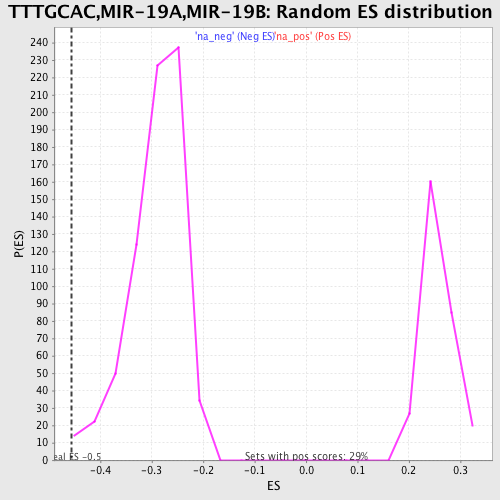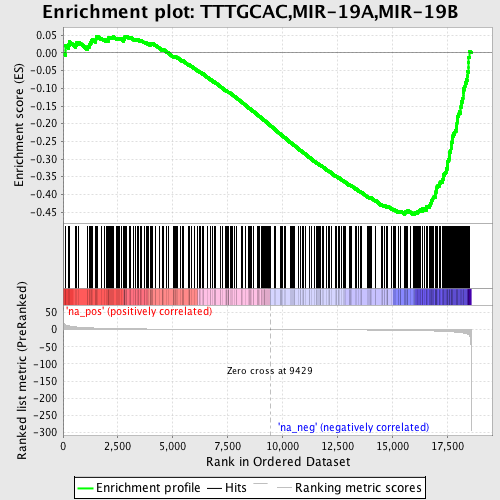
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_WTpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | TTTGCAC,MIR-19A,MIR-19B |
| Enrichment Score (ES) | -0.45695275 |
| Normalized Enrichment Score (NES) | -1.5755587 |
| Nominal p-value | 0.004237288 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PRC1 | 118 | 13.404 | 0.0070 | No | ||
| 2 | DOCK3 | 119 | 13.386 | 0.0204 | No | ||
| 3 | SFRS6 | 261 | 10.573 | 0.0232 | No | ||
| 4 | EPN2 | 287 | 10.284 | 0.0322 | No | ||
| 5 | CCRN4L | 576 | 7.789 | 0.0242 | No | ||
| 6 | AKAP1 | 625 | 7.430 | 0.0290 | No | ||
| 7 | PAPD4 | 719 | 6.927 | 0.0308 | No | ||
| 8 | NME7 | 1116 | 5.394 | 0.0145 | No | ||
| 9 | DNAJA2 | 1129 | 5.355 | 0.0192 | No | ||
| 10 | LRRK1 | 1185 | 5.159 | 0.0214 | No | ||
| 11 | ST3GAL5 | 1211 | 5.080 | 0.0251 | No | ||
| 12 | IVNS1ABP | 1236 | 5.009 | 0.0288 | No | ||
| 13 | CLOCK | 1275 | 4.902 | 0.0316 | No | ||
| 14 | CBX7 | 1292 | 4.862 | 0.0356 | No | ||
| 15 | ATP2C1 | 1334 | 4.735 | 0.0381 | No | ||
| 16 | PIK3R3 | 1472 | 4.347 | 0.0350 | No | ||
| 17 | CRSP7 | 1492 | 4.289 | 0.0382 | No | ||
| 18 | ROBO2 | 1499 | 4.283 | 0.0422 | No | ||
| 19 | GRSF1 | 1501 | 4.277 | 0.0464 | No | ||
| 20 | SYT11 | 1589 | 4.082 | 0.0457 | No | ||
| 21 | HNRPU | 1743 | 3.772 | 0.0411 | No | ||
| 22 | FOXF2 | 1867 | 3.516 | 0.0379 | No | ||
| 23 | RHEBL1 | 1956 | 3.372 | 0.0365 | No | ||
| 24 | BRWD1 | 2002 | 3.293 | 0.0373 | No | ||
| 25 | BZRAP1 | 2073 | 3.183 | 0.0367 | No | ||
| 26 | YTHDF2 | 2076 | 3.178 | 0.0397 | No | ||
| 27 | CCNL1 | 2078 | 3.176 | 0.0429 | No | ||
| 28 | NIPA1 | 2135 | 3.095 | 0.0429 | No | ||
| 29 | FASTK | 2173 | 3.034 | 0.0439 | No | ||
| 30 | RGL1 | 2210 | 2.986 | 0.0449 | No | ||
| 31 | NR4A2 | 2266 | 2.899 | 0.0448 | No | ||
| 32 | SOX6 | 2288 | 2.874 | 0.0465 | No | ||
| 33 | ITGA6 | 2431 | 2.689 | 0.0415 | No | ||
| 34 | TGOLN2 | 2475 | 2.648 | 0.0418 | No | ||
| 35 | DDX3X | 2518 | 2.596 | 0.0421 | No | ||
| 36 | L3MBTL3 | 2582 | 2.538 | 0.0412 | No | ||
| 37 | SFMBT1 | 2645 | 2.479 | 0.0402 | No | ||
| 38 | PHTF2 | 2772 | 2.373 | 0.0357 | No | ||
| 39 | RASSF1 | 2775 | 2.370 | 0.0380 | No | ||
| 40 | PLAA | 2788 | 2.362 | 0.0397 | No | ||
| 41 | TGM3 | 2799 | 2.344 | 0.0415 | No | ||
| 42 | OGT | 2811 | 2.337 | 0.0432 | No | ||
| 43 | TOR1B | 2815 | 2.333 | 0.0454 | No | ||
| 44 | DICER1 | 2854 | 2.304 | 0.0456 | No | ||
| 45 | PITPNM2 | 2883 | 2.277 | 0.0464 | No | ||
| 46 | KIF3A | 2900 | 2.264 | 0.0478 | No | ||
| 47 | ATG16L1 | 3025 | 2.150 | 0.0431 | No | ||
| 48 | B4GALT5 | 3056 | 2.134 | 0.0436 | No | ||
| 49 | FMR1 | 3089 | 2.112 | 0.0440 | No | ||
| 50 | DTNA | 3225 | 2.010 | 0.0386 | No | ||
| 51 | ZFYVE9 | 3299 | 1.949 | 0.0365 | No | ||
| 52 | PHF12 | 3318 | 1.933 | 0.0375 | No | ||
| 53 | TAF4 | 3376 | 1.891 | 0.0363 | No | ||
| 54 | MIB1 | 3416 | 1.866 | 0.0360 | No | ||
| 55 | RUNX3 | 3422 | 1.865 | 0.0376 | No | ||
| 56 | MACF1 | 3524 | 1.806 | 0.0339 | No | ||
| 57 | IGSF3 | 3531 | 1.802 | 0.0353 | No | ||
| 58 | CAMSAP1 | 3579 | 1.771 | 0.0345 | No | ||
| 59 | WNT1 | 3690 | 1.713 | 0.0302 | No | ||
| 60 | SLC26A7 | 3712 | 1.702 | 0.0308 | No | ||
| 61 | SLMAP | 3782 | 1.664 | 0.0287 | No | ||
| 62 | PCDHA10 | 3848 | 1.628 | 0.0268 | No | ||
| 63 | SLC9A2 | 3876 | 1.615 | 0.0269 | No | ||
| 64 | PITX1 | 3968 | 1.573 | 0.0235 | No | ||
| 65 | SPG20 | 3973 | 1.570 | 0.0248 | No | ||
| 66 | WEE1 | 3977 | 1.567 | 0.0262 | No | ||
| 67 | MINK1 | 4023 | 1.546 | 0.0253 | No | ||
| 68 | DLC1 | 4036 | 1.542 | 0.0262 | No | ||
| 69 | ITSN1 | 4067 | 1.526 | 0.0261 | No | ||
| 70 | ASXL2 | 4086 | 1.520 | 0.0266 | No | ||
| 71 | ATRX | 4191 | 1.476 | 0.0224 | No | ||
| 72 | ATP10A | 4231 | 1.460 | 0.0217 | No | ||
| 73 | IGF2R | 4392 | 1.395 | 0.0144 | No | ||
| 74 | ASNA1 | 4514 | 1.346 | 0.0091 | No | ||
| 75 | POU3F2 | 4523 | 1.343 | 0.0100 | No | ||
| 76 | KPNA3 | 4550 | 1.334 | 0.0099 | No | ||
| 77 | DDX3Y | 4586 | 1.321 | 0.0093 | No | ||
| 78 | BNC2 | 4711 | 1.271 | 0.0038 | No | ||
| 79 | SMOC1 | 4787 | 1.240 | 0.0009 | No | ||
| 80 | DPYSL5 | 5011 | 1.166 | -0.0101 | No | ||
| 81 | MAP3K12 | 5043 | 1.158 | -0.0106 | No | ||
| 82 | SPEN | 5044 | 1.158 | -0.0095 | No | ||
| 83 | CAB39 | 5096 | 1.142 | -0.0111 | No | ||
| 84 | CSNK1G1 | 5120 | 1.133 | -0.0113 | No | ||
| 85 | TNRC6A | 5134 | 1.130 | -0.0108 | No | ||
| 86 | GDA | 5162 | 1.120 | -0.0112 | No | ||
| 87 | RUNX2 | 5229 | 1.100 | -0.0137 | No | ||
| 88 | EXOC5 | 5339 | 1.070 | -0.0186 | No | ||
| 89 | PCDHA12 | 5439 | 1.031 | -0.0230 | No | ||
| 90 | SPIRE1 | 5446 | 1.029 | -0.0223 | No | ||
| 91 | SBF2 | 5470 | 1.022 | -0.0225 | No | ||
| 92 | WDFY3 | 5480 | 1.020 | -0.0220 | No | ||
| 93 | NRBF2 | 5717 | 0.949 | -0.0340 | No | ||
| 94 | POU4F1 | 5741 | 0.943 | -0.0343 | No | ||
| 95 | TNPO2 | 5742 | 0.942 | -0.0334 | No | ||
| 96 | TSC1 | 5773 | 0.933 | -0.0341 | No | ||
| 97 | PPARA | 5865 | 0.906 | -0.0381 | No | ||
| 98 | NPAS2 | 5966 | 0.878 | -0.0427 | No | ||
| 99 | CALM1 | 6121 | 0.834 | -0.0503 | No | ||
| 100 | PCDHA7 | 6124 | 0.832 | -0.0496 | No | ||
| 101 | RAP1A | 6204 | 0.813 | -0.0531 | No | ||
| 102 | OLFM1 | 6208 | 0.812 | -0.0525 | No | ||
| 103 | ARC | 6254 | 0.800 | -0.0541 | No | ||
| 104 | PRICKLE2 | 6354 | 0.775 | -0.0588 | No | ||
| 105 | EREG | 6381 | 0.769 | -0.0594 | No | ||
| 106 | RBMS1 | 6386 | 0.768 | -0.0589 | No | ||
| 107 | DCBLD2 | 6591 | 0.711 | -0.0693 | No | ||
| 108 | PCDH10 | 6714 | 0.682 | -0.0753 | No | ||
| 109 | ARHGAP5 | 6737 | 0.677 | -0.0759 | No | ||
| 110 | ARHGEF12 | 6825 | 0.653 | -0.0800 | No | ||
| 111 | PTPRG | 6878 | 0.639 | -0.0822 | No | ||
| 112 | ARHGAP21 | 6882 | 0.637 | -0.0817 | No | ||
| 113 | STK35 | 6946 | 0.621 | -0.0845 | No | ||
| 114 | LRCH2 | 6958 | 0.617 | -0.0845 | No | ||
| 115 | HNRPF | 7168 | 0.567 | -0.0954 | No | ||
| 116 | TRPS1 | 7169 | 0.567 | -0.0948 | No | ||
| 117 | LRP2 | 7283 | 0.541 | -0.1005 | No | ||
| 118 | RBMS3 | 7387 | 0.517 | -0.1056 | No | ||
| 119 | IGF1 | 7424 | 0.506 | -0.1071 | No | ||
| 120 | FNDC3A | 7427 | 0.504 | -0.1067 | No | ||
| 121 | TBK1 | 7451 | 0.497 | -0.1074 | No | ||
| 122 | TBC1D8 | 7479 | 0.489 | -0.1084 | No | ||
| 123 | CLTC | 7528 | 0.477 | -0.1106 | No | ||
| 124 | KCNJ2 | 7543 | 0.474 | -0.1109 | No | ||
| 125 | WDR26 | 7550 | 0.473 | -0.1107 | No | ||
| 126 | RXRA | 7557 | 0.471 | -0.1106 | No | ||
| 127 | PCDHA3 | 7611 | 0.456 | -0.1130 | No | ||
| 128 | KCNA4 | 7677 | 0.438 | -0.1162 | No | ||
| 129 | BACE1 | 7703 | 0.430 | -0.1171 | No | ||
| 130 | PLCB1 | 7808 | 0.401 | -0.1224 | No | ||
| 131 | PGM2L1 | 7904 | 0.380 | -0.1272 | No | ||
| 132 | ANXA7 | 7912 | 0.378 | -0.1272 | No | ||
| 133 | ADAM12 | 8127 | 0.326 | -0.1386 | No | ||
| 134 | SV2A | 8152 | 0.317 | -0.1396 | No | ||
| 135 | PLXNC1 | 8169 | 0.312 | -0.1402 | No | ||
| 136 | ITPR1 | 8317 | 0.274 | -0.1479 | No | ||
| 137 | IGFBP3 | 8459 | 0.244 | -0.1554 | No | ||
| 138 | CBLN2 | 8480 | 0.238 | -0.1563 | No | ||
| 139 | SEMA4C | 8516 | 0.229 | -0.1580 | No | ||
| 140 | DLX3 | 8552 | 0.219 | -0.1597 | No | ||
| 141 | ENPP5 | 8558 | 0.217 | -0.1597 | No | ||
| 142 | LRIG1 | 8568 | 0.215 | -0.1600 | No | ||
| 143 | GRK6 | 8677 | 0.189 | -0.1657 | No | ||
| 144 | SLC24A3 | 8699 | 0.183 | -0.1667 | No | ||
| 145 | KIT | 8853 | 0.149 | -0.1749 | No | ||
| 146 | ZFPM2 | 8855 | 0.148 | -0.1748 | No | ||
| 147 | USP33 | 8908 | 0.136 | -0.1775 | No | ||
| 148 | PDE5A | 8918 | 0.133 | -0.1779 | No | ||
| 149 | SHANK2 | 8938 | 0.129 | -0.1788 | No | ||
| 150 | LPP | 9024 | 0.105 | -0.1834 | No | ||
| 151 | ZBTB4 | 9059 | 0.096 | -0.1851 | No | ||
| 152 | HLF | 9105 | 0.085 | -0.1875 | No | ||
| 153 | RAPGEF4 | 9137 | 0.072 | -0.1891 | No | ||
| 154 | RIN2 | 9161 | 0.066 | -0.1903 | No | ||
| 155 | KPNA4 | 9176 | 0.063 | -0.1910 | No | ||
| 156 | RNF2 | 9179 | 0.063 | -0.1911 | No | ||
| 157 | F3 | 9221 | 0.052 | -0.1933 | No | ||
| 158 | TESK2 | 9279 | 0.037 | -0.1964 | No | ||
| 159 | SLC24A4 | 9327 | 0.026 | -0.1989 | No | ||
| 160 | SMOC2 | 9382 | 0.012 | -0.2018 | No | ||
| 161 | CSMD1 | 9419 | 0.002 | -0.2038 | No | ||
| 162 | SYT1 | 9441 | -0.004 | -0.2050 | No | ||
| 163 | NCALD | 9466 | -0.013 | -0.2063 | No | ||
| 164 | CTGF | 9648 | -0.059 | -0.2161 | No | ||
| 165 | ABCA1 | 9699 | -0.072 | -0.2188 | No | ||
| 166 | NRK | 9887 | -0.124 | -0.2289 | No | ||
| 167 | ARIH2 | 9898 | -0.128 | -0.2293 | No | ||
| 168 | GRM7 | 9951 | -0.140 | -0.2320 | No | ||
| 169 | BTBD7 | 9962 | -0.143 | -0.2324 | No | ||
| 170 | STAT5B | 9990 | -0.149 | -0.2338 | No | ||
| 171 | ZFYVE26 | 10108 | -0.179 | -0.2400 | No | ||
| 172 | CS | 10116 | -0.181 | -0.2402 | No | ||
| 173 | ATXN1 | 10372 | -0.234 | -0.2539 | No | ||
| 174 | MAPK6 | 10388 | -0.237 | -0.2545 | No | ||
| 175 | ARHGAP12 | 10444 | -0.249 | -0.2573 | No | ||
| 176 | SH3D19 | 10489 | -0.258 | -0.2594 | No | ||
| 177 | TMEPAI | 10534 | -0.268 | -0.2616 | No | ||
| 178 | ARFGEF1 | 10547 | -0.272 | -0.2619 | No | ||
| 179 | COL19A1 | 10737 | -0.317 | -0.2720 | No | ||
| 180 | EIF2C1 | 10796 | -0.332 | -0.2748 | No | ||
| 181 | SOCS3 | 10804 | -0.334 | -0.2749 | No | ||
| 182 | PNRC1 | 10889 | -0.355 | -0.2791 | No | ||
| 183 | MLLT6 | 10950 | -0.374 | -0.2820 | No | ||
| 184 | KLF10 | 10954 | -0.374 | -0.2818 | No | ||
| 185 | NEUROD1 | 11032 | -0.392 | -0.2856 | No | ||
| 186 | EPHB3 | 11210 | -0.438 | -0.2949 | No | ||
| 187 | ERBB4 | 11236 | -0.444 | -0.2958 | No | ||
| 188 | ATP11A | 11319 | -0.466 | -0.2998 | No | ||
| 189 | CGN | 11457 | -0.498 | -0.3068 | No | ||
| 190 | CCNT2 | 11544 | -0.518 | -0.3110 | No | ||
| 191 | EDG1 | 11553 | -0.521 | -0.3110 | No | ||
| 192 | RAI2 | 11573 | -0.527 | -0.3115 | No | ||
| 193 | BMPR2 | 11649 | -0.548 | -0.3150 | No | ||
| 194 | VPS4B | 11702 | -0.563 | -0.3173 | No | ||
| 195 | DHX40 | 11723 | -0.568 | -0.3178 | No | ||
| 196 | KCNS2 | 11733 | -0.571 | -0.3178 | No | ||
| 197 | ST8SIA3 | 11745 | -0.575 | -0.3178 | No | ||
| 198 | BAMBI | 11809 | -0.594 | -0.3206 | No | ||
| 199 | HIPK1 | 11861 | -0.608 | -0.3228 | No | ||
| 200 | RAB2B | 12026 | -0.654 | -0.3312 | No | ||
| 201 | STK38 | 12098 | -0.676 | -0.3344 | No | ||
| 202 | RTN1 | 12130 | -0.684 | -0.3354 | No | ||
| 203 | BCL3 | 12157 | -0.691 | -0.3361 | No | ||
| 204 | PFN1 | 12242 | -0.713 | -0.3400 | No | ||
| 205 | RFX1 | 12393 | -0.766 | -0.3474 | No | ||
| 206 | SLC35F1 | 12447 | -0.779 | -0.3496 | No | ||
| 207 | PCDHA5 | 12471 | -0.788 | -0.3500 | No | ||
| 208 | MID1IP1 | 12549 | -0.814 | -0.3534 | No | ||
| 209 | MAP3K14 | 12563 | -0.819 | -0.3533 | No | ||
| 210 | THRAP2 | 12576 | -0.823 | -0.3532 | No | ||
| 211 | EMX2 | 12668 | -0.850 | -0.3573 | No | ||
| 212 | PTP4A1 | 12768 | -0.880 | -0.3618 | No | ||
| 213 | PCDHA1 | 12822 | -0.893 | -0.3638 | No | ||
| 214 | TRIM33 | 12860 | -0.905 | -0.3650 | No | ||
| 215 | SOX21 | 13030 | -0.952 | -0.3733 | No | ||
| 216 | ELMOD2 | 13052 | -0.957 | -0.3735 | No | ||
| 217 | ADAMTS18 | 13078 | -0.967 | -0.3739 | No | ||
| 218 | RFX4 | 13110 | -0.981 | -0.3746 | No | ||
| 219 | DBN1 | 13160 | -1.002 | -0.3763 | No | ||
| 220 | WDR20 | 13335 | -1.064 | -0.3847 | No | ||
| 221 | DAG1 | 13353 | -1.070 | -0.3846 | No | ||
| 222 | GULP1 | 13483 | -1.119 | -0.3905 | No | ||
| 223 | PHF13 | 13551 | -1.149 | -0.3930 | No | ||
| 224 | ATP2B2 | 13603 | -1.174 | -0.3947 | No | ||
| 225 | PDE7B | 13607 | -1.175 | -0.3936 | No | ||
| 226 | NRP2 | 13872 | -1.274 | -0.4068 | No | ||
| 227 | RAB34 | 13913 | -1.290 | -0.4077 | No | ||
| 228 | PCDHA11 | 13971 | -1.311 | -0.4095 | No | ||
| 229 | SEPT7 | 14013 | -1.325 | -0.4105 | No | ||
| 230 | QKI | 14028 | -1.333 | -0.4099 | No | ||
| 231 | CNTFR | 14034 | -1.336 | -0.4088 | No | ||
| 232 | CYLD | 14228 | -1.408 | -0.4180 | No | ||
| 233 | ARRDC4 | 14237 | -1.413 | -0.4170 | No | ||
| 234 | SPRYD3 | 14493 | -1.517 | -0.4295 | No | ||
| 235 | MAPK14 | 14537 | -1.541 | -0.4303 | No | ||
| 236 | CCND2 | 14567 | -1.552 | -0.4303 | No | ||
| 237 | CBX1 | 14637 | -1.593 | -0.4325 | No | ||
| 238 | WNT3 | 14643 | -1.597 | -0.4312 | No | ||
| 239 | UCP3 | 14755 | -1.656 | -0.4356 | No | ||
| 240 | FLNC | 14771 | -1.664 | -0.4347 | No | ||
| 241 | TNIP1 | 14783 | -1.671 | -0.4337 | No | ||
| 242 | RAF1 | 14806 | -1.685 | -0.4332 | No | ||
| 243 | SULF1 | 14968 | -1.779 | -0.4402 | No | ||
| 244 | FZD8 | 15046 | -1.825 | -0.4426 | No | ||
| 245 | PCDHA6 | 15091 | -1.853 | -0.4432 | No | ||
| 246 | ELAVL1 | 15167 | -1.909 | -0.4454 | No | ||
| 247 | MECP2 | 15279 | -1.996 | -0.4494 | No | ||
| 248 | SNPH | 15303 | -2.014 | -0.4487 | No | ||
| 249 | IMPDH1 | 15354 | -2.060 | -0.4494 | No | ||
| 250 | CHST1 | 15387 | -2.079 | -0.4490 | No | ||
| 251 | CAST | 15399 | -2.088 | -0.4475 | No | ||
| 252 | GRIN2A | 15566 | -2.220 | -0.4544 | Yes | ||
| 253 | ESR1 | 15580 | -2.232 | -0.4529 | Yes | ||
| 254 | RNF111 | 15586 | -2.236 | -0.4509 | Yes | ||
| 255 | ZNF3 | 15601 | -2.250 | -0.4494 | Yes | ||
| 256 | LIF | 15662 | -2.303 | -0.4504 | Yes | ||
| 257 | PHLDA3 | 15675 | -2.316 | -0.4488 | Yes | ||
| 258 | MEF2D | 15700 | -2.341 | -0.4477 | Yes | ||
| 259 | INHBB | 15712 | -2.351 | -0.4460 | Yes | ||
| 260 | ZBTB10 | 15828 | -2.459 | -0.4498 | Yes | ||
| 261 | SLC31A2 | 15955 | -2.600 | -0.4541 | Yes | ||
| 262 | ELL2 | 16008 | -2.674 | -0.4543 | Yes | ||
| 263 | ARMC8 | 16028 | -2.693 | -0.4526 | Yes | ||
| 264 | BSN | 16054 | -2.726 | -0.4513 | Yes | ||
| 265 | LNPEP | 16107 | -2.785 | -0.4513 | Yes | ||
| 266 | ARHGAP1 | 16157 | -2.853 | -0.4511 | Yes | ||
| 267 | RAB18 | 16166 | -2.866 | -0.4487 | Yes | ||
| 268 | SLC6A8 | 16193 | -2.900 | -0.4472 | Yes | ||
| 269 | ARPP-19 | 16229 | -2.939 | -0.4462 | Yes | ||
| 270 | SNIP1 | 16276 | -2.998 | -0.4457 | Yes | ||
| 271 | SMAD5 | 16280 | -3.000 | -0.4429 | Yes | ||
| 272 | PFKFB3 | 16375 | -3.169 | -0.4449 | Yes | ||
| 273 | ADCY9 | 16377 | -3.172 | -0.4417 | Yes | ||
| 274 | AP1G1 | 16381 | -3.175 | -0.4387 | Yes | ||
| 275 | CNOT7 | 16488 | -3.355 | -0.4412 | Yes | ||
| 276 | TMEM32 | 16554 | -3.480 | -0.4412 | Yes | ||
| 277 | BLCAP | 16562 | -3.493 | -0.4381 | Yes | ||
| 278 | ABR | 16566 | -3.497 | -0.4348 | Yes | ||
| 279 | DDX6 | 16619 | -3.595 | -0.4340 | Yes | ||
| 280 | CBFB | 16690 | -3.725 | -0.4341 | Yes | ||
| 281 | SLC9A6 | 16710 | -3.749 | -0.4314 | Yes | ||
| 282 | ARFIP1 | 16728 | -3.781 | -0.4286 | Yes | ||
| 283 | NDFIP2 | 16745 | -3.798 | -0.4257 | Yes | ||
| 284 | DNAJB1 | 16768 | -3.862 | -0.4230 | Yes | ||
| 285 | SMARCD2 | 16797 | -3.912 | -0.4206 | Yes | ||
| 286 | CACNA1C | 16801 | -3.921 | -0.4168 | Yes | ||
| 287 | FBXO8 | 16836 | -3.994 | -0.4147 | Yes | ||
| 288 | ZMYND11 | 16857 | -4.033 | -0.4118 | Yes | ||
| 289 | FEM1C | 16862 | -4.045 | -0.4079 | Yes | ||
| 290 | RAB8B | 16903 | -4.139 | -0.4060 | Yes | ||
| 291 | RAB33B | 16970 | -4.305 | -0.4053 | Yes | ||
| 292 | TNFRSF12A | 16971 | -4.305 | -0.4010 | Yes | ||
| 293 | EDARADD | 16974 | -4.309 | -0.3968 | Yes | ||
| 294 | RHOB | 16989 | -4.350 | -0.3932 | Yes | ||
| 295 | SPHK2 | 17001 | -4.388 | -0.3894 | Yes | ||
| 296 | FOXP1 | 17003 | -4.392 | -0.3850 | Yes | ||
| 297 | ABCB7 | 17029 | -4.480 | -0.3819 | Yes | ||
| 298 | EFNB2 | 17039 | -4.519 | -0.3779 | Yes | ||
| 299 | ELOVL5 | 17082 | -4.629 | -0.3756 | Yes | ||
| 300 | ACBD5 | 17138 | -4.773 | -0.3738 | Yes | ||
| 301 | EPS15 | 17161 | -4.849 | -0.3701 | Yes | ||
| 302 | SLC9A1 | 17175 | -4.881 | -0.3660 | Yes | ||
| 303 | SPATA2 | 17218 | -4.994 | -0.3633 | Yes | ||
| 304 | MBNL1 | 17291 | -5.195 | -0.3620 | Yes | ||
| 305 | TRAK2 | 17304 | -5.247 | -0.3574 | Yes | ||
| 306 | GPR137B | 17337 | -5.352 | -0.3538 | Yes | ||
| 307 | ZDHHC18 | 17344 | -5.367 | -0.3488 | Yes | ||
| 308 | ARID4B | 17348 | -5.382 | -0.3435 | Yes | ||
| 309 | ADCY7 | 17389 | -5.510 | -0.3402 | Yes | ||
| 310 | PKNOX1 | 17429 | -5.589 | -0.3368 | Yes | ||
| 311 | ADIPOR2 | 17462 | -5.761 | -0.3327 | Yes | ||
| 312 | WDR44 | 17477 | -5.807 | -0.3277 | Yes | ||
| 313 | RAB21 | 17501 | -5.914 | -0.3230 | Yes | ||
| 314 | WDR1 | 17521 | -5.990 | -0.3181 | Yes | ||
| 315 | ARL6IP2 | 17528 | -6.024 | -0.3124 | Yes | ||
| 316 | CNOT4 | 17532 | -6.032 | -0.3065 | Yes | ||
| 317 | UBE2A | 17557 | -6.086 | -0.3017 | Yes | ||
| 318 | EPC2 | 17589 | -6.197 | -0.2972 | Yes | ||
| 319 | CD69 | 17611 | -6.271 | -0.2921 | Yes | ||
| 320 | RAP1B | 17617 | -6.283 | -0.2861 | Yes | ||
| 321 | RAP2C | 17622 | -6.298 | -0.2800 | Yes | ||
| 322 | ST8SIA4 | 17665 | -6.412 | -0.2759 | Yes | ||
| 323 | SNX17 | 17679 | -6.437 | -0.2701 | Yes | ||
| 324 | RNF11 | 17690 | -6.488 | -0.2642 | Yes | ||
| 325 | PPP2R5E | 17693 | -6.501 | -0.2578 | Yes | ||
| 326 | TTYH3 | 17705 | -6.553 | -0.2518 | Yes | ||
| 327 | ACSL4 | 17740 | -6.705 | -0.2470 | Yes | ||
| 328 | TGFBR2 | 17742 | -6.708 | -0.2403 | Yes | ||
| 329 | PSAP | 17765 | -6.774 | -0.2347 | Yes | ||
| 330 | ATP6V1B2 | 17782 | -6.835 | -0.2288 | Yes | ||
| 331 | KLF13 | 17831 | -7.092 | -0.2243 | Yes | ||
| 332 | UBE2D2 | 17862 | -7.224 | -0.2187 | Yes | ||
| 333 | LBH | 17911 | -7.514 | -0.2138 | Yes | ||
| 334 | FBXO28 | 17925 | -7.598 | -0.2069 | Yes | ||
| 335 | RNF38 | 17948 | -7.675 | -0.2004 | Yes | ||
| 336 | SDC1 | 17954 | -7.690 | -0.1930 | Yes | ||
| 337 | SGK | 17966 | -7.716 | -0.1859 | Yes | ||
| 338 | ETV1 | 17982 | -7.808 | -0.1789 | Yes | ||
| 339 | MBNL2 | 18014 | -7.993 | -0.1726 | Yes | ||
| 340 | MYLIP | 18045 | -8.182 | -0.1660 | Yes | ||
| 341 | RAB5B | 18127 | -8.884 | -0.1616 | Yes | ||
| 342 | ARRDC3 | 18128 | -8.916 | -0.1526 | Yes | ||
| 343 | SOCS5 | 18141 | -9.002 | -0.1443 | Yes | ||
| 344 | SOCS1 | 18162 | -9.161 | -0.1362 | Yes | ||
| 345 | CCM2 | 18214 | -9.613 | -0.1294 | Yes | ||
| 346 | WBP2 | 18228 | -9.816 | -0.1203 | Yes | ||
| 347 | UBL3 | 18231 | -9.848 | -0.1105 | Yes | ||
| 348 | ELK3 | 18247 | -9.988 | -0.1013 | Yes | ||
| 349 | SOX4 | 18311 | -10.671 | -0.0941 | Yes | ||
| 350 | RALGPS1 | 18322 | -10.854 | -0.0838 | Yes | ||
| 351 | BTG1 | 18371 | -11.636 | -0.0748 | Yes | ||
| 352 | ZDHHC7 | 18426 | -12.800 | -0.0649 | Yes | ||
| 353 | VGLL4 | 18437 | -12.997 | -0.0524 | Yes | ||
| 354 | GJA1 | 18473 | -14.071 | -0.0403 | Yes | ||
| 355 | PTK2B | 18476 | -14.219 | -0.0261 | Yes | ||
| 356 | HBP1 | 18488 | -15.154 | -0.0116 | Yes | ||
| 357 | SOX5 | 18531 | -18.503 | 0.0047 | Yes |