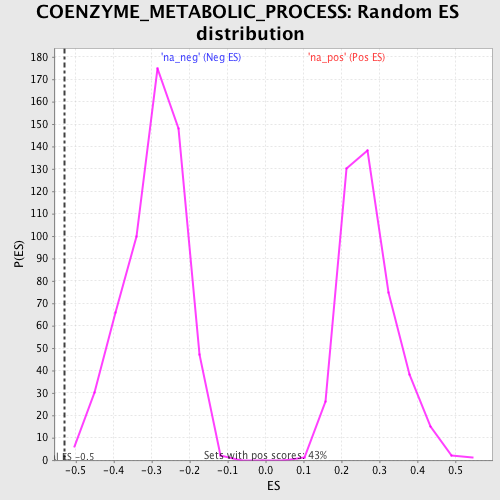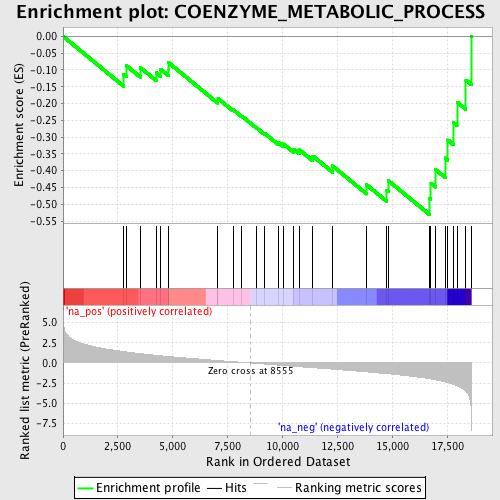
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMpreB_versus_DMpreB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_neg |
| GeneSet | COENZYME_METABOLIC_PROCESS |
| Enrichment Score (ES) | -0.5303094 |
| Normalized Enrichment Score (NES) | -1.806572 |
| Nominal p-value | 0.0017421603 |
| FDR q-value | 0.07428543 |
| FWER p-Value | 0.774 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GCLC | 2758 | 1.406 | -0.1139 | No | ||
| 2 | ACLY | 2882 | 1.357 | -0.0872 | No | ||
| 3 | ALDH1L1 | 3516 | 1.153 | -0.0929 | No | ||
| 4 | GPX1 | 4238 | 0.938 | -0.1087 | No | ||
| 5 | ACOT12 | 4440 | 0.886 | -0.0978 | No | ||
| 6 | GLYAT | 4800 | 0.797 | -0.0975 | No | ||
| 7 | ACOT2 | 4802 | 0.796 | -0.0781 | No | ||
| 8 | GSTA1 | 7043 | 0.294 | -0.1914 | No | ||
| 9 | PGD | 7058 | 0.289 | -0.1850 | No | ||
| 10 | SDHA | 7744 | 0.158 | -0.2180 | No | ||
| 11 | GCLM | 8150 | 0.082 | -0.2378 | No | ||
| 12 | SDHD | 8824 | -0.054 | -0.2727 | No | ||
| 13 | PDSS1 | 9191 | -0.133 | -0.2891 | No | ||
| 14 | COQ3 | 9802 | -0.265 | -0.3154 | No | ||
| 15 | MTHFD2 | 10038 | -0.317 | -0.3203 | No | ||
| 16 | FTCD | 10518 | -0.415 | -0.3358 | No | ||
| 17 | ACO2 | 10777 | -0.470 | -0.3382 | No | ||
| 18 | CTNS | 11384 | -0.591 | -0.3563 | No | ||
| 19 | COASY | 12280 | -0.764 | -0.3857 | No | ||
| 20 | ACOT4 | 13836 | -1.110 | -0.4421 | No | ||
| 21 | MLYCD | 14745 | -1.323 | -0.4585 | No | ||
| 22 | ACOT9 | 14824 | -1.343 | -0.4297 | No | ||
| 23 | SDHC | 16695 | -1.962 | -0.4821 | Yes | ||
| 24 | SOD1 | 16765 | -1.988 | -0.4371 | Yes | ||
| 25 | PGLS | 16981 | -2.098 | -0.3971 | Yes | ||
| 26 | PDHB | 17412 | -2.373 | -0.3620 | Yes | ||
| 27 | GLRX2 | 17534 | -2.442 | -0.3086 | Yes | ||
| 28 | NNT | 17791 | -2.673 | -0.2567 | Yes | ||
| 29 | MOCS2 | 17954 | -2.839 | -0.1957 | Yes | ||
| 30 | COQ7 | 18359 | -3.556 | -0.1302 | Yes | ||
| 31 | SDHB | 18604 | -5.865 | 0.0006 | Yes |