | No | ||
| 10 | GRB10 | 6685 | 0.688 | -0.2254 | No | ||
| 11 | CNTNAP1 | 6862 | 0.644 | -0.2297 | No | ||
| 12 | LAT | 9409 | 0.005 | -0.3667 | No | ||
| 13 | SH3BGR | 10242 | -0.209 | -0.4098 | No | ||
| 14 | GRAP | 10568 | -0.278 | -0.4251 | No | ||
| 15 | CHN1 | 10608 | -0.287 | -0.4249 | No | ||
| 16 | CHN2 | 11088 | -0.404 | -0.4475 | No | ||
| 17 | SNX9 | 11906 | -0.620 | -0.4865 | No | ||
| 18 | BAIAP2 | 12215 | -0.706 | -0.4975 | No | ||
| 19 | ABI2 | 13082 | -0.969 | -0.5363 | No | ||
| 20 | IRS4 | 13356 | -1.072 | -0.5425 | No | ||
| 21 | GRB14 | 13992 | -1.318 | -0.5661 | No | ||
| 22 | RUSC1 | 14473 | -1.511 | -0.5799 | No | ||
| 23 | ARHGAP6 | 14779 | -1.669 | -0.5830 | No | ||
| 24 | SHC1 | 14889 | -1.736 | -0.5750 | No | ||
| 25 | ARHGAP1 | 16157 | -2.853 | -0.6204 | Yes | ||
| 26 | SH2D3C | 16197 | -2.906 | -0.5993 | Yes | ||
| 27 | GRB2 | 16640 | -3.636 | -0.5940 | Yes | ||
| 28 | TOB1 | 16781 | -3.893 | -0.5704 | Yes | ||
| 29 | SH3BP2 | 16850 | -4.017 | -0.5420 | Yes | ||
| 30 | STAM | 16889 | -4.107 | -0.5112 | Yes | ||
| 31 | ARHGAP4 | 16904 | -4.143 | -0.4789 | Yes | ||
| 32 | SH3BGRL | 16960 | -4.275 | -0.4477 | Yes | ||
| 33 | BLNK | 17018 | -4.436 | -0.4153 | Yes | ||
| 34 | EPS8 | 17183 | -4.893 | -0.3850 | Yes | ||
| 35 | CRKL | 17222 | -5.001 | -0.3471 | Yes | ||
| 36 | SH2D2A | 17241 | -5.040 | -0.3078 | Yes | ||
| 37 | LASP1 | 17764 | -6.772 | -0.2818 | Yes | ||
| 38 | GAB1 | 18256 | -10.039 | -0.2281 | Yes | ||
| 39 | SLA | 18451 | -13.415 | -0.1313 | Yes | ||
| 40 | VAV3 | 18521 | -17.543 | 0.0051 | Yes |