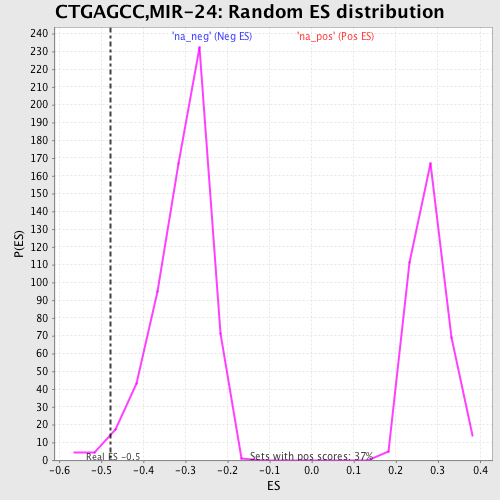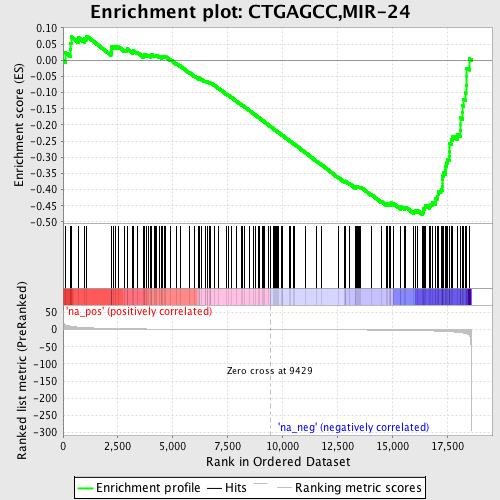
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_WTpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | CTGAGCC,MIR-24 |
| Enrichment Score (ES) | -0.47694772 |
| Normalized Enrichment Score (NES) | -1.5539281 |
| Nominal p-value | 0.017350158 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | DOCK3 | 119 | 13.386 | 0.0231 | No | ||
| 2 | XPO4 | 335 | 9.750 | 0.0330 | No | ||
| 3 | CCT3 | 350 | 9.521 | 0.0532 | No | ||
| 4 | AARS | 377 | 9.333 | 0.0724 | No | ||
| 5 | NEK4 | 710 | 6.962 | 0.0698 | No | ||
| 6 | PRKRIP1 | 977 | 5.887 | 0.0684 | No | ||
| 7 | CDV3 | 1084 | 5.496 | 0.0748 | No | ||
| 8 | SLC19A2 | 2184 | 3.018 | 0.0219 | No | ||
| 9 | RHBDL1 | 2189 | 3.009 | 0.0284 | No | ||
| 10 | ING5 | 2192 | 3.005 | 0.0349 | No | ||
| 11 | DHX30 | 2195 | 3.003 | 0.0414 | No | ||
| 12 | MAFG | 2285 | 2.877 | 0.0429 | No | ||
| 13 | MATR3 | 2407 | 2.715 | 0.0424 | No | ||
| 14 | PPM1G | 2533 | 2.586 | 0.0413 | No | ||
| 15 | OGT | 2811 | 2.337 | 0.0315 | No | ||
| 16 | INSIG1 | 2911 | 2.253 | 0.0311 | No | ||
| 17 | HAP1 | 2919 | 2.245 | 0.0357 | No | ||
| 18 | POGZ | 3184 | 2.043 | 0.0259 | No | ||
| 19 | TAF5 | 3186 | 2.040 | 0.0303 | No | ||
| 20 | CMTM4 | 3386 | 1.885 | 0.0237 | No | ||
| 21 | PTGER4 | 3681 | 1.719 | 0.0116 | No | ||
| 22 | RNF138 | 3706 | 1.705 | 0.0140 | No | ||
| 23 | NDST1 | 3710 | 1.703 | 0.0176 | No | ||
| 24 | PER2 | 3798 | 1.654 | 0.0166 | No | ||
| 25 | PIM2 | 3907 | 1.598 | 0.0142 | No | ||
| 26 | PTPRD | 3987 | 1.564 | 0.0134 | No | ||
| 27 | DLC1 | 4036 | 1.542 | 0.0142 | No | ||
| 28 | SEMA4A | 4037 | 1.541 | 0.0176 | No | ||
| 29 | RGL2 | 4150 | 1.498 | 0.0149 | No | ||
| 30 | MAPK7 | 4205 | 1.468 | 0.0152 | No | ||
| 31 | STC2 | 4247 | 1.452 | 0.0162 | No | ||
| 32 | SP6 | 4373 | 1.401 | 0.0125 | No | ||
| 33 | FGFR3 | 4482 | 1.356 | 0.0096 | No | ||
| 34 | SEMA6B | 4504 | 1.350 | 0.0115 | No | ||
| 35 | KPNA3 | 4550 | 1.334 | 0.0120 | No | ||
| 36 | JUB | 4620 | 1.308 | 0.0111 | No | ||
| 37 | PHC1 | 4686 | 1.280 | 0.0104 | No | ||
| 38 | INPP5B | 4876 | 1.205 | 0.0029 | No | ||
| 39 | TCF2 | 5155 | 1.122 | -0.0097 | No | ||
| 40 | EXOC5 | 5339 | 1.070 | -0.0173 | No | ||
| 41 | PRSS8 | 5767 | 0.936 | -0.0383 | No | ||
| 42 | PIGS | 6005 | 0.866 | -0.0493 | No | ||
| 43 | BHLHB5 | 6176 | 0.820 | -0.0567 | No | ||
| 44 | FST | 6202 | 0.813 | -0.0562 | No | ||
| 45 | RAP1A | 6204 | 0.813 | -0.0545 | No | ||
| 46 | HIC2 | 6327 | 0.781 | -0.0594 | No | ||
| 47 | NELL2 | 6467 | 0.745 | -0.0653 | No | ||
| 48 | KIF21B | 6478 | 0.743 | -0.0642 | No | ||
| 49 | CLCN6 | 6558 | 0.721 | -0.0668 | No | ||
| 50 | DCBLD2 | 6591 | 0.711 | -0.0670 | No | ||
| 51 | DNAJB12 | 6663 | 0.695 | -0.0693 | No | ||
| 52 | PCDH10 | 6714 | 0.682 | -0.0705 | No | ||
| 53 | CALCR | 6734 | 0.678 | -0.0701 | No | ||
| 54 | TOP1 | 6905 | 0.631 | -0.0779 | No | ||
| 55 | FBXL19 | 7082 | 0.590 | -0.0861 | No | ||
| 56 | HIP1R | 7452 | 0.497 | -0.1050 | No | ||
| 57 | YBX2 | 7539 | 0.475 | -0.1086 | No | ||
| 58 | COL11A2 | 7657 | 0.444 | -0.1140 | No | ||
| 59 | LRFN2 | 7914 | 0.377 | -0.1270 | No | ||
| 60 | APBA1 | 8109 | 0.331 | -0.1368 | No | ||
| 61 | PAK4 | 8161 | 0.314 | -0.1389 | No | ||
| 62 | AMMECR1 | 8279 | 0.282 | -0.1446 | No | ||
| 63 | ARID5B | 8479 | 0.238 | -0.1548 | No | ||
| 64 | SFXN5 | 8685 | 0.187 | -0.1655 | No | ||
| 65 | KLHL1 | 8766 | 0.169 | -0.1695 | No | ||
| 66 | PER1 | 8898 | 0.138 | -0.1763 | No | ||
| 67 | RASGRP4 | 8966 | 0.120 | -0.1796 | No | ||
| 68 | SUV420H2 | 9069 | 0.094 | -0.1849 | No | ||
| 69 | SMCR8 | 9151 | 0.068 | -0.1892 | No | ||
| 70 | RNF2 | 9179 | 0.063 | -0.1905 | No | ||
| 71 | MNT | 9368 | 0.015 | -0.2007 | No | ||
| 72 | ADAM19 | 9440 | -0.004 | -0.2045 | No | ||
| 73 | RARG | 9458 | -0.011 | -0.2054 | No | ||
| 74 | BCL2L11 | 9608 | -0.049 | -0.2134 | No | ||
| 75 | MMP16 | 9651 | -0.060 | -0.2155 | No | ||
| 76 | EPHA8 | 9700 | -0.072 | -0.2179 | No | ||
| 77 | NEK9 | 9728 | -0.079 | -0.2192 | No | ||
| 78 | PLOD2 | 9790 | -0.096 | -0.2223 | No | ||
| 79 | PDGFRA | 9795 | -0.098 | -0.2223 | No | ||
| 80 | MARK4 | 9930 | -0.136 | -0.2293 | No | ||
| 81 | GPX3 | 9987 | -0.149 | -0.2320 | No | ||
| 82 | ARHGEF5 | 10013 | -0.156 | -0.2330 | No | ||
| 83 | PTPRF | 10334 | -0.228 | -0.2498 | No | ||
| 84 | CACNB1 | 10345 | -0.230 | -0.2499 | No | ||
| 85 | IGFBP5 | 10518 | -0.264 | -0.2586 | No | ||
| 86 | PURA | 10543 | -0.271 | -0.2593 | No | ||
| 87 | NEUROD1 | 11032 | -0.392 | -0.2849 | No | ||
| 88 | EDG1 | 11553 | -0.521 | -0.3119 | No | ||
| 89 | TRPM6 | 11789 | -0.588 | -0.3233 | No | ||
| 90 | SP1 | 12546 | -0.813 | -0.3625 | No | ||
| 91 | RASA1 | 12828 | -0.895 | -0.3757 | No | ||
| 92 | CDX2 | 12847 | -0.902 | -0.3747 | No | ||
| 93 | TRIM33 | 12860 | -0.905 | -0.3734 | No | ||
| 94 | KCNK2 | 13053 | -0.957 | -0.3816 | No | ||
| 95 | EDA | 13314 | -1.056 | -0.3934 | No | ||
| 96 | ZXDA | 13315 | -1.057 | -0.3911 | No | ||
| 97 | SCML2 | 13360 | -1.073 | -0.3911 | No | ||
| 98 | CLK2 | 13404 | -1.090 | -0.3910 | No | ||
| 99 | SYP | 13467 | -1.115 | -0.3919 | No | ||
| 100 | GPC4 | 13517 | -1.135 | -0.3921 | No | ||
| 101 | APOA5 | 13574 | -1.159 | -0.3925 | No | ||
| 102 | CNTFR | 14034 | -1.336 | -0.4144 | No | ||
| 103 | WDTC1 | 14513 | -1.528 | -0.4370 | No | ||
| 104 | RAB5C | 14738 | -1.646 | -0.4455 | No | ||
| 105 | DUSP16 | 14775 | -1.668 | -0.4437 | No | ||
| 106 | BCL2L2 | 14888 | -1.736 | -0.4460 | No | ||
| 107 | AMOTL2 | 14935 | -1.759 | -0.4446 | No | ||
| 108 | RNF150 | 14943 | -1.762 | -0.4411 | No | ||
| 109 | RANBP10 | 15069 | -1.841 | -0.4438 | No | ||
| 110 | PTPN9 | 15391 | -2.084 | -0.4566 | No | ||
| 111 | RASSF2 | 15397 | -2.087 | -0.4522 | No | ||
| 112 | ABCB9 | 15553 | -2.206 | -0.4558 | No | ||
| 113 | GBA2 | 15614 | -2.266 | -0.4540 | No | ||
| 114 | MXI1 | 15985 | -2.644 | -0.4682 | No | ||
| 115 | BSN | 16054 | -2.726 | -0.4659 | No | ||
| 116 | ACVR1B | 16163 | -2.860 | -0.4654 | No | ||
| 117 | ADCY9 | 16377 | -3.172 | -0.4699 | Yes | ||
| 118 | MEN1 | 16423 | -3.244 | -0.4652 | Yes | ||
| 119 | GRIA3 | 16434 | -3.270 | -0.4585 | Yes | ||
| 120 | MMP14 | 16492 | -3.360 | -0.4542 | Yes | ||
| 121 | LYPLA2 | 16528 | -3.432 | -0.4485 | Yes | ||
| 122 | PIP5K1B | 16696 | -3.734 | -0.4493 | Yes | ||
| 123 | MOCS1 | 16765 | -3.851 | -0.4445 | Yes | ||
| 124 | DLGAP4 | 16829 | -3.977 | -0.4391 | Yes | ||
| 125 | FURIN | 16978 | -4.325 | -0.4376 | Yes | ||
| 126 | PIM1 | 16981 | -4.328 | -0.4282 | Yes | ||
| 127 | MAGI1 | 17043 | -4.526 | -0.4215 | Yes | ||
| 128 | WDR23 | 17086 | -4.638 | -0.4135 | Yes | ||
| 129 | PARP6 | 17127 | -4.749 | -0.4052 | Yes | ||
| 130 | TMEM9 | 17224 | -5.004 | -0.3994 | Yes | ||
| 131 | MAP3K7IP2 | 17275 | -5.161 | -0.3907 | Yes | ||
| 132 | LENG4 | 17285 | -5.183 | -0.3797 | Yes | ||
| 133 | TRPC4AP | 17286 | -5.185 | -0.3683 | Yes | ||
| 134 | RAB6IP1 | 17305 | -5.247 | -0.3577 | Yes | ||
| 135 | B3GNT6 | 17343 | -5.365 | -0.3478 | Yes | ||
| 136 | PLK3 | 17436 | -5.629 | -0.3404 | Yes | ||
| 137 | ATP6V0A2 | 17444 | -5.685 | -0.3282 | Yes | ||
| 138 | SESN1 | 17480 | -5.817 | -0.3173 | Yes | ||
| 139 | SLC12A6 | 17531 | -6.030 | -0.3067 | Yes | ||
| 140 | TOB2 | 17612 | -6.271 | -0.2971 | Yes | ||
| 141 | RAP1B | 17617 | -6.283 | -0.2835 | Yes | ||
| 142 | RAP2C | 17622 | -6.298 | -0.2698 | Yes | ||
| 143 | CDKN1B | 17623 | -6.299 | -0.2559 | Yes | ||
| 144 | RNF11 | 17690 | -6.488 | -0.2451 | Yes | ||
| 145 | PSAP | 17765 | -6.774 | -0.2342 | Yes | ||
| 146 | NEDD9 | 17973 | -7.778 | -0.2282 | Yes | ||
| 147 | SSH2 | 18101 | -8.657 | -0.2160 | Yes | ||
| 148 | VAV1 | 18125 | -8.843 | -0.1977 | Yes | ||
| 149 | RAB5B | 18127 | -8.884 | -0.1782 | Yes | ||
| 150 | CSK | 18203 | -9.505 | -0.1612 | Yes | ||
| 151 | H2AFX | 18211 | -9.573 | -0.1405 | Yes | ||
| 152 | TNFRSF19 | 18239 | -9.899 | -0.1201 | Yes | ||
| 153 | MIDN | 18342 | -11.212 | -0.1009 | Yes | ||
| 154 | PLCL2 | 18363 | -11.488 | -0.0766 | Yes | ||
| 155 | ALS2CR2 | 18398 | -12.251 | -0.0514 | Yes | ||
| 156 | SBK1 | 18401 | -12.359 | -0.0242 | Yes | ||
| 157 | ACBD4 | 18511 | -16.219 | 0.0057 | Yes |