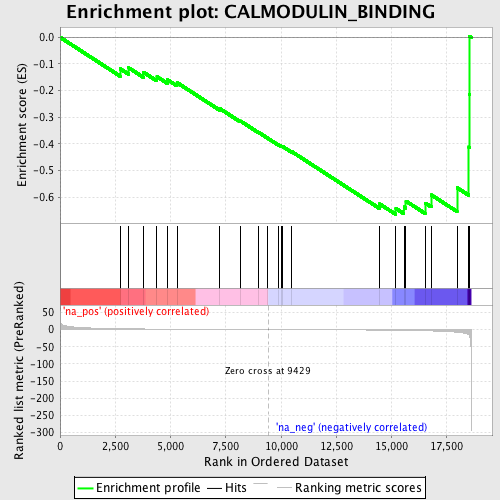
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_WTpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | CALMODULIN_BINDING |
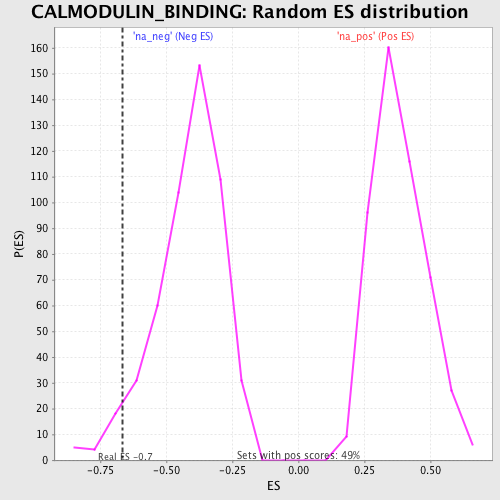
| Enrichment Score (ES) | -0.6653009 |
| Normalized Enrichment Score (NES) | -1.6041411 |
| Nominal p-value | 0.0407767 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TRPV4 | 2719 | 2.419 | -0.1183 | No | ||
| 2 | MYO9B | 3106 | 2.101 | -0.1147 | No | ||
| 3 | DAPK2 | 3773 | 1.669 | -0.1313 | No | ||
| 4 | TRPV6 | 4384 | 1.397 | -0.1479 | No | ||
| 5 | RGS16 | 4857 | 1.213 | -0.1593 | No | ||
| 6 | IQGAP1 | 5291 | 1.085 | -0.1700 | No | ||
| 7 | MYO7A | 7223 | 0.554 | -0.2675 | No | ||
| 8 | RIT2 | 8160 | 0.315 | -0.3142 | No | ||
| 9 | MYO3A | 8959 | 0.122 | -0.3557 | No | ||
| 10 | CAMKK2 | 9391 | 0.010 | -0.3788 | No | ||
| 11 | STRN4 | 9883 | -0.123 | -0.4038 | No | ||
| 12 | RGS1 | 10010 | -0.156 | -0.4087 | No | ||
| 13 | CALD1 | 10058 | -0.167 | -0.4093 | No | ||
| 14 | DAPK1 | 10479 | -0.256 | -0.4290 | No | ||
| 15 | SPHK1 | 14459 | -1.506 | -0.6256 | No | ||
| 16 | MYO6 | 15199 | -1.937 | -0.6429 | Yes | ||
| 17 | TTN | 15564 | -2.218 | -0.6368 | Yes | ||
| 18 | RGS4 | 15654 | -2.295 | -0.6151 | Yes | ||
| 19 | ATPIF1 | 16523 | -3.424 | -0.6222 | Yes | ||
| 20 | CACNA1C | 16801 | -3.921 | -0.5918 | Yes | ||
| 21 | RIT1 | 17968 | -7.749 | -0.5649 | Yes | ||
| 22 | NRGN | 18502 | -15.718 | -0.4119 | Yes | ||
| 23 | MARCKS | 18519 | -17.134 | -0.2147 | Yes | ||
| 24 | RGS2 | 18534 | -19.018 | 0.0044 | Yes |