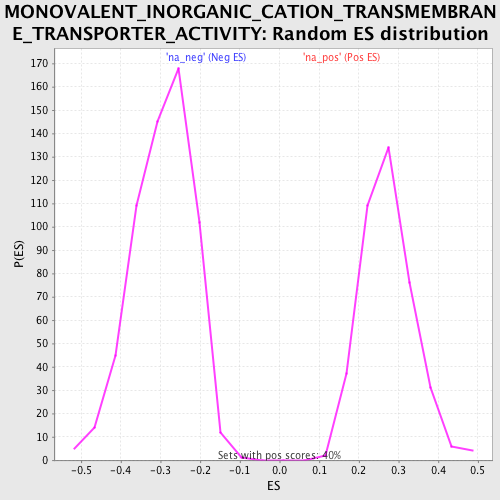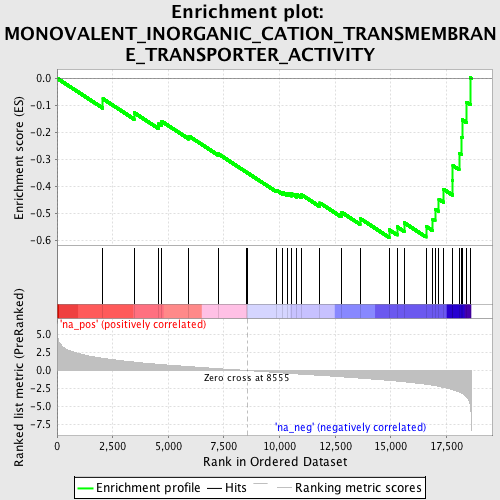
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | MONOVALENT_INORGANIC_CATION_TRANSMEMBRANE_TRANSPORTER_ACTIVITY |
| Enrichment Score (ES) | -0.58883727 |
| Normalized Enrichment Score (NES) | -2.002691 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.023575075 |
| FWER p-Value | 0.126 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SLC13A4 | 2061 | 1.688 | -0.0761 | No | ||
| 2 | COX10 | 3462 | 1.172 | -0.1272 | No | ||
| 3 | COX7A2L | 4564 | 0.854 | -0.1689 | No | ||
| 4 | COX8A | 4698 | 0.822 | -0.1591 | No | ||
| 5 | COX4I2 | 5923 | 0.543 | -0.2137 | No | ||
| 6 | SLC9A3 | 7243 | 0.253 | -0.2795 | No | ||
| 7 | UQCRC1 | 8522 | 0.006 | -0.3481 | No | ||
| 8 | ATP4B | 8544 | 0.002 | -0.3492 | No | ||
| 9 | ATP6V0C | 9859 | -0.276 | -0.4142 | No | ||
| 10 | SLC9A7 | 10134 | -0.337 | -0.4220 | No | ||
| 11 | ATP6V1B2 | 10348 | -0.385 | -0.4255 | No | ||
| 12 | COX11 | 10536 | -0.420 | -0.4269 | No | ||
| 13 | SLC9A2 | 10771 | -0.469 | -0.4299 | No | ||
| 14 | COX7B | 10983 | -0.515 | -0.4306 | No | ||
| 15 | SLC9A6 | 11813 | -0.673 | -0.4613 | No | ||
| 16 | ATP4A | 12773 | -0.873 | -0.4949 | No | ||
| 17 | COX5A | 13617 | -1.060 | -0.5184 | No | ||
| 18 | UQCRH | 14927 | -1.368 | -0.5606 | Yes | ||
| 19 | COX5B | 15284 | -1.461 | -0.5496 | Yes | ||
| 20 | COX15 | 15601 | -1.563 | -0.5344 | Yes | ||
| 21 | ATP6V0E1 | 16583 | -1.910 | -0.5478 | Yes | ||
| 22 | ACCN4 | 16889 | -2.053 | -0.5219 | Yes | ||
| 23 | COX7A1 | 16987 | -2.101 | -0.4837 | Yes | ||
| 24 | SLC8A1 | 17162 | -2.201 | -0.4477 | Yes | ||
| 25 | ABCC8 | 17362 | -2.338 | -0.4102 | Yes | ||
| 26 | KCNA1 | 17773 | -2.652 | -0.3775 | Yes | ||
| 27 | NNT | 17791 | -2.673 | -0.3233 | Yes | ||
| 28 | SURF1 | 18068 | -2.986 | -0.2765 | Yes | ||
| 29 | ABCB11 | 18177 | -3.143 | -0.2175 | Yes | ||
| 30 | COX4I1 | 18211 | -3.196 | -0.1534 | Yes | ||
| 31 | CYB5A | 18407 | -3.651 | -0.0885 | Yes | ||
| 32 | ATP6V1C1 | 18570 | -4.835 | 0.0025 | Yes |