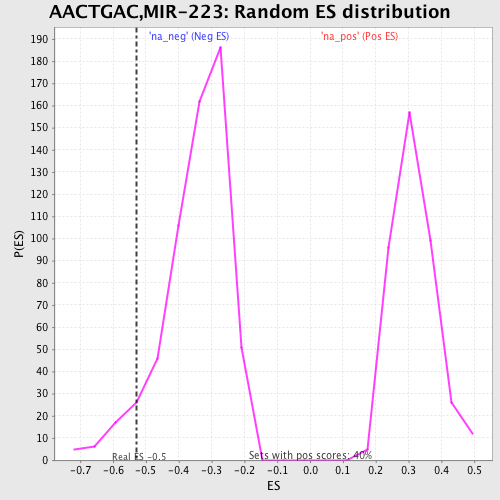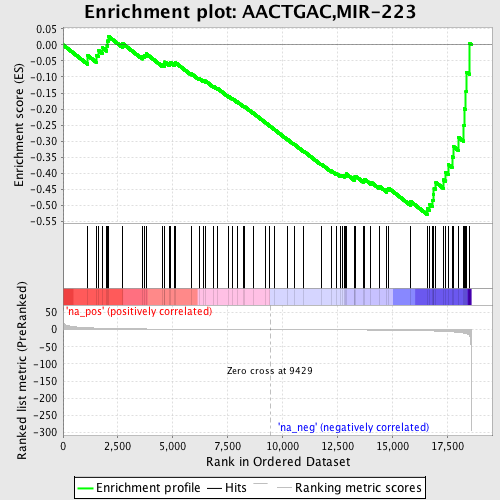
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_WTpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | AACTGAC,MIR-223 |
| Enrichment Score (ES) | -0.52893275 |
| Normalized Enrichment Score (NES) | -1.519657 |
| Nominal p-value | 0.0661157 |
| FDR q-value | 1.0 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RASSF4 | 1123 | 5.388 | -0.0332 | No | ||
| 2 | APRIN | 1509 | 4.258 | -0.0324 | No | ||
| 3 | PLEKHH1 | 1606 | 4.048 | -0.0170 | No | ||
| 4 | RPS6KB1 | 1785 | 3.698 | -0.0079 | No | ||
| 5 | UBTF | 1999 | 3.301 | -0.0026 | No | ||
| 6 | FBXW7 | 2029 | 3.259 | 0.0124 | No | ||
| 7 | ANKRD17 | 2053 | 3.218 | 0.0274 | No | ||
| 8 | SLC8A1 | 2711 | 2.428 | 0.0043 | No | ||
| 9 | PLAGL2 | 3609 | 1.755 | -0.0351 | No | ||
| 10 | SLC26A7 | 3712 | 1.702 | -0.0320 | No | ||
| 11 | KIF4A | 3787 | 1.661 | -0.0276 | No | ||
| 12 | KPNA3 | 4550 | 1.334 | -0.0619 | No | ||
| 13 | IL6ST | 4634 | 1.303 | -0.0598 | No | ||
| 14 | STMN1 | 4636 | 1.302 | -0.0532 | No | ||
| 15 | RNF4 | 4849 | 1.214 | -0.0585 | No | ||
| 16 | INPP5B | 4876 | 1.205 | -0.0538 | No | ||
| 17 | IGF1R | 5072 | 1.151 | -0.0584 | No | ||
| 18 | CSNK1G1 | 5120 | 1.133 | -0.0552 | No | ||
| 19 | ZCCHC14 | 5839 | 0.913 | -0.0893 | No | ||
| 20 | OLFM1 | 6208 | 0.812 | -0.1050 | No | ||
| 21 | PCTK2 | 6405 | 0.762 | -0.1117 | No | ||
| 22 | SLC23A2 | 6487 | 0.741 | -0.1124 | No | ||
| 23 | ATP7A | 6874 | 0.641 | -0.1299 | No | ||
| 24 | SON | 7053 | 0.595 | -0.1365 | No | ||
| 25 | WDR26 | 7550 | 0.473 | -0.1608 | No | ||
| 26 | STK39 | 7699 | 0.431 | -0.1666 | No | ||
| 27 | EFNA1 | 7934 | 0.374 | -0.1774 | No | ||
| 28 | ACVR2A | 8226 | 0.296 | -0.1916 | No | ||
| 29 | TSPAN7 | 8273 | 0.284 | -0.1926 | No | ||
| 30 | CALML4 | 8654 | 0.195 | -0.2121 | No | ||
| 31 | F3 | 9221 | 0.052 | -0.2423 | No | ||
| 32 | NFIB | 9405 | 0.006 | -0.2522 | No | ||
| 33 | MMP16 | 9651 | -0.060 | -0.2651 | No | ||
| 34 | NFIA | 10240 | -0.209 | -0.2957 | No | ||
| 35 | SOX11 | 10544 | -0.271 | -0.3107 | No | ||
| 36 | ANKRD16 | 10970 | -0.378 | -0.3317 | No | ||
| 37 | RBM16 | 11765 | -0.580 | -0.3716 | No | ||
| 38 | SCN3A | 12230 | -0.710 | -0.3930 | No | ||
| 39 | SLC35F1 | 12447 | -0.779 | -0.4007 | No | ||
| 40 | SEPT6 | 12651 | -0.845 | -0.4073 | No | ||
| 41 | SMARCD1 | 12724 | -0.866 | -0.4068 | No | ||
| 42 | RASA1 | 12828 | -0.895 | -0.4078 | No | ||
| 43 | IL28RA | 12882 | -0.913 | -0.4061 | No | ||
| 44 | SP3 | 12898 | -0.916 | -0.4022 | No | ||
| 45 | PRDM1 | 13269 | -1.039 | -0.4169 | No | ||
| 46 | SPRED1 | 13275 | -1.041 | -0.4119 | No | ||
| 47 | PTBP2 | 13320 | -1.059 | -0.4089 | No | ||
| 48 | RCN2 | 13682 | -1.203 | -0.4223 | No | ||
| 49 | VNN1 | 13726 | -1.223 | -0.4184 | No | ||
| 50 | QKI | 14028 | -1.333 | -0.4278 | No | ||
| 51 | CRIM1 | 14398 | -1.477 | -0.4403 | No | ||
| 52 | ALCAM | 14756 | -1.657 | -0.4511 | No | ||
| 53 | PDE4D | 14829 | -1.703 | -0.4463 | No | ||
| 54 | ZBTB10 | 15828 | -2.459 | -0.4877 | No | ||
| 55 | SLC39A1 | 16594 | -3.550 | -0.5109 | Yes | ||
| 56 | CBFB | 16690 | -3.725 | -0.4972 | Yes | ||
| 57 | FBXO8 | 16836 | -3.994 | -0.4847 | Yes | ||
| 58 | PURB | 16872 | -4.070 | -0.4660 | Yes | ||
| 59 | RAB8B | 16903 | -4.139 | -0.4466 | Yes | ||
| 60 | RHOB | 16989 | -4.350 | -0.4291 | Yes | ||
| 61 | MTPN | 17346 | -5.377 | -0.4210 | Yes | ||
| 62 | PKNOX1 | 17429 | -5.589 | -0.3971 | Yes | ||
| 63 | UBE2A | 17557 | -6.086 | -0.3731 | Yes | ||
| 64 | HHEX | 17732 | -6.662 | -0.3487 | Yes | ||
| 65 | STIM1 | 17786 | -6.867 | -0.3167 | Yes | ||
| 66 | PDPK1 | 18025 | -8.051 | -0.2887 | Yes | ||
| 67 | ATP1B1 | 18264 | -10.083 | -0.2504 | Yes | ||
| 68 | RNF34 | 18290 | -10.412 | -0.1989 | Yes | ||
| 69 | LMO2 | 18358 | -11.460 | -0.1444 | Yes | ||
| 70 | MEF2C | 18390 | -12.088 | -0.0848 | Yes | ||
| 71 | DUSP10 | 18536 | -19.103 | 0.0043 | Yes |