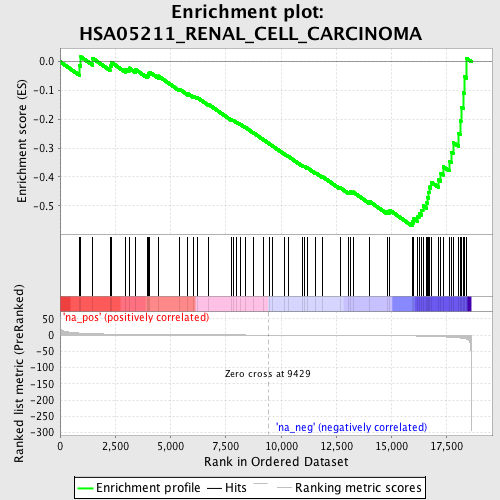
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_WTpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | HSA05211_RENAL_CELL_CARCINOMA |
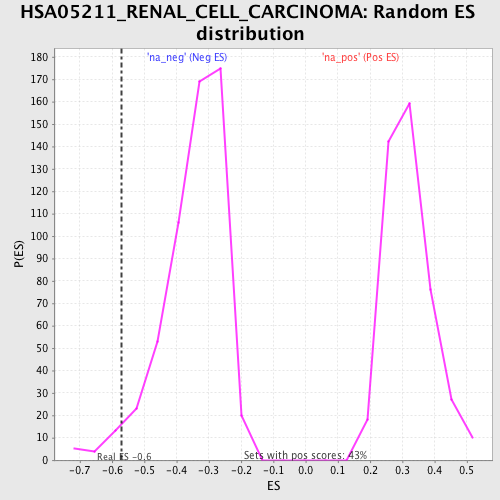
| Enrichment Score (ES) | -0.5701007 |
| Normalized Enrichment Score (NES) | -1.6427057 |
| Nominal p-value | 0.022887323 |
| FDR q-value | 0.46237373 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PIK3CB | 882 | 6.246 | -0.0133 | No | ||
| 2 | VEGFB | 938 | 6.030 | 0.0169 | No | ||
| 3 | PIK3R3 | 1472 | 4.347 | 0.0120 | No | ||
| 4 | CDC42 | 2263 | 2.900 | -0.0147 | No | ||
| 5 | CRK | 2332 | 2.817 | -0.0029 | No | ||
| 6 | AKT1 | 2974 | 2.198 | -0.0254 | No | ||
| 7 | EGLN1 | 3120 | 2.088 | -0.0218 | No | ||
| 8 | MAP2K1 | 3408 | 1.872 | -0.0269 | No | ||
| 9 | VEGFA | 3952 | 1.581 | -0.0475 | No | ||
| 10 | TCEB2 | 4007 | 1.551 | -0.0419 | No | ||
| 11 | RAPGEF1 | 4062 | 1.529 | -0.0365 | No | ||
| 12 | HIF1A | 4444 | 1.376 | -0.0494 | No | ||
| 13 | MAP2K2 | 5396 | 1.043 | -0.0950 | No | ||
| 14 | FIGF | 5786 | 0.929 | -0.1108 | No | ||
| 15 | PAK1 | 6045 | 0.855 | -0.1201 | No | ||
| 16 | RAP1A | 6204 | 0.813 | -0.1241 | No | ||
| 17 | PIK3R5 | 6715 | 0.682 | -0.1479 | No | ||
| 18 | PGF | 7744 | 0.419 | -0.2010 | No | ||
| 19 | PAK3 | 7834 | 0.395 | -0.2036 | No | ||
| 20 | PIK3R2 | 7997 | 0.358 | -0.2104 | No | ||
| 21 | PAK4 | 8161 | 0.314 | -0.2174 | No | ||
| 22 | TGFB2 | 8377 | 0.261 | -0.2276 | No | ||
| 23 | TCEB1 | 8771 | 0.168 | -0.2478 | No | ||
| 24 | HRAS | 9216 | 0.053 | -0.2715 | No | ||
| 25 | EPAS1 | 9487 | -0.019 | -0.2859 | No | ||
| 26 | NRAS | 9596 | -0.046 | -0.2915 | No | ||
| 27 | PAK7 | 10147 | -0.187 | -0.3201 | No | ||
| 28 | SOS1 | 10324 | -0.226 | -0.3284 | No | ||
| 29 | ARNT2 | 10976 | -0.380 | -0.3614 | No | ||
| 30 | HGF | 11038 | -0.393 | -0.3625 | No | ||
| 31 | ARAF | 11175 | -0.429 | -0.3675 | No | ||
| 32 | CUL2 | 11578 | -0.529 | -0.3862 | No | ||
| 33 | PIK3R1 | 11865 | -0.608 | -0.3983 | No | ||
| 34 | RBX1 | 12669 | -0.850 | -0.4369 | No | ||
| 35 | PTPN11 | 13063 | -0.961 | -0.4528 | No | ||
| 36 | MET | 13134 | -0.992 | -0.4512 | No | ||
| 37 | TGFA | 13256 | -1.034 | -0.4520 | No | ||
| 38 | PDGFB | 13980 | -1.314 | -0.4838 | No | ||
| 39 | RAF1 | 14806 | -1.685 | -0.5190 | No | ||
| 40 | RAC1 | 14926 | -1.756 | -0.5158 | No | ||
| 41 | TGFB3 | 15935 | -2.580 | -0.5559 | Yes | ||
| 42 | ETS1 | 16011 | -2.677 | -0.5453 | Yes | ||
| 43 | ARNT | 16185 | -2.887 | -0.5388 | Yes | ||
| 44 | PAK2 | 16246 | -2.958 | -0.5258 | Yes | ||
| 45 | SOS2 | 16367 | -3.158 | -0.5149 | Yes | ||
| 46 | PIK3CA | 16426 | -3.246 | -0.5002 | Yes | ||
| 47 | PIK3CD | 16600 | -3.558 | -0.4900 | Yes | ||
| 48 | GRB2 | 16640 | -3.636 | -0.4722 | Yes | ||
| 49 | SLC2A1 | 16684 | -3.714 | -0.4541 | Yes | ||
| 50 | VEGFC | 16707 | -3.748 | -0.4348 | Yes | ||
| 51 | CREBBP | 16786 | -3.900 | -0.4176 | Yes | ||
| 52 | JUN | 17135 | -4.762 | -0.4102 | Yes | ||
| 53 | CRKL | 17222 | -5.001 | -0.3874 | Yes | ||
| 54 | MAPK3 | 17335 | -5.344 | -0.3641 | Yes | ||
| 55 | RAP1B | 17617 | -6.283 | -0.3448 | Yes | ||
| 56 | TGFB1 | 17710 | -6.583 | -0.3136 | Yes | ||
| 57 | MAPK1 | 17787 | -6.868 | -0.2800 | Yes | ||
| 58 | AKT2 | 18046 | -8.193 | -0.2490 | Yes | ||
| 59 | AKT3 | 18107 | -8.687 | -0.2046 | Yes | ||
| 60 | PIK3CG | 18147 | -9.020 | -0.1572 | Yes | ||
| 61 | GAB1 | 18256 | -10.039 | -0.1079 | Yes | ||
| 62 | EGLN3 | 18317 | -10.740 | -0.0522 | Yes | ||
| 63 | KRAS | 18407 | -12.449 | 0.0113 | Yes |