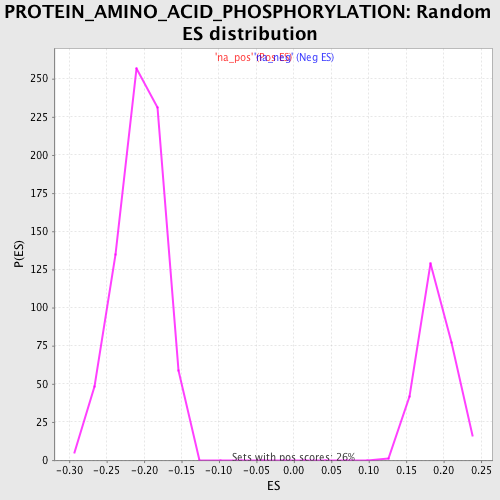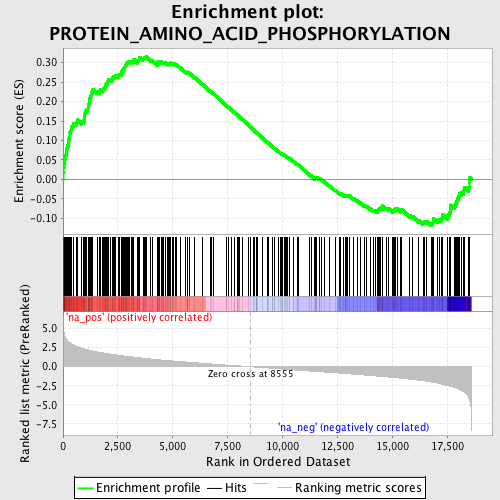
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | PROTEIN_AMINO_ACID_PHOSPHORYLATION |
| Enrichment Score (ES) | 0.31505203 |
| Normalized Enrichment Score (NES) | 1.6647406 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.45504692 |
| FWER p-Value | 0.999 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCND1 | 2 | 6.319 | 0.0185 | Yes | ||
| 2 | CDC2L2 | 34 | 4.486 | 0.0299 | Yes | ||
| 3 | TYK2 | 44 | 4.347 | 0.0422 | Yes | ||
| 4 | PRKCB1 | 67 | 4.060 | 0.0529 | Yes | ||
| 5 | MAP3K11 | 104 | 3.838 | 0.0623 | Yes | ||
| 6 | STK10 | 138 | 3.647 | 0.0712 | Yes | ||
| 7 | BTK | 166 | 3.564 | 0.0802 | Yes | ||
| 8 | CAMK2A | 208 | 3.401 | 0.0879 | Yes | ||
| 9 | STK11 | 253 | 3.248 | 0.0951 | Yes | ||
| 10 | RPS6KA4 | 265 | 3.219 | 0.1039 | Yes | ||
| 11 | MAST2 | 305 | 3.135 | 0.1110 | Yes | ||
| 12 | PCTK1 | 310 | 3.130 | 0.1200 | Yes | ||
| 13 | MARK2 | 357 | 3.024 | 0.1264 | Yes | ||
| 14 | WNK1 | 365 | 3.008 | 0.1348 | Yes | ||
| 15 | ROCK2 | 450 | 2.892 | 0.1387 | Yes | ||
| 16 | CCNT1 | 490 | 2.842 | 0.1450 | Yes | ||
| 17 | JAK2 | 591 | 2.705 | 0.1475 | Yes | ||
| 18 | ERC1 | 634 | 2.654 | 0.1530 | Yes | ||
| 19 | CLCF1 | 840 | 2.447 | 0.1490 | Yes | ||
| 20 | TNK2 | 921 | 2.374 | 0.1516 | Yes | ||
| 21 | IL3 | 958 | 2.345 | 0.1566 | Yes | ||
| 22 | PKN1 | 978 | 2.335 | 0.1624 | Yes | ||
| 23 | CSNK1G2 | 995 | 2.323 | 0.1683 | Yes | ||
| 24 | VRK1 | 1001 | 2.317 | 0.1749 | Yes | ||
| 25 | ROCK1 | 1057 | 2.284 | 0.1786 | Yes | ||
| 26 | NEK4 | 1134 | 2.203 | 0.1809 | Yes | ||
| 27 | MAP4K2 | 1136 | 2.202 | 0.1873 | Yes | ||
| 28 | TWF1 | 1149 | 2.194 | 0.1931 | Yes | ||
| 29 | MKNK2 | 1184 | 2.166 | 0.1976 | Yes | ||
| 30 | PICK1 | 1191 | 2.160 | 0.2037 | Yes | ||
| 31 | ABI3 | 1220 | 2.138 | 0.2084 | Yes | ||
| 32 | EIF2AK4 | 1253 | 2.118 | 0.2129 | Yes | ||
| 33 | MAP3K2 | 1271 | 2.105 | 0.2182 | Yes | ||
| 34 | MAPK3 | 1303 | 2.079 | 0.2226 | Yes | ||
| 35 | RAGE | 1327 | 2.062 | 0.2274 | Yes | ||
| 36 | TNKS | 1359 | 2.044 | 0.2317 | Yes | ||
| 37 | MAP2K6 | 1583 | 1.919 | 0.2252 | Yes | ||
| 38 | GSG2 | 1657 | 1.881 | 0.2267 | Yes | ||
| 39 | GMFG | 1685 | 1.864 | 0.2308 | Yes | ||
| 40 | MYOD1 | 1788 | 1.816 | 0.2305 | Yes | ||
| 41 | MARK4 | 1821 | 1.798 | 0.2341 | Yes | ||
| 42 | NF2 | 1899 | 1.761 | 0.2351 | Yes | ||
| 43 | PRKCD | 1917 | 1.749 | 0.2393 | Yes | ||
| 44 | SRPK1 | 1948 | 1.732 | 0.2427 | Yes | ||
| 45 | CSK | 1969 | 1.722 | 0.2467 | Yes | ||
| 46 | MINK1 | 2030 | 1.699 | 0.2484 | Yes | ||
| 47 | EPHB2 | 2043 | 1.695 | 0.2527 | Yes | ||
| 48 | DYRK1A | 2046 | 1.693 | 0.2576 | Yes | ||
| 49 | PIM2 | 2153 | 1.648 | 0.2567 | Yes | ||
| 50 | TLK2 | 2242 | 1.609 | 0.2566 | Yes | ||
| 51 | MAP3K3 | 2251 | 1.606 | 0.2609 | Yes | ||
| 52 | SRP72 | 2282 | 1.591 | 0.2639 | Yes | ||
| 53 | TLK1 | 2351 | 1.563 | 0.2648 | Yes | ||
| 54 | ULK1 | 2377 | 1.550 | 0.2680 | Yes | ||
| 55 | STK4 | 2510 | 1.502 | 0.2652 | Yes | ||
| 56 | MATK | 2547 | 1.488 | 0.2677 | Yes | ||
| 57 | SGK3 | 2575 | 1.476 | 0.2705 | Yes | ||
| 58 | STK16 | 2651 | 1.446 | 0.2707 | Yes | ||
| 59 | FES | 2667 | 1.442 | 0.2741 | Yes | ||
| 60 | SNF1LK | 2669 | 1.442 | 0.2783 | Yes | ||
| 61 | MAP3K8 | 2710 | 1.426 | 0.2803 | Yes | ||
| 62 | MKNK1 | 2750 | 1.409 | 0.2823 | Yes | ||
| 63 | RAF1 | 2767 | 1.403 | 0.2856 | Yes | ||
| 64 | PRKACB | 2804 | 1.387 | 0.2877 | Yes | ||
| 65 | FASTK | 2820 | 1.380 | 0.2909 | Yes | ||
| 66 | IRAK2 | 2851 | 1.369 | 0.2933 | Yes | ||
| 67 | TEC | 2878 | 1.359 | 0.2959 | Yes | ||
| 68 | CCND2 | 2900 | 1.351 | 0.2987 | Yes | ||
| 69 | JAK3 | 2916 | 1.346 | 0.3018 | Yes | ||
| 70 | CCL2 | 2994 | 1.321 | 0.3015 | Yes | ||
| 71 | CDC42BPA | 3041 | 1.308 | 0.3029 | Yes | ||
| 72 | EIF2AK3 | 3099 | 1.293 | 0.3036 | Yes | ||
| 73 | ITGB2 | 3183 | 1.262 | 0.3028 | Yes | ||
| 74 | PRKX | 3215 | 1.250 | 0.3047 | Yes | ||
| 75 | ABL2 | 3224 | 1.247 | 0.3080 | Yes | ||
| 76 | IL12A | 3393 | 1.195 | 0.3023 | Yes | ||
| 77 | PRKG1 | 3395 | 1.194 | 0.3058 | Yes | ||
| 78 | CTBP1 | 3447 | 1.175 | 0.3065 | Yes | ||
| 79 | CRKRS | 3459 | 1.173 | 0.3093 | Yes | ||
| 80 | MAP3K9 | 3485 | 1.163 | 0.3114 | Yes | ||
| 81 | CHUK | 3488 | 1.162 | 0.3147 | Yes | ||
| 82 | GRK1 | 3641 | 1.113 | 0.3097 | Yes | ||
| 83 | PIM1 | 3685 | 1.096 | 0.3105 | Yes | ||
| 84 | BMX | 3687 | 1.095 | 0.3137 | Yes | ||
| 85 | PKN3 | 3753 | 1.076 | 0.3133 | Yes | ||
| 86 | CCL11 | 3780 | 1.068 | 0.3151 | Yes | ||
| 87 | SNF1LK2 | 4003 | 1.002 | 0.3059 | No | ||
| 88 | PLK3 | 4067 | 0.984 | 0.3054 | No | ||
| 89 | ERBB3 | 4288 | 0.929 | 0.2961 | No | ||
| 90 | MAP4K5 | 4309 | 0.923 | 0.2978 | No | ||
| 91 | CAMK4 | 4316 | 0.922 | 0.3001 | No | ||
| 92 | LTK | 4329 | 0.919 | 0.3022 | No | ||
| 93 | CAMK1 | 4388 | 0.902 | 0.3017 | No | ||
| 94 | PDPK1 | 4391 | 0.901 | 0.3042 | No | ||
| 95 | PTK2B | 4470 | 0.877 | 0.3025 | No | ||
| 96 | CARD14 | 4548 | 0.857 | 0.3009 | No | ||
| 97 | TSSK3 | 4585 | 0.850 | 0.3014 | No | ||
| 98 | FYB | 4655 | 0.836 | 0.3001 | No | ||
| 99 | PRKCZ | 4746 | 0.809 | 0.2976 | No | ||
| 100 | IRAK3 | 4811 | 0.793 | 0.2964 | No | ||
| 101 | ERN2 | 4851 | 0.783 | 0.2966 | No | ||
| 102 | RET | 4873 | 0.777 | 0.2978 | No | ||
| 103 | IL31RA | 4887 | 0.774 | 0.2993 | No | ||
| 104 | PAK1 | 4985 | 0.751 | 0.2962 | No | ||
| 105 | AURKC | 4998 | 0.748 | 0.2978 | No | ||
| 106 | MAP4K3 | 5030 | 0.741 | 0.2983 | No | ||
| 107 | FGR | 5123 | 0.719 | 0.2954 | No | ||
| 108 | CCND3 | 5169 | 0.709 | 0.2950 | No | ||
| 109 | HCLS1 | 5328 | 0.668 | 0.2884 | No | ||
| 110 | HCK | 5588 | 0.613 | 0.2761 | No | ||
| 111 | CBLC | 5647 | 0.602 | 0.2747 | No | ||
| 112 | CDK2AP1 | 5679 | 0.596 | 0.2748 | No | ||
| 113 | TXK | 5686 | 0.595 | 0.2762 | No | ||
| 114 | MAPK12 | 5765 | 0.579 | 0.2736 | No | ||
| 115 | PCTK2 | 6008 | 0.521 | 0.2620 | No | ||
| 116 | MAP3K6 | 6361 | 0.443 | 0.2442 | No | ||
| 117 | MAP4K4 | 6702 | 0.368 | 0.2267 | No | ||
| 118 | EPHA5 | 6749 | 0.359 | 0.2253 | No | ||
| 119 | CSNK1D | 6835 | 0.340 | 0.2217 | No | ||
| 120 | TNK1 | 7455 | 0.208 | 0.1886 | No | ||
| 121 | ACVRL1 | 7464 | 0.206 | 0.1888 | No | ||
| 122 | BCR | 7556 | 0.191 | 0.1844 | No | ||
| 123 | CDKL5 | 7682 | 0.170 | 0.1781 | No | ||
| 124 | FRK | 7800 | 0.148 | 0.1721 | No | ||
| 125 | MYLK | 7818 | 0.144 | 0.1716 | No | ||
| 126 | SNRK | 7925 | 0.122 | 0.1662 | No | ||
| 127 | MAP4K1 | 7938 | 0.119 | 0.1659 | No | ||
| 128 | LYN | 7998 | 0.109 | 0.1630 | No | ||
| 129 | TRIB2 | 8056 | 0.099 | 0.1602 | No | ||
| 130 | DAPK1 | 8175 | 0.078 | 0.1540 | No | ||
| 131 | LIMK1 | 8436 | 0.023 | 0.1399 | No | ||
| 132 | RIPK4 | 8535 | 0.003 | 0.1346 | No | ||
| 133 | EPHA8 | 8670 | -0.026 | 0.1274 | No | ||
| 134 | DAPK2 | 8742 | -0.039 | 0.1236 | No | ||
| 135 | BMP4 | 8794 | -0.048 | 0.1210 | No | ||
| 136 | TGFB1 | 8803 | -0.050 | 0.1207 | No | ||
| 137 | CAMK2B | 8806 | -0.050 | 0.1208 | No | ||
| 138 | DAPK3 | 8873 | -0.064 | 0.1174 | No | ||
| 139 | PRKCH | 9066 | -0.105 | 0.1072 | No | ||
| 140 | AKT1 | 9294 | -0.158 | 0.0953 | No | ||
| 141 | TESK2 | 9315 | -0.163 | 0.0947 | No | ||
| 142 | CAMKK2 | 9342 | -0.167 | 0.0938 | No | ||
| 143 | CSNK2A1 | 9542 | -0.208 | 0.0836 | No | ||
| 144 | STK38 | 9546 | -0.209 | 0.0840 | No | ||
| 145 | ERG | 9635 | -0.228 | 0.0799 | No | ||
| 146 | PRKCE | 9636 | -0.228 | 0.0806 | No | ||
| 147 | MYO3A | 9807 | -0.266 | 0.0721 | No | ||
| 148 | GSK3B | 9910 | -0.287 | 0.0674 | No | ||
| 149 | IGF1R | 9965 | -0.299 | 0.0653 | No | ||
| 150 | TRIB1 | 10013 | -0.312 | 0.0637 | No | ||
| 151 | STK17B | 10014 | -0.312 | 0.0646 | No | ||
| 152 | LIPE | 10088 | -0.328 | 0.0616 | No | ||
| 153 | IL20 | 10132 | -0.337 | 0.0603 | No | ||
| 154 | IKBKE | 10197 | -0.350 | 0.0578 | No | ||
| 155 | PRKCI | 10237 | -0.360 | 0.0567 | No | ||
| 156 | CDK9 | 10324 | -0.380 | 0.0532 | No | ||
| 157 | TSSK1A | 10331 | -0.381 | 0.0540 | No | ||
| 158 | SOCS1 | 10482 | -0.409 | 0.0470 | No | ||
| 159 | ADAM10 | 10666 | -0.446 | 0.0384 | No | ||
| 160 | MAPKAPK2 | 10672 | -0.448 | 0.0394 | No | ||
| 161 | PTK6 | 10738 | -0.461 | 0.0372 | No | ||
| 162 | ABI2 | 11227 | -0.561 | 0.0123 | No | ||
| 163 | EGFR | 11316 | -0.578 | 0.0092 | No | ||
| 164 | MERTK | 11478 | -0.610 | 0.0022 | No | ||
| 165 | EIF2AK1 | 11482 | -0.611 | 0.0039 | No | ||
| 166 | TSSK6 | 11492 | -0.613 | 0.0052 | No | ||
| 167 | ABL1 | 11541 | -0.621 | 0.0044 | No | ||
| 168 | CD80 | 11565 | -0.625 | 0.0050 | No | ||
| 169 | MAP3K12 | 11663 | -0.645 | 0.0016 | No | ||
| 170 | IGFBP3 | 11687 | -0.647 | 0.0022 | No | ||
| 171 | STAT1 | 11783 | -0.667 | -0.0010 | No | ||
| 172 | LMTK2 | 11917 | -0.693 | -0.0062 | No | ||
| 173 | PRKG2 | 12155 | -0.736 | -0.0169 | No | ||
| 174 | CDC42BPB | 12395 | -0.789 | -0.0276 | No | ||
| 175 | PRKD1 | 12618 | -0.841 | -0.0372 | No | ||
| 176 | COL4A3BP | 12657 | -0.848 | -0.0368 | No | ||
| 177 | TGFBR1 | 12782 | -0.875 | -0.0410 | No | ||
| 178 | TDGF1 | 12856 | -0.891 | -0.0423 | No | ||
| 179 | CSNK1A1 | 12887 | -0.897 | -0.0413 | No | ||
| 180 | ABI1 | 12929 | -0.904 | -0.0409 | No | ||
| 181 | MAST1 | 12973 | -0.916 | -0.0405 | No | ||
| 182 | MAP3K10 | 13057 | -0.932 | -0.0423 | No | ||
| 183 | MAPK4 | 13237 | -0.973 | -0.0492 | No | ||
| 184 | SRPK2 | 13395 | -1.008 | -0.0548 | No | ||
| 185 | BMPR1B | 13550 | -1.041 | -0.0601 | No | ||
| 186 | AURKA | 13742 | -1.086 | -0.0673 | No | ||
| 187 | AKTIP | 13835 | -1.109 | -0.0691 | No | ||
| 188 | INSR | 13987 | -1.145 | -0.0739 | No | ||
| 189 | ALS2CR2 | 14156 | -1.185 | -0.0796 | No | ||
| 190 | ILK | 14233 | -1.204 | -0.0802 | No | ||
| 191 | PRPF4B | 14315 | -1.224 | -0.0810 | No | ||
| 192 | HIPK3 | 14349 | -1.232 | -0.0792 | No | ||
| 193 | IL5 | 14372 | -1.238 | -0.0767 | No | ||
| 194 | BMPR1A | 14412 | -1.250 | -0.0752 | No | ||
| 195 | PLK4 | 14449 | -1.257 | -0.0734 | No | ||
| 196 | TSSK2 | 14540 | -1.279 | -0.0746 | No | ||
| 197 | UHMK1 | 14561 | -1.284 | -0.0719 | No | ||
| 198 | MAPK8 | 14563 | -1.284 | -0.0682 | No | ||
| 199 | NEK11 | 14740 | -1.323 | -0.0739 | No | ||
| 200 | MAPK6 | 14840 | -1.347 | -0.0753 | No | ||
| 201 | WNK4 | 15018 | -1.394 | -0.0808 | No | ||
| 202 | DYRK1B | 15059 | -1.401 | -0.0789 | No | ||
| 203 | RPS6KA5 | 15107 | -1.413 | -0.0773 | No | ||
| 204 | PINK1 | 15158 | -1.428 | -0.0758 | No | ||
| 205 | PRKD3 | 15233 | -1.444 | -0.0756 | No | ||
| 206 | ACVR1B | 15394 | -1.499 | -0.0799 | No | ||
| 207 | IL22RA2 | 15410 | -1.505 | -0.0763 | No | ||
| 208 | PAK2 | 15809 | -1.628 | -0.0932 | No | ||
| 209 | BRSK2 | 15933 | -1.673 | -0.0950 | No | ||
| 210 | PRKAA1 | 16211 | -1.769 | -0.1049 | No | ||
| 211 | TTN | 16405 | -1.833 | -0.1100 | No | ||
| 212 | MARK1 | 16453 | -1.851 | -0.1071 | No | ||
| 213 | NEK6 | 16584 | -1.910 | -0.1086 | No | ||
| 214 | DYRK3 | 16791 | -1.999 | -0.1139 | No | ||
| 215 | PXK | 16848 | -2.033 | -0.1110 | No | ||
| 216 | F2 | 16859 | -2.037 | -0.1055 | No | ||
| 217 | STK38L | 16863 | -2.038 | -0.0997 | No | ||
| 218 | LATS2 | 17066 | -2.144 | -0.1044 | No | ||
| 219 | PMVK | 17145 | -2.191 | -0.1022 | No | ||
| 220 | PRDX4 | 17242 | -2.251 | -0.1008 | No | ||
| 221 | CD81 | 17273 | -2.273 | -0.0958 | No | ||
| 222 | CDKL1 | 17305 | -2.292 | -0.0907 | No | ||
| 223 | VRK2 | 17528 | -2.440 | -0.0957 | No | ||
| 224 | MAP3K13 | 17541 | -2.451 | -0.0891 | No | ||
| 225 | NDUFS4 | 17625 | -2.531 | -0.0862 | No | ||
| 226 | ERBB2 | 17637 | -2.544 | -0.0793 | No | ||
| 227 | ICK | 17653 | -2.556 | -0.0726 | No | ||
| 228 | IKBKAP | 17667 | -2.566 | -0.0658 | No | ||
| 229 | LCK | 17843 | -2.721 | -0.0673 | No | ||
| 230 | PRKAG1 | 17877 | -2.749 | -0.0611 | No | ||
| 231 | GMFB | 17939 | -2.821 | -0.0561 | No | ||
| 232 | F2R | 17972 | -2.869 | -0.0494 | No | ||
| 233 | ACVR1 | 18021 | -2.923 | -0.0434 | No | ||
| 234 | PKN2 | 18056 | -2.965 | -0.0366 | No | ||
| 235 | WNK3 | 18167 | -3.132 | -0.0334 | No | ||
| 236 | ERN1 | 18265 | -3.292 | -0.0290 | No | ||
| 237 | AKT3 | 18272 | -3.301 | -0.0196 | No | ||
| 238 | STK3 | 18491 | -4.032 | -0.0196 | No | ||
| 239 | STK19 | 18535 | -4.455 | -0.0089 | No | ||
| 240 | CSNK1E | 18540 | -4.504 | 0.0041 | No |